Abstract
Since the revolutionary discovery of the CRISPR-Cas technology for programmable genome editing, its range of applications has been extended by multiple biotechnological tools that go far beyond its original function as “genetic scissors”. One of these further developments of the CRISPR-Cas system allows genes to be activated in a targeted and efficient manner. These gene-activating CRISPR-Cas modules (CRISPRa) are based on a programmable recruitment of transcription factors to specific loci and offer several key advantages that make them particularly attractive for therapeutic applications. These advantages include inter alia low off-target effects, independence of the target gene size as well as the potential to develop gene- and mutation-independent therapeutic strategies. Herein, I will give an overview on the currently available CRISPRa modules and discuss recent developments, future potentials and limitations of this approach with a focus on therapeutic applications and in vivo delivery.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Main text
CRISPR-Cas tools for therapy
The discovery of the CRISPR (clustered regularly interspaced short palindromic repeats)/Cas (CRISPR-associated protein) system has revolutionized the way genetic or acquired diseases can be treated. For its natural function in adaptive bacterial immune defense against viruses, the Cas enzyme requires CRISPR RNA (crRNA) that contains sequence segments from the viral genome responsible for binding to the target virus-derived DNA (spacer), and trans-activating crRNA (tracrRNA), which originates from the bacterial genome. tracrRNA and crRNA act together to recruit the Cas complex to the viral genome so that it can be cut and rendered harmless [1, 2]. For programmable manipulation of the bacterial and mammalian cell genomes, single guide (sg)RNA was developed by fusing the tracr and crRNA elements [1]. Since then, a wide range of different CRISPR-Cas derivatives were developed which differ in the species from which the Cas enzyme is derived, the architecture of the single guide RNAs, the number and type of proteins fused to Cas, or the endonuclease activity of the enzyme [3, 4]. Since its FDA-approval for investigational new drug application in 2019 for treatment of Leber congenital amaurosis, a severe form of retinal dystrophy caused by a mutation in the CEP290 gene (ClinicalTrials.gov Identifier: NCT03872479), the CRISPR-Cas technology has finally cleared the crucial regulatory hurdle for human application. Since then it has been applied in several clinical trials utilizing CRISPR-Cas to target genetic or acquired diseases (NCT04601051, NCT04191148, NCT03057912).
The currently available CRISPR-Cas modules can be used to excise or correct genome segments or to induce changes in the gene expression at specific loci [3, 4]. The gene transcription-modifying Cas modules typically contain the endonuclease-deficient Cas9 protein from Streptococcus pyogenes (SpdCas9), to which functional domains capable of transcriptional activation (CRISPRa) or repression (CRISPRi) are fused. Endonuclease deficiency of Cas9 is usually achieved by introducing mutations in the corresponding functional domains of Cas9 [1, 5]. However, the enzyme also loses its nuclease activity in the presence of sgRNAs with truncated spacer sequences (< 16 bp) allowing the development of more sophisticated strategies for the treatment of specific diseases [6]. Transcriptional changes can e.g. be induced by recruitment of gene activating or repressive transcriptional factors or indirectly, i.e., by modifying the chromatin structure. While this review focuses on CRISPRa, it is worth noting here that the basic principles regarding the rational design of individual CRISPRa modules, their application spectra as well as their advantages and disadvantages largely apply to their CRISPRi counterparts.
CRISPRa modules for epigenome editing
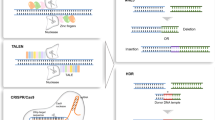
The proof-of-concept for programmable gene activation was established well before the CRISPR-Cas era using zinc-finger nucleases (ZFNs) and transcription activator-like effector nucleases (TALENs) fused to trans-activating domains [7]. One advantage of gene activation via ZFNs and TALENs is the fact that these modules do not require dual AAV vectors for in vivo expression due to the small size of these proteins. Nevertheless, ZFNs and TALENs require a design of new protein arrays for each new target sequence in the genome. In comparison, the CRISPRa system allows for a much simpler design, as target specificity can be achieved only by replacing the protospacer sequence within the sgRNA cassette. Another key advantage of CRISPR-Cas-based systems over ZFNs or TALENs is the ease of multiplexing, i.e., editing multiple genes simultaneously using different sgRNAs [8]. The first CRISPRa module described consists of dCas9 fused at its C-terminus to four VP16 transcription activating (trans-activating) domains derived from the herpes simplex virus (dCas9-VP64) [9, 10]. dCas9-VP64 is characterized by low to moderate gene activation efficiency. To increase gene activation and thus expand the range of applications of CRISPRa, more effective CRISPRa modules have been engineered by fusing additional trans-activating domains or their binding sites to dCas9. This has led to the development of several second-generation CRISPRa variants, of which some lead representatives will be described in this review. One important representative is the dCas9-VPR module, in which the NF-κB (nuclear factor kappa-light-chain-enhancer of activated B cells) p65 subunit and R transactivator (Rta) domains from Epstein-Barr virus were fused to dCas9-VP64 [11]. By comparison, the dCas9-SunTag module contains a tag with multiple binding sites for VP64 [12]. In other CRISPRa systems, in addition to fusing trans-activating domains to the Cas protein, the sgRNAs tetraloops and stemloops have been modified to contain binding sites (protein binding RNA aptamers) for other trans-activating molecules. This includes the dCas9-SAM (synergistic activation mediator) module, which consists of dCas9-VP64 and an additional fusion protein containing the coat protein of the RNA bacteriophage MS2, p65, and the Heat Shock Transcription Factor 1 (HSF1). This MS2-p65-HSF1 (MPH) complex can bind to the tetraloops and stemloops of