Abstract
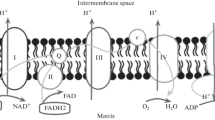
ATP synthases are unique rotatory molecular machines that supply biochemical reactions with adenosine triphosphate (ATP)—the universal “currency”, which cells use for synthesis of vital molecules and sustaining life. ATP synthases of F-type (FOF1) are found embedded in bacterial cellular membrane, in thylakoid membranes of chloroplasts, and in mitochondrial inner membranes in eukaryotes. The main functions of ATP synthases are control of the ATP synthesis and transmembrane potential. Although the key subunits of the enzyme remain highly conserved, subunit composition and structural organization of ATP synthases and their assemblies are significantly different. In addition, there are hypotheses that the enzyme might be involved in the formation of the mitochondrial permeability transition pore and play a role in regulation of the cell death processes. Dysfunctions of this enzyme lead to numerous severe disorders with high fatality levels. In our review, we focus on FOF1-structure-based approach towards development of new therapies by using FOF1 structural features inherited by the representatives of this enzyme family from different taxonomy groups. We analyzed and systematized the most relevant information about the structural organization of FOF1 to discuss how this approach might help in the development of new therapies targeting ATP synthases and design tools for cellular bioenergetics control.

The figure is reprinted from [4]

The figure was slightly modified from ref. [4]


The figure was taken from ref. [4]


The figure is from [77]




The figure was reprinted from ref. [4] with modifications

The figure was reprinted from ref. [4] with modifications

The figure was reprinted from ref. [4]

The figure was reprinted from ref. [4]

The figure was modified from [25]