the modified sgRNA via MS2 and facilitate transcription via p65 and HSF1 domains [13]. Recently, different CRISPRa modules were compared side-by-side with the result that dCas9-VPR, dCas9-SAM, and dCas9-SunTag consistently provided the highest gene activation efficiencies between different cell types and species [14]. Comparing only these three most potent modules with each other, the trans activation efficiency was largely gene-specific. These modules have been applied in numerous in vitro studies to date, however only a few in vivo studies exist. In the translationally focused in vivo studies to date, CRISPRa has been successfully applied to treat genetic or acquired diseases such as diabetes mellitus, cancer, epilepsy, Dravet syndrome, muscular dystrophy, acute kidney injury, obesity or retinitis pigmentosa [15,16,17,18,19,20,21,22]. In this context, recombinant adeno-associated viral (rAAV) vectors were used to express Staphylococcus aureus (Sa)dCas9-VP64 [18, 20], SpdCas9-VP64, Sp(d)Cas9-MPH, a truncated variant of the dCas9-SAM module lacking the VP64 domain [19], and SpdCas9-VPR [15] (Fig. 1). In the study by Matharu et al., two different AAV vectors were used to rescue the obesity phenotype in the heterozygous loss of function Sim1 mouse model by overactivation of the remaining functional allele. The first vector expresses SpCas9-VP64 or SadCas9-2xVP64 and the second one the sgRNA cassette. A similar SpCas9-VP64 dual AAV vector system was used in two recent studies to activate the Kcn1 or the Scn1a gene to treat the respective mouse model for epilepsy or Dravet syndrome [16, 17]. From the dual AAV vector perspective, the main difference from the Mathary et al. study was that SpCas9-VP64 expression was driven by a doxycycline-dependent TetOn (TRE) promoter (Fig. 1A). Yamagata et al. also established a therapy for Dravet syndrome in the Scn1a-haplodeficient mouse model by overactivating the intact endogenous Scn1a locus. For this purpose, the authors crossed transgenic mice expressing dCas9-VPR with Scn1a-haplodeficient mice and applied only the Scn1a-targeting sgRNAs via AAV vectors [22]. Nevertheless, the results from CRISPRa-expressing transgenic mice, although helpful to provide proof-of-concept, are less meaningful from a therapeutic perspective, as such a strategy bypasses CRISPRa delivery, which is one of the major hurdles to translation. In comparison, Kemaledawi et al. either used (i) a single rAAV vector expressing SadCas9-VP64 and one sgRNA or (ii) two separate AAVs, the first encoding SadCas9-VP64 and the second expressing three sgRNAs, to activate the Lama1 gene in a mouse model of congenital muscular dystrophy type 1A (MDC1A). This disease is caused by a mutation in Lama2 gene, which is functionally related to Lama1 (Fig. 1B). In another study, Böhm et al. used dual split-intein-based AAV vectors to activate the rhodopsin-related Opn1mw (middle wavelength opsin, M-opsin) gene in rod photoreceptors in the rhodopsin-deficient mouse model for retinitis pigmentosa, the most common form of inherited retinal blindness. The first vector contains three sgRNAs and one half of dCas9-VPR fused to a split-intein, and the second one encodes the remaining half of the dCas9-VPR-intein fusion protein. In this approach, the split-inteins are required for scarless reconstitution of SpdCas9-VPR when dual rAAVs are co-administered (Fig. 1C). Split-intein-based dual AAV vectors were also used in another study to express a SpdCas9 CRISPRa version containing VP64 at the C- and N-termini of Cas9 and additionally the p65 domain at the C-terminus. Using this approach, moderate activation of the Afp locus was observed in the mouse retina, but the therapeutic potential of this activation was not investigated [23]. In the study by Liao et al., most in vivo experiments were performed in transgenic mice expressing Cas9. These mice were crossed with mouse models for different diseases and injected with an AAV vector expressing gene-specific sgRNAs and MPH to investigate the basic potential of gene activation for the therapy of the respective disorders. In the injected mice, activation of the follistatin (Fst), utrophin, or klotho genes was shown to increase the muscle mass or mitigate the muscle wasting phenotype (e.g., Duchenne muscular dystrophy) or, in the case of klotho, to attenuate the disease phenotype of acute kidney injury. In addition, the pancreatic and duodenal homeobox gene 1 (Pdx1) was also activated in liver cells to induce the formation of insulin-producing cells, which could represent a new therapeutic option for diabetes mellitus type I. Recently, another study utilized a very similar SpdCas9-MPH-based dual AAV vector strategy for multiplexed activation of endogenous genes to induce an anti-tumor adaptive immunity [21] (Fig. 1D).
Overview of CRISPRa modules that have been successfully applied to treat genetic or acquired diseases. Right panel, strategies and expression cassettes used for in vivo delivery of the individual modules. The red asterisk highlights the expression cassette which exceeds the packaging capacity of rAAV vectors (4.7 kb). TRE tetracycline response element. UbProm ubiquitous promoter (e.g., CAG or EF1a). pA polyadenylation signal, U6 human Pol III promoter, CMV cytomegalovirus immediate-early promoter, RHO human Rhodopsin promoter, N- or C-SpdCas9 N-or C-terminal part of split SpdCas9, N-or C-int N-or C-terminal part of split intein, dgRNA “dead” sgRNA with short spacer sequence (14–15 bp) harboring a MS2 stem-loop
Compared to SpdCas9-VPR, SpdCas9-SAM, or SpdCas9-SunTag modules, SpdCas9-VP64 results in substantially lower gene activation efficiencies [14] and is therefore less suitable for the treatment of diseases that require higher expression levels of target genes to achieve a therapeutic effect. Due to its considerably shorter coding sequence (3159 bp) SaCas9 offers a key advantage over SpCas9 (4104 bp) in terms of gene delivery using rAAVs, as these gold standard gene therapy vectors have a comparatively low genome packaging capacity (4.7 kb). Accordingly, in contrast to Sp(d)Cas9-VP64, Sa(d)Cas9-VP64 expression cassettes including all regulatory elements needed for effective expression (e.g., inverted terminal repeats, promoter, polyadenylation signal) typically do not exceed the payload of rAAVs when promoters shorter than 800 bp are used [24].
Given that the most potent VPR, SAM or SunTag-based CRISPRa variants exceed the packaging limit of rAAVs even if using SaCas9 or other alternatives with similar size such as Campylobacter jejuni CjCas9, [25], Geobacillus stearothermophilus GeoCas9 [26], or CasX [27] these Cas9 variants thus do not offer a noticeable advantage over SpCas9 for in vivo expression of these modules. Additionally, compared to SpCas9, SaCas9 has a more restrictive protospacer adjacent motif (PAM) sequence (5'-NNGRRT-3' for SaCas9 vs 5'-NGG-3' for SpCas9), which limits the number of suitable sgRNAs for activation of specific loci. Nevertheless, many studies have identified Cas variants with less restrictive PAM sequences (e.g. [28, 29]) and the field of identification and optimization of different therapeutically relevant Cas proteins is still very dynamic. Hence, it remains to be seen which of these Cas variants will win the race in the long run.
In addition, gene expression can also be regulated via recruitment of enzymes directly affecting DNA methylation or inducing histone modifications [30]. DNA methylation of specific regions results in gene silencing [31], whereas different histone modifications (e.g., acetylation, methylation, phosphorylation, formylation) lead to gene activation [32]. Several CRISPR-Cas modules exist that can be used for these purposes [33]. Among those, CRISPRa modules developed in the past contain Cas9 linked to demethylases such as dCas9 fused to the catalytic domain of ten-eleven translocation dioxygenase1 (TET1), [34], to catalytic histone acetyltransferase (HAT) core domain of the human E1A-associated protein p300 [35], or to histone methyltransferase PR domain zinc finger protein 9 (PRDM9) [36].
Advantages of CRISPRa for therapeutic applications
Compared to other existing strategies for the therapy of genetic or acquired diseases, CRISPRa-based approaches offer several important advantages: (i) They are independent of target gene size and can be used in multiplexing experiments to activate different genes simultaneously. (ii) They induce little to no off-target effects in vivo [18,19,20]. This is partly because potential off-target sequences, in addition to matching the target sequence, must also be part of a promoter or enhancer region. (iii) They can be used for simultaneous activation and knockdown of different genes in target cells. This requires the catalytically active Cas enzyme and the use of sgRNAs with spacer sequences of different lengths. For spacers less than 16 bp in length, Cas loses its catalytic activity, but the CRISPRa module is still capable of gene activation [6, 37]. Such an approach can be used, for example, to treat gain-of-function mutations that require simultaneous knockout of the mutant allele in addition to gene supplementation. (iv) In contrast to CRISPR/Cas modules which rely on the endonuclease activity, CRISPRa generally do not induce permanent changes in the genome. While permanent modification of the target locus is the desired effect in some diseases (e.g., to accomplish efficient gene knockdown), this can also become a problem if such modifications cause undesirable effects (e.g., large deletions) or occur at undesirable locations in the genome [38,39,40,41,42]. (v) In contrast to gene complementation strategies, where only the coding sequence is introduced, gene activation involves all natural processes of gene expression including mRNA splicing or the expression of long non-coding RNAs. (vi) In contrast to CRISPR/Cas-based strategies designed for excision or correction of the disease-causing mutation, CRISPRa approaches are mutation-independent and thus allow the treatment of a much larger number of patients.
Limitations of the CRISPRa strategy
Despite the important advantages listed above, CRISPRa approaches also have some notable limitations: (i) They require permanent expression of the gene activating module in target cells. Therefore, in contrast to approaches using the Cas endonuclease activity, delivery techniques tailored to short-term transgene expression (e.g., delivery of protein or mRNA by liposomes, virus-like particles, or nanoparticles) are not appropriate for CRISPRa. To date, studies in mouse models have not observed any negative effects on target cells despite prolonged expression of the CRISPRa components [15, 18,19,20]. Nevertheless, this circumstance needs to be further investigated before using CRISPRa modules in initial non-humane primates or clinical studies. A hit-and-run strategy that enables a long-term effect at the epigenome level has already been developed for the CRISPRi system [43], and it remains to be seen whether a similar approach can be achieved for CRISPRa in a therapeutic context. (ii) Similar to all other CRISPR-Cas systems, there is a risk of an immune reaction against the bacterial Cas protein in target cells [44]. (iii) They require appropriate delivery methods for efficient and long-lasting expression of CRISPRa cassettes in vivo. This is particularly difficult to be achieved for the large expression cassettes exceeding the packaging capacity of rAAV vectors [8]. (iv) When using vectors that are not integrated into the genome of the target cell (e.g., rAAVs), the application of CRISPRa is usually limited to non-diving cells, unless the therapeutic effect can be achieved by short-term expression of the target gene. Nevertheless, usage of AAV and CRISPR/Cas-based strategies is also limited due to the preexisting immunity to both components in the human population [45,46,47,48,49,50]. This can be partially overcome by developing new AAV vectors capsids [51] or using immune orthogonal AAVs and Cas proteins [52]. (v) The (over)expression of a gene in the target cell can also potentially have cell-damaging effects. In these cases, fine-tuning of the expression of affected genes may be required.
Strategies for in vivo delivery of CRISPRa modules for therapeutic purposes
Most therapeutic studies currently use rAAV vectors for stable (long-term) expression of transgenes in target cells [53]. Accordingly, despite many differences in the design of the individual vectors, all the studies reporting beneficial effects of the CRISPRa system in a therapeutically relevant experimental setup utilized AAV vectors so far.