Similar content being viewed by others
Availability of data and materials
Not applicable.
References
Walker JE (2013) The ATP synthase: the understood, the uncertain and the unknown. Biochem Soc Trans 41:1–16
Junge W, Nelson N (2015) ATP synthase. Annu Rev Biochem 84:631–657
Hahn A, Vonck J, Mills DJ, Meier T, Kühlbrandt W (2018) Structure, mechanism, and regulation of the chloroplast ATP synthase. Science 360:eaat4318
Vlasov AV (2021) New structural insights in chloroplast F1FO-ATP synthases—RWTH Publications. RWTH Aachen University. https://doi.org/10.18154/RWTH-2021-00849
Kühlbrandt W (2019) Structure and mechanisms of F-Type ATP synthases. Annu Rev Biochem 88:515–549
Morales-Rios E, Montgomery MG, Leslie AGW, Walker JE (2015) Structure of ATP synthase from Paracoccus denitrificans determined by X-ray crystallography at 4.0 Å resolution. Proc Natl Acad Sci USA 112:13231–13236
Sobti M et al (2020) Cryo-EM structures provide insight into how E. coli F1Fo ATP synthase accommodates symmetry mismatch. Nat Commun 11:1–10
Daum B, Nicastro D, Austin J, Richard MJ, Kühlbrandt W (2010) Arrangement of photosystem II and ATP synthase in chloroplast membranes of spinach and pea. Plant Cell 22:1299–1312
Seelert H, Dencher NA (2011) ATP synthase superassemblies in animals and plants: two or more are better. Biochim Biophys Acta Bioenerg 1807:1185–1197
Mühleip A et al (2021) ATP synthase hexamer assemblies shape cristae of Toxoplasma mitochondria. Nat Commun 12:1–13
Flygaard RK, Mühleip A, Tobiasson V, Amunts A (2020) Type III ATP synthase is a symmetry-deviated dimer that induces membrane curvature through tetramerization. Nat Commun 11:1–11
Nuskova H et al (2019) Biochemical thresholds for pathological presentation of ATP synthase deficiencies. Biochem Biophys Res Commun. https://doi.org/10.1016/j.bbrc.2019.11.033
Neupane P, Bhuju S, Thapa N, Bhattarai HK (2019) ATP synthase: structure, function and inhibition. Biomol Concepts 10:1–10
Gerle C (2016) On the structural possibility of pore-forming mitochondrial FoF1 ATP synthase. Biochim Biophys Acta Bioenerg 1857:1191–1196
Carraro M, Carrer A, Urbani A, Bernardi P (2020) Molecular nature and regulation of the mitochondrial permeability transition pore(s), drug target(s) in cardioprotection. J Mol Cell Cardiol 144:76–86
Walker JE, Carroll J, He J (2020) Reply to Bernardi: the mitochondrial permeability transition pore and the ATP synthase. Proc Natl Acad Sci 117:2745–2746
Petronilli V, Szabò I, Zoratti M (1989) The inner mitochondrial membrane contains ion-conducting channels similar to those found in bacteria. FEBS Lett 259:137–143
Szabo I, Zoratti M (1991) The giant channel of the inner mitochondrial membrane is inhibited by cyclosporin A. J Biol Chem 266:3376–3379
Szabó I, Zoratti M (1992) The mitochondrial megachannel is the permeability transition pore. J Bioenerg Biomembr 24:111–117
Szabo I, Bernardi P, Zoratti M (1992) Modulation of the mitochondrial megachannel by divalent cations and protons. J Biol Chem 267:2940–2946
Kinnally KW, Zorov DYA, Perini S (1991) Calcium modulation of mitochondrial inner membrane channel activity. Biochem Biophys Res Commun 176:1183–1188
Bernardi P et al (1992) Modulation of the mitochondrial permeability transition pore. Effect of protons and divalent cations. J Biol Chem 267:2934–2939
Vlasov AV et al (2019) Unusual features of the c-ring of F1FO ATP synthases. Sci Rep 9:18547
Preiss L et al (2015) Structure of the mycobacterial ATP synthase Fo rotor ring in complex with the anti-TB drug bedaquiline. Sci Adv 1:e1500106–e1500106
Guo H et al (2021) Structure of mycobacterial ATP synthase bound to the tuberculosis drug bedaquiline. Nature 589:143–147
Grinkova YV, Denisov IG, Sligar SG (2010) Engineering extended membrane scaffold proteins for self-assembly of soluble nanoscale lipid bilayers. Protein Eng Des Sel 23:843–848
Muñoz-Gómez SA, Wideman JG, Roger AJ, Slamovits CH (2017) The origin of mitochondrial cristae from alphaproteobacteria. Mol Biol Evol 34:943–956
Eydt K, Davies KM, Behrendt C, Wittig I, Reichert AS (2017) Cristae architecture is determined by an interplay of the MICOS complex and the F1Fo ATP synthase via Mic27 and Mic10. Microb Cell 4:259–272
Davies KM, Anselmi C, Wittig I, Faraldo-Gómez JD, Kühlbrandt W (2012) Structure of the yeast F1Fo-ATP synthase dimer and its role in shaping the mitochondrial cristae. Proc Natl Acad Sci USA 109:13602–13607
Schoch CL et al (2020) NCBI taxonomy: a comprehensive update on curation, resources and tools. Database 2020: baaa062. PubMed: 32761142 PMC: PMC7408187