The advantages of this method include inter alia high transduction efficiency, low immunogenicity, good tolerability, low risk of genomic integration, long-term expression, etc. [4, 54]. Moreover, just like CRISPR/Cas technology itself, the AAV vectors have already been approved by the FDA. Thus, the combination of both is well suited for translationally oriented studies. However, the limited DNA uptake capacity of these vectors requires alternative solutions for expression of large cassettes. Among the CRISPRa modules that have been successfully evaluated in vivo, Sa(d)Cas9-VP64 does not exceed the packaging capacity of rAAVs (Fig. 1A, B). However, due to its restrictive PAM sequence and low gene activation efficiency of VP64 alone, this module is less flexible and less broadly applicable. In comparison, expression of other more effective CRISPRa modules, such as SpdCas9-VPR or SpdCas9-MPH, requires co-application of two different AAV vectors, each encoding only a portion of the entire module (Fig. 1C, D). Such dual AAV vector approaches depend on high co-transduction efficiency of both vectors in target cells. High co-transduction rates (> 90%) were observed after local administration of rAAVs into some organs or tissues, such as the retina or brain [55,56,57,58]. From the efficacy perspective of dual AAV vectors, local administration (if possible) should therefore be preferred when dealing with diseases affecting only one or a few tissues or organs. However, local administration is also advantageous from a safety point of view, as fewer global immune reactions or side effects are to be expected compared to systemic application.
The transduction efficiency of dual rAAV vectors after systemic administration is understudied, but it is reasonable to assume that significantly higher AAV vector amounts are necessary to co-transduce a considerable percentage of target cells. However, higher rAAV dosage is associated with higher side effect risks, e.g., immune responses [59]. Therefore, further studies are necessary to investigate the principal suitability and the side effect profile of dual rAAV vectors after systemic administration. Some new strategies, such as co-packaging of multiple AAVs into extracellular vesicles, might help to decrease the dosage of dual AAVs needed for systemic application [60].
Despite their differences in the underlying molecular mechanisms, dual rAAV vectors expressing split genes follow the same principle: after the gene fragments are packaged into two different rAAV vectors and co-applied to the target tissue, they can be reconstituted at one of three levels: genomic, mRNA or protein level [61]. Historically, gene reconstitution at the genome level is the eldest dual rAAV vector strategy. There are several versions of this method and all of them depend on two processes called concatemerization and/or homologous recombination [62, 63]. In recent years another approach aiming at the reconstitution of the split rAAV genome at the protein level attracted increasing attention. In this context, each of the split coding sequences is fused to split inteins, which are small affinity tags capable of autocatalytic and scarless reconstitution of the split fragments at the protein level [64]. Split inteins have been shown to operate at high efficiency in vivo using rAAVs [15, 44, 65,66,67,68,69]. Nevertheless, the efficiency of this method strongly depends on the split site within the protein and relies on the presence of specific residues at the split and the neighboring positions. In addition, the reconstitution efficiency of proteins provided with split inteins also depends on protein folding. Optimal results are only achieved if both separate parts of a protein are folded independently of each other and if the exposed split inteins have good steric access to each other [64, 70]. Therefore, finding an efficient split position for split intein approaches requires an elaborative and time-consuming screening process for each gene. Finally, the split intein approach creates equimolar amounts of potentially immunogenic or pathogenic proteins (inteins), raising safety concerns for translational studies. Such concerns may be even more pronounced if split inteins are combined with Cas9-based approaches, as the co-expression of Cas9 and inteins could lead to additive or even synergistic effects with respect to the host immune responses. Conclusively, there is an unmet need for developing alternative approaches for reconstitution of split genes using dual rAAV vectors. Reconstitution of the split fragments at the mRNA level via a process called mRNA trans-splicing is a largely unexplored option for dual rAAV vectors and it remains to be shown whether this technology can overcome the drawbacks of currently used dual rAAV vector approaches.
Aside from their limited DNA packaging capacity, other factors exist that impede the widespread use of AAVs in clinical trials. One reason is the preexisting immunity to naturally occurring AAVs in the human population [45, 47, 50]. Although many AAV capsid variants have been reported that show a partial immune escape (e.g., [71,72,73,74,75,76,77,78,79,80,81,82]), their performance has to be further investigated in more advanced preclinical and clinical settings. Another bottleneck for the broad usage of AAVs in clinical trials are their high manufacturing costs, which drives up the cost of such therapies to unimaginable heights that hardly any healthcare system in the world can currently afford [83]. However, increasing competition and the number of production sites, as well as improved production technologies and processes, could lead to a significant reduction in these costs in the near future and thus make such therapies accessible to the broad mass of the population.
Apart from AAVs, to my knowledge, no other strategies have been used to deliver CRISPRa in vivo in a therapeutically relevant setting. This is most likely due to the fact that gene activation requires sustained expression of CRISPRa, which is unlikely to be achieved with non-viral systems, e.g., by delivery of the corresponding protein and sgRNAs using other techniques such as nanoparticles or virus-like particles [8]. In contrast, for other CRISPR-Cas-based systems designed to induce permanent changes in the genome sequence, transient expression is certainly an attractive solution to reduce the risk of adverse effects. Accordingly, several non-viral gene delivery strategies have been used for such purposes with varying success rates [84,85,86,87,88,89,90,91,92].
Other commonly used virus-based alternatives to AAVs for in vivo gene delivery are lentiviral or adenoviral vectors (LVs or Ad) [93]. An advantage of LVs and Ad is their much higher genome packaging capacity (10 kb and 36–37 kb, respectively) [94, 95]. This allows packaging of all relevant CRISPRa modules and corresponding sgRNA expression cassettes into a single vector. However, compared to AAVs, LVs and Ad also have some important drawbacks. These include the less preferable safety profile of these vectors due to their higher immunogenic activity. In addition, due to their size, Ad and LV are less able to cross biological barriers or diffuse away from the injection site (lateral spreading), which is required for the transduction of a larger area after a single local injection [93]. In addition, LVs carry a higher risk of integrating the transgenes into the host genome (insertional mutagenesis). Integrase-deficient LV variants have also been developed, but these lead to transient transgene expression and are therefore less suitable for CRISPRa approaches [96, 97]. This also applies to Ad variants which inherently show transient transgene expression [94].
Final remarks and future directions
CRISPRa is a modular and broadly applicable strategy for the therapy of genetic or acquired diseases. The very high therapeutic potential of CRISPRa approaches faces several challenges, mainly related to delivery, long-term expression, and immunity. AAV vectors remain the gold standard for CRISPRa delivery and long-term expression. Their limitations, such as limited cargo capacity or pre-existing immunity in the human population, may be overcome in the future by the development of novel AAV capsid variants with improved tissue tropism, higher packaging capacity, lower immunogenicity, and/or higher immune escape [83]. To minimize the potential negative effects of long-term CRISPRa expression, the use of controllable, i.e., demand-driven, expression systems, such as doxy- or tetracycline-dependent promoters or other inducible expression systems, would be beneficial [98]. However, this requires concomitant administration with the appropriate drugs, all of which may also cause side effects themselves and should therefore be balanced with the expected therapeutic benefit. Another approach that may facilitate gene delivery in the future is the development of new (shorter) CRISPRa variants that can be packaged into a single AAV vector without noticeable sacrifice in flexibility or efficacy. The new CRISPRa variants should meet similar requirements as the new AAV capsids in terms of their immunogenicity and immune escape. In addition to AAV-based approaches, the development of novel viral or non-viral delivery strategies could substantially advance the field and further exploit the potential and capabilities of CRISPRa in a therapeutic setting. However, given the advantages of AAVs, any new approaches will have to compete with these vectors in terms of efficacy and safety, and it remains to be seen whether this ambitious goal can be achieved in the foreseeable future.
Conclusively, although some barriers to gene delivery still need to be overcome and the immunogenicity better investigated, CRISPRa holds great translational potential and its implementation into first clinical trials could benefit millions of patients worldwide.
Availability of data and materials
Not applicable.
References
Jinek M, Chylinski K, Fonfara I, Hauer M, Doudna JA, Charpentier E (2012) A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Science 337(6096):816–821
Mali P, Yang L, Esvelt KM, Aach J, Guell M, DiCarlo JE et al (2013) RNA-guided human genome engineering via Cas9. Science 339(6121):823–826
Adli M (2018) The CRISPR tool kit for genome editing and beyond. Nat Commun 9(1):1911
Wang D, Zhang F, Gao G (2020) CRISPR-based therapeutic genome editing: strategies and in vivo delivery by AAV vectors. Cell 181(1):136–150
Qi LS, Larson MH, Gilbert LA, Doudna JA, Weissman JS, Arkin AP et al (2013) Repurposing CRISPR as an RNA-guided platform for sequence-specific control of gene expression. Cell 152(5):1173–1183
Dahlman JE, Abudayyeh OO, Joung J, Gootenberg JS, Zhang F, Konermann S (2015) Orthogonal gene knockout and activation with a catalytically active Cas9 nuclease. Nat Biotechnol 33(11):1159–1161
Gersbach CA, Perez-Pinera P (2014) Activating human genes with zinc finger proteins, transcription activator-like effectors and CRISPR/Cas9 for gene therapy and regenerative medicine. Expert Opin Ther Targets 18(8):835–839
van Haasteren J, Li J, Scheideler OJ, Murthy N, Schaffer DV (2020) The delivery challenge: fulfilling the promise of therapeutic genome editing. Nat Biotechnol 38(7):845–855
Maeder ML, Linder SJ, Cascio VM, Fu Y, Ho QH, Joung JK (2013) CRISPR RNA-guided activation of endogenous human genes. Nat Methods 10(10):977–979
Perez-Pinera P, Kocak DD, Vockley CM, Adler AF, Kabadi AM, Polstein LR et al (2013) RNA-guided gene activation by CRISPR-Cas9-based transcription factors. Nat Methods 10(10):973–976
Chavez A, Scheiman J, Vora S, Pruitt BW, Tuttle M, Eswar PRI et al (2015) Highly efficient Cas9-mediated transcriptional programming. Nat Methods 12(4):326–328
Tanenbaum ME, Gilbert LA, Qi LS, Weissman JS, Vale RD (2014) A protein-tagging system for signal amplification in gene expression and fluorescence imaging. Cell 159(3):635–646
Konermann S, Brigham MD, Trevino AE, Joung J, Abudayyeh OO, Barcena C et al (2015) Genome-scale transcriptional activation by an engineered CRISPR-Cas9 complex. Nature 517(7536):583–588
Chavez A, Tuttle M, Pruitt BW, Ewen-Campen B, Chari R, Ter-Ovanesyan D et al (2016) Comparison of Cas9 activators in multiple species. Nat Methods 13(7):563–567
Bohm S, Splith V, Riedmayr LM, Rotzer RD, Gasparoni G, Nordstrom KJV et al (2020) A gene therapy for inherited blindness using dCas9-VPR-mediated transcriptional activation. Sci Adv 6(34):eaba5614
Colasante G, Lignani G, Brusco S, Di Berardino C, Carpenter J, Giannelli S et al (2020) dCas9-based Scn1a gene activation restores inhibitory interneuron excitability and attenuates seizures in dravet syndrome mice. Mol Ther 28(1):235–253
Colasante G, Qiu Y, Massimino L, Di Berardino C, Cornford JH, Snowball A et al (2020) In vivo CRISPRa decreases seizures and rescues cognitive deficits in a rodent model of epilepsy. Brain 143(3):891–905