Sobti M et al (2016) Cryo-EM structures of the autoinhibited E. coli ATP synthase in three rotational states. Elife 5:e21598
Guo H, Suzuki T, Rubinstein JL (2019) Structure of a bacterial ATP synthase. Elife 8:e43128
Guo H, Bueler SA, Rubinstein JL (2017) Atomic model for the dimeric FOregion of mitochondrial ATP synthase. Science 358:936–940
Gu J et al (2019) Cryo-EM structure of the mammalian ATP synthase tetramer bound with inhibitory protein IF1. Science 364:1068–1075
Rexroth S et al (2004) Dimeric H+-ATP synthase in the chloroplast of Chlamydomonas reinhardtii. Biochim Biophys Acta Bioenerg 1658:202–211
Schwaßmann HJ, Rexroth S, Seelert H, Dencher NA (2007) Metabolism controls dimerization of the chloroplast FoF1 ATP synthase in Chlamydomonas reinhardtii. FEBS Lett 581:1391–1396
Murphy BJ et al (2019) Rotary substates of mitochondrial ATP synthase reveal the basis of flexible F1–Fo coupling. Science 364:eaaw9128
Blum TB, Hahn A, Meier T, Davies KM, Kühlbrandt W (2019) Dimers of mitochondrial ATP synthase induce membrane curvature and self-assemble into rows. Proc Natl Acad Sci USA 116:4250–4255
Mühleip AW, Dewar CE, Schnaufer A, Kühlbrandt W, Davies KM (2017) In situ structure of trypanosomal ATP synthase dimer reveals a unique arrangement of catalytic subunits. Proc Natl Acad Sci 114:992–997
Mühleip A, McComas SE, Amunts A (2019) Structure of a mitochondrial ATP synthase with bound native cardiolipin. Elife 8:e51179
Vlasov AV, Ryzhykau YL, Gordeliy VI, Kuklin AI (2017) Spinach ATP-synthases form dimers in nanodiscs. Small-angle X-ray and neutron scattering investigations. FEBS J 284:87
Yanyushin MF (1993) Subunit structure of ATP synthase from chloroflexus aurantiacus. FEBS Lett 335:85–88
Yanyushin MF (1997) Determination of subunit composition of the F 1 and F 0 moieties of ATP synthase from Chloroflexus aurantiacus. Biochem 62:285–288
Abrahams JP, Leslie AGW, Lutter R, Walker JE (1994) Structure at 28 Â resolution of F1-ATPase from bovine heart mitochondria. Nature 370:621–628
Stock D, Leslie AGW, Walker JE (1999) Molecular architecture of the rotary motor in ATP synthase. Science 286:1700–1705
Dautant A, Velours J, Giraud MF (2010) Crystal structure of the Mg·ADP-inhibited state of the yeast F 1c10-ATP synthase. J Biol Chem 285:29502–29510
Watt IN, Montgomery MG, Runswick MJ, Leslie AGW, Walker JE (2010) Bioenergetic cost of making an adenosine triphosphate molecule in animal mitochondria. Proc Natl Acad Sci USA 107:16823–16827
Giraud MF et al (2012) Rotor architecture in the yeast and bovine F 1-c-ring complexes of F-ATP synthase. J Struct Biol 177:490–497
Morales-Rios E et al (2015) Purification, characterization and crystallization of the F-ATPase from Paracoccus denitrificans. Open Biol 5:150119
Hahn A et al (2016) Structure of a complete ATP synthase dimer reveals the molecular basis of inner mitochondrial membrane morphology. Mol Cell 63:445–456
Seelert H et al (2000) Proton-powered turbine of a plant motor. Nature 405:418–419
Zhou A et al (2015) Structure and conformational states of the bovine mitochondrial ATP synthase by cryo-EM. Elife 4:e10180
Allegretti M et al (2015) Horizontal membrane-intrinsic α-helices in the stator a-subunit of an F-type ATP synthase. Nature 521:237–240
Vinothkumar KR, Montgomery MG, Liu S, Walker JE (2016) Structure of the mitochondrial ATP synthase from Pichia angusta determined by electron cryo-microscopy. Proc Natl Acad Sci USA 113:12709–12714
Srivastava AP et al (2018) High-resolution cryo-EM analysis of the yeast ATP synthase in a lipid membrane. Science 360:6389
Mellwig C, Böttcher B (2003) A unique resting position of the ATP-synthase from Chloroplasts*. J Biol Chem 278:18544–18549
Guo H, Bueler SA, Rubinstein JL (2017) Atomic model for the dimeric FO region of mitochondrial ATP synthase. Science 358:936–940
Allen RD, Schroeder CC, Fok AK (1989) An investigation of mitochondrial inner membranes by rapid-freeze deep-etch techniques. J Cell Biol 108:2233–2240
Nicastro D, Frangakis AS, Typke D, Baumeister W (2000) Cryo-electron tomography of neurospora mitochondria. J Struct Biol 129:48–56
Giraud MF et al (2002) Is there a relationship between the supramolecular organization of the mitochondrial ATP synthase and the formation of cristae? Biochim Biophys Acta Bioenerg 1555:174–180
Benvenuti M, Mangani S (2007) Crystallization of soluble proteins in vapor diffusion for X-ray crystallography. Nat Protoc 2:1633–1651