Kemaladewi DU, Bassi PS, Erwood S, Al-Basha D, Gawlik KI, Lindsay K et al (2019) A mutation-independent approach for muscular dystrophy via upregulation of a modifier gene. Nature 572(7767):125–130
Liao HK, Hatanaka F, Araoka T, Reddy P, Wu MZ, Sui Y et al (2017) In vivo target gene activation via CRISPR/Cas9-mediated trans-epigenetic modulation. Cell 171(7):1495–1507 (e15)
Matharu N, Rattanasopha S, Tamura S, Maliskova L, Wang Y, Bernard A et al (2019) CRISPR-mediated activation of a promoter or enhancer rescues obesity caused by haploinsufficiency. Science. https://doi.org/10.1126/science.aau0629
Wang G, Chow RD, Bai Z, Zhu L, Errami Y, Dai X et al (2019) Multiplexed activation of endogenous genes by CRISPRa elicits potent antitumor immunity. Nat Immunol 20(11):1494–1505
Yamagata T, Raveau M, Kobayashi K, Miyamoto H, Tatsukawa T, Ogiwara I et al (2020) CRISPR/dCas9-based Scn1a gene activation in inhibitory neurons ameliorates epileptic and behavioral phenotypes of Dravet syndrome model mice. Neurobiol Dis 141:104954
Moreno AM, Fu X, Zhu J, Katrekar D, Shih YV, Marlett J et al (2018) In situ gene therapy via AAV-CRISPR-Cas9-mediated targeted gene regulation. Mol Ther 26(7):1818–1827
Ran FA, Cong L, Yan WX, Scott DA, Gootenberg JS, Kriz AJ et al (2015) In vivo genome editing using Staphylococcus aureus Cas9. Nature 520(7546):186–191
Kim E, Koo T, Park SW, Kim D, Kim K, Cho HY et al (2017) In vivo genome editing with a small Cas9 orthologue derived from Campylobacter jejuni. Nat Commun 8:14500
Harrington LB, Paez-Espino D, Staahl BT, Chen JS, Ma E, Kyrpides NC et al (2017) A thermostable Cas9 with increased lifetime in human plasma. Nat Commun 8(1):1424
Liu JJ, Orlova N, Oakes BL, Ma E, Spinner HB, Baney KLM et al (2019) CasX enzymes comprise a distinct family of RNA-guided genome editors. Nature 566(7743):218–223
Hu JH, Miller SM, Geurts MH, Tang W, Chen L, Sun N et al (2018) Evolved Cas9 variants with broad PAM compatibility and high DNA specificity. Nature 556(7699):57–63
Schmidt MJ, Gupta A, Bednarski C, Gehrig-Giannini S, Richter F, Pitzler C et al (2021) Improved CRISPR genome editing using small highly active and specific engineered RNA-guided nucleases. Nat Commun 12(1):4219
Hashimshony T, Zhang J, Keshet I, Bustin M, Cedar H (2003) The role of DNA methylation in setting up chromatin structure during development. Nat Genet 34(2):187–192
Siegfried Z, Eden S, Mendelsohn M, Feng X, Tsuberi BZ, Cedar H (1999) DNA methylation represses transcription in vivo. Nat Genet 22(2):203–206
Tessarz P, Kouzarides T (2014) Histone core modifications regulating nucleosome structure and dynamics. Nat Rev Mol Cell Biol 15(11):703–708
Nakamura M, Gao Y, Dominguez AA, Qi LS (2021) CRISPR technologies for precise epigenome editing. Nat Cell Biol 23(1):11–22
Liu XS, Wu H, Ji X, Stelzer Y, Wu X, Czauderna S et al (2016) Editing DNA methylation in the mammalian genome. Cell 167(1):233–247 (e17)
Hilton IB, D’Ippolito AM, Vockley CM, Thakore PI, Crawford GE, Reddy TE et al (2015) Epigenome editing by a CRISPR-Cas9-based acetyltransferase activates genes from promoters and enhancers. Nat Biotechnol 33(5):510–517
Cano-Rodriguez D, Gjaltema RA, Jilderda LJ, Jellema P, Dokter-Fokkens J, Ruiters MH et al (2016) Writing of H3K4Me3 overcomes epigenetic silencing in a sustained but context-dependent manner. Nat Commun 7:12284
Kiani S, Chavez A, Tuttle M, Hall RN, Chari R, Ter-Ovanesyan D et al (2015) Cas9 gRNA engineering for genome editing, activation and repression. Nat Methods 12(11):1051–1054
Kang SH, Lee WJ, An JH, Lee JH, Kim YH, Kim H et al (2020) Prediction-based highly sensitive CRISPR off-target validation using target-specific DNA enrichment. Nat Commun 11(1):3596
Kim D, Bae S, Park J, Kim E, Kim S, Yu HR et al (2015) Digenome-seq: genome-wide profiling of CRISPR-Cas9 off-target effects in human cells. Nat Methods 12(3):237–243 (1 p following 43)
Pattanayak V, Lin S, Guilinger JP, Ma E, Doudna JA, Liu DR (2013) High-throughput profiling of off-target DNA cleavage reveals RNA-programmed Cas9 nuclease specificity. Nat Biotechnol 31(9):839–843
Tsai SQ, Nguyen NT, Malagon-Lopez J, Topkar VV, Aryee MJ, Joung JK (2017) CIRCLE-seq: a highly sensitive in vitro screen for genome-wide CRISPR-Cas9 nuclease off-targets. Nat Methods 14(6):607–614
Wienert B, Wyman SK, Richardson CD, Yeh CD, Akcakaya P, Porritt MJ et al (2019) Unbiased detection of CRISPR off-targets in vivo using DISCOVER-Seq. Science 364(6437):286–289
Amabile A, Migliara A, Capasso P, Biffi M, Cittaro D, Naldini L et al (2016) Inheritable silencing of endogenous genes by hit-and-run targeted epigenetic editing. Cell 167(1):219–232 (e14)
Chew WL, Tabebordbar M, Cheng JK, Mali P, Wu EY, Ng AH et al (2016) A multifunctional AAV-CRISPR-Cas9 and its host response. Nat Methods 13(10):868–874
Calcedo R, Vandenberghe LH, Gao G, Lin J, Wilson JM (2009) Worldwide epidemiology of neutralizing antibodies to adeno-associated viruses. J Infect Dis 199(3):381–390
Charlesworth CT, Deshpande PS, Dever DP, Camarena J, Lemgart VT, Cromer MK et al (2019) Identification of preexisting adaptive immunity to Cas9 proteins in humans. Nat Med 25(2):249–254
Ronzitti G, Gross DA, Mingozzi F (2020) Human immune responses to adeno-associated virus (AAV) vectors. Front Immunol 11:670
Simhadri VL, McGill J, McMahon S, Wang J, Jiang H, Sauna ZE (2018) Prevalence of pre-existing antibodies to CRISPR-associated nuclease Cas9 in the USA population. Mol Ther Methods Clin Dev 10:105–112
Wagner DL, Amini L, Wendering DJ, Burkhardt LM, Akyuz L, Reinke P et al (2019) High prevalence of Streptococcus pyogenes Cas9-reactive T cells within the adult human population. Nat Med 25(2):242–248
Wang L, Calcedo R, Wang H, Bell P, Grant R, Vandenberghe LH et al (2010) The pleiotropic effects of natural AAV infections on liver-directed gene transfer in macaques. Mol Ther 18(1):126–134
Barnes C, Scheideler O, Schaffer D (2019) Engineering the AAV capsid to evade immune responses. Curr Opin Biotechnol 60:99–103
Moreno AM, Palmer N, Aleman F, Chen G, Pla A, Jiang N et al (2019) Immune-orthogonal orthologues of AAV capsids and of Cas9 circumvent the immune response to the administration of gene therapy. Nat Biomed Eng 3(10):806–816
High KA, Roncarolo MG (2019) Gene therapy. N Engl J Med 381(5):455–464
Kuzmin DA, Shutova MV, Johnston NR, Smith OP, Fedorin VV, Kukushkin YS et al (2021) The clinical landscape for AAV gene therapies. Nat Rev Drug Discov 20(3):173–174
Becirovic E, Bohm S, Nguyen ON, Riedmayr LM, Hammelmann V, Schon C et al (2016) AAV vectors for FRET-based analysis of protein-protein interactions in photoreceptor outer segments. Front Neurosci 10:356
Becirovic E, Bohm S, Nguyen ON, Riedmayr LM, Koch MA, Schulze E et al (2016) In vivo analysis of disease-associated point mutations unveils profound differences in mRNA splicing of peripherin-2 in rod and cone photoreceptors. PLoS Genet 12(1):e1005811
Bohm S, Riedmayr LM, Nguyen ONP, Giessl A, Liebscher T, Butz ES et al (2017) Peripherin-2 and Rom-1 have opposing effects on rod outer segment targeting of retinitis pigmentosa-linked peripherin-2 mutants. Sci Rep 7(1):2321
Maturana CJ, Verpeut JL, Kooshkbaghi M, Engel EA (2021) Novel tool to quantify with single-cell resolution the number of incoming AAV genomes co-expressed in the mouse nervous system. Gene Ther. https://doi.org/10.1038/s41434-021-00272-8
Ertl HCJ (2021) T cell-mediated immune responses to AAV and AAV vectors. Front Immunol 12:666666
Gyorgy B, Maguire CA (2018) Extracellular vesicles: nature’s nanoparticles for improving gene transfer with adeno-associated virus vectors. Wiley Interdiscip Rev Nanomed Nanobiotechnol 10(3):e1488
Trapani I, Tornabene P, Auricchio A (2021) Large gene delivery to the retina with AAV vectors: are we there yet? Gene Ther 28(5):220–222
Li H, Yang Y, Hong W, Huang M, Wu M, Zhao X (2020) Applications of genome editing technology in the targeted therapy of human diseases: mechanisms, advances and prospects. Signal Transduct Target Ther 5(1):1
Patel A, Zhao J, Duan D, Lai Y (2019) Design of AAV vectors for delivery of large or multiple transgenes. Methods Mol Biol 1950:19–33
Muralidharan V, Muir TW (2006) Protein ligation: an enabling technology for the biophysical analysis of proteins. Nat Methods 3(6):429–438
Levy JM, Yeh WH, Pendse N, Davis JR, Hennessey E, Butcher R et al (2020) Cytosine and adenine base editing of the brain, liver, retina, heart and skeletal muscle of mice via adeno-associated viruses. Nat Biomed Eng 4(1):97–110
Lim CKW, Gapinske M, Brooks AK, Woods WS, Powell JE, Zeballos CM et al (2020) Treatment of a mouse model of ALS by in vivo base editing. Mol Ther 28(4):1177–1189
Schmelas C, Grimm D (2018) Split Cas9, not hairs—advancing the therapeutic index of CRISPR technology. Biotechnol J 13(9):e1700432
Tornabene P, Trapani I, Minopoli R, Centrulo M, Lupo M, de Simone S et al (2019) Intein-mediated protein trans-splicing expands adeno-associated virus transfer capacity in the retina. Sci Transl Med. https://doi.org/10.1126/scitranslmed.aav4523
Villiger L, Grisch-Chan HM, Lindsay H, Ringnalda F, Pogliano CB, Allegri G et al (2018) Treatment of a metabolic liver disease by in vivo genome base editing in adult mice. Nat Med 24(10):1519–1525
Shah NH, Muir TW (2014) Inteins: nature’s gift to protein chemists. Chem Sci 5(1):446–461
Bowles DE, McPhee SW, Li C, Gray SJ, Samulski JJ, Camp AS et al (2012) Phase 1 gene therapy for Duchenne muscular dystrophy using a translational optimized AAV vector. Mol Ther 20(2):443–455
Hsu HL, Brown A, Loveland AB, Lotun A, Xu M, Luo L et al (2020) Structural characterization of a novel human adeno-associated virus capsid with neurotropic properties. Nat Commun 11(1):3279
Huttner NA, Girod A, Perabo L, Edbauer D, Kleinschmidt JA, Buning H et al (2003) Genetic modifications of the adeno-associated virus type 2 capsid reduce the affinity and the neutralizing effects of human serum antibodies. Gene Ther 10(26):2139–2147
Katrekar D, Moreno AM, Chen G, Worlikar A, Mali P (2018) Oligonucleotide conjugated multi-functional adeno-associated viruses. Sci Rep 8(1):3589
Lee GK, Maheshri N, Kaspar B, Schaffer DV (2005) PEG conjugation moderately protects adeno-associated viral vectors against antibody neutralization. Biotechnol Bioeng 92(1):24–34
Li C, Wu S, Albright B, Hirsch M, Li W, Tseng YS et al (2016) Development of patient-specific AAV vectors after neutralizing antibody selection for enhanced muscle gene transfer. Mol Ther 24(1):53–65
Meliani A, Boisgerault F, Fitzpatrick Z, Marmier S, Leborgne C, Collaud F et al (2017) Enhanced liver gene transfer and evasion of preexisting humoral immunity with exosome-enveloped AAV vectors. Blood Adv 1(23):2019–2031
Mingozzi F, Anguela XM, Pavani G, Chen Y, Davidson RJ, Hui DJ et al (2013) Overcoming preexisting humoral immunity to AAV using capsid decoys. Sci Transl Med 5(194):194–192
Ojala DS, Sun S, Santiago-Ortiz JL, Shapiro MG, Romero PA, Schaffer DV (2018) In vivo selection of a computationally designed SCHEMA AAV library yields a novel variant for infection of adult neural stem cells in the SVZ. Mol Ther 26(1):304–319
Pavlou M, Schon C, Occelli LM, Rossi A, Meumann N, Boyd RF et al (2021) Novel AAV capsids for intravitreal gene therapy of photoreceptor disorders. EMBO Mol Med 13(4):e13392
Selot R, Arumugam S, Mary B, Cheemadan S, Jayandharan GR (2017) Optimized AAV rh.10 vectors that partially evade neutralizing antibodies during hepatic gene transfer. Front Pharmacol 8:441
Tse LV, Klinc KA, Madigan VJ, Castellanos Rivera RM, Wells LF, Havlik LP et al (2017) Structure-guided evolution of antigenically distinct adeno-associated virus variants for immune evasion. Proc Natl Acad Sci U S A 114(24):E4812–E4821
Wang D, Tai PWL, Gao G (2019) Adeno-associated virus vector as a platform for gene therapy delivery. Nat Rev Drug Discov 18(5):358–378
Banskota S, Raguram A, Suh S, Du SW, Davis JR, Choi EH et al (2022) Engineered virus-like particles for efficient in vivo delivery of therapeutic proteins. Cell. https://doi.org/10.1016/j.cell.2021.12.021
Campbell LA, Coke LM, Richie CT, Fortuno LV, Park AY, Harvey BK (2019) Gesicle-mediated delivery of CRISPR/Cas9 ribonucleoprotein complex for inactivating the HIV provirus. Mol Ther 27(1):151–163
Choi JG, Dang Y, Abraham S, Ma H, Zhang J, Guo H et al (2016) Lentivirus pre-packed with Cas9 protein for safer gene editing. Gene Ther 23(7):627–633
Gee P, Lung MSY, Okuzaki Y, Sasakawa N, Iguchi T, Makita Y et al (2020) Extracellular nanovesicles for packaging of CRISPR-Cas9 protein and sgRNA to induce therapeutic exon skipping. Nat Commun 11(1):1334
Hamilton JR, Tsuchida CA, Nguyen DN, Shy BR, McGarrigle ER, Sandoval Espinoza CR et al (2021) Targeted delivery of CRISPR-Cas9 and transgenes enables complex immune cell engineering. Cell Rep 35(9):109207
Indikova I, Indik S (2020) Highly efficient “hit-and-run” genome editing with unconcentrated lentivectors carrying Vpr.Prot.Cas9 protein produced from RRE-containing transcripts. Nucleic Acids Res 48(14):8178–8187
Lyu P, Lu Z, Cho SI, Yadav M, Yoo KW, Atala A et al (2021) Adenine base editor ribonucleoproteins delivered by lentivirus-like particles show high on-target base editing and undetectable RNA off-target activities. CRISPR J 4(1):69–81
Mangeot PE, Risson V, Fusil F, Marnef A, Laurent E, Blin J et al (2019) Genome editing in primary cells and in vivo using viral-derived nanoblades loaded with Cas9-sgRNA ribonucleoproteins. Nat Commun 10(1):45
Yao X, Lyu P, Yoo K, Yadav MK, Singh R, Atala A et al (2021) Engineered extracellular vesicles as versatile ribonucleoprotein delivery vehicles for efficient and safe CRISPR genome editing. J Extracell Vesicles 10(5):e12076
Bulcha JT, Wang Y, Ma H, Tai PWL, Gao G (2021) Viral vector platforms within the gene therapy landscape. Signal Transduct Target Ther 6(1):53
Lee CS, Bishop ES, Zhang R, Yu X, Farina EM, Yan S et al (2017) Adenovirus-mediated gene delivery: potential applications for gene and cell-based therapies in the new era of personalized medicine. Genes Dis 4(2):43–63
Milone MC, O’Doherty U (2018) Clinical use of lentiviral vectors. Leukemia 32(7):1529–1541
Chen F, Qi X, Zhang R, Wu ZY, Yan CE, Li J et al (2017) Episomal lentiviral vectors confer erythropoietin expression in dividing cells. Plasmid 90:15–19
Philippe S, Sarkis C, Barkats M, Mammeri H, Ladroue C, Petit C et al (2006) Lentiviral vectors with a defective integrase allow efficient and sustained transgene expression in vitro and in vivo. Proc Natl Acad Sci USA 103(47):17684–17689
Kallunki T, Barisic M, Jaattela M, Liu B (2019) How to choose the right inducible gene expression system for mammalian studies? Cells 8(8):796
Acknowledgements
I thank Stefanie Fenske, Lisa Riedmayr and Martin Biel for their thorough review of this manuscript.
Funding
Open Access funding enabled and organized by Projekt DEAL. This article was funded by the German Research Foundation, Individual Grant BE 4830/5-1 and Priority Program “Gene and cell based therapies to counteract neuroretinal degeneration” SPP2127, BE 4830/2-1.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The author declare has no conflict of interest.
Ethics approval and consent to participate
Not applicable.
Consent for publication
Not applicable.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Becirovic, E. Maybe you can turn me on: CRISPRa-based strategies for therapeutic applications. Cell. Mol. Life Sci. 79, 130 (2022). https://doi.org/10.1007/s00018-022-04175-8
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00018-022-04175-8