Balakrishna AM, Seelert H, Marx S-H, Dencher NA, Grüber G (2014) Crystallographic structure of the turbine C-ring from spinach chloroplast F-ATP synthase. Biosci Rep 34:e00102
Caffrey M, Cherezov V (2009) Crystallizing membrane proteins using lipidic mesophases. Nat Protoc 4:706–731
Li D, Shah STA, Caffrey M (2013) Host lipid and temperature as important screening variables for crystallizing integral membrane proteins in lipidic mesophases. Trials with diacylglycerol kinase. Cryst Growth Des 13:2846–2857
Ishchenko A et al (2017) Chemically stable lipids for membrane protein crystallization. Cryst Growth Des 17:3502–3511
Zabara A et al (2018) Design of ultra-swollen lipidic mesophases for the crystallization of membrane proteins with large extracellular domains. Nat Commun 9:1–9
Johnson DE et al (2012) High-throughput characterization of intrinsic disorder in proteins from the protein structure initiative. J Struct Biol 180:201–215
Sedzik J, Kirschner DA (1992) Is myelin basic protein crystallizable? Neurochem Res 17:157–166
Harauz G et al (2004) Myelin basic protein-diverse conformational states of an intrinsically unstructured protein and its roles in myelin assembly and multiple sclerosis. Micron 35:503–542
Le Gall T, Romero PR, Cortese MS, Uversky VN, Dunker AK (2007) Intrinsic disorder in the protein data bank. J Biomol Struct Dyn 24:325–341
DeForte S, Uversky VN (2016) Resolving the ambiguity: making sense of intrinsic disorder when PDB structures disagree. Protein Sci 25:676–688
Romero P et al (2000) Sequence complexity of disordered protein. Proteins Struct Funct Bioinform 42:38–48
Peng K, Radivojac P, Vucetic S, Dunker AK, Obradovic Z (2006) Length-dependent prediction of protein intrinsic disorder. BMC Bioinform 7:208–212
Peng K et al (2005) Optimizing long intrinsic disorder predictors with protein evolutionary information. J Bioinform Comput Biol 3:35–60
Xue B, Dunbrack RL, Williams RW, Dunker AK, Uversky VN (2010) PONDR-FIT: a meta-predictor of intrinsically disordered amino acids. Biochim Biophys Acta 1804:996–1010
Mészáros B, Erdos G, Dosztányi Z (2018) IUPred2A: context-dependent prediction of protein disorder as a function of redox state and protein binding. Nucleic Acids Res 46:W329–W337
Mendoza-Hoffmann F, Zarco-Zavala M, Ortega R, García-Trejo JJ (2018) Control of rotation of the F1FO-ATP synthase nanomotor by an inhibitory α-helix from unfolded ε or intrinsically disordered ζ and IF1 proteins. J Bioenerg Biomembr 50:403–424
Yang J-H, Williams D, Kandiah E, Fromme P, Chiu P-L (2020) Structural basis of redox modulation on chloroplast ATP synthase. Commun Biol 3:1–12
Fischer S et al (1994) ATP synthesis catalyzed by the ATP synthase of Escherichia coli reconstituted into liposomes. Eur J Biochem 225:167–172
Poetsch A et al (2003) Characterisation of subunit III and its oligomer from spinach chloroplast ATP synthase. Biochim Biophys Acta Biomembr 1618:59–66
Suhai T, Dencher NA, Poetsch A, Seelert H (2008) Remarkable stability of the proton translocating F1FO-ATP synthase from the thermophilic cyanobacterium Thermosynechococcus elongatus BP-1. Biochim Biophys Acta Biomembr 1778:1131–1140
Suhai T, Heidrich NG, Dencher NA, Seelert H (2009) Highly sensitive detection of ATPase activity in native gels. Electrophoresis 30:3622–3625
Schmidt G, Gräber P (1985) The rate of ATP synthesis by reconstituted CF0F1 liposomes. Biochim Biophys Acta Bioenerg 808:46–51
Förster K et al (2010) Proton transport coupled ATP synthesis by the purified yeast H+-ATP synthase in proteoliposomes. Biochim Biophys Acta Bioenerg 1797:1828–1837
Turina P, Samoray D, Gräber P (2003) H+/ATP ratio of proton transport-coupled ATP synthesis and hydrolysis catalysed by CF0F1-liposomes. EMBO J. https://doi.org/10.1093/emboj/cdg073
Groth G, Walker JE (1996) ATP synthase from bovine heart mitochondria: reconstitution into unilamellar phospholipid vesicles of the pure enzyme in a functional state. Biochem J 318:351–357
Varco-Merth B, Fromme R, Wang M, Fromme P (2008) Crystallization of the c14-rotor of the chloroplast ATP synthase reveals that it contains pigments. Biochim Biophys Acta Bioenerg 1777:605–612
Grotjohann I, Gräber P (2002) The H+-ATPase from chloroplasts: effect of different reconstitution procedures on ATP synthesis activity and on phosphate dependence of ATP synthesis. Biochim Biophys Acta Bioenerg 1556:208–216
Pogoryelov D et al (2012) Engineering rotor ring stoichiometries in the ATP synthase. Proc Natl Acad Sci USA 109:E1599–E1608
Luz AL, Lagido C, Hirschey MD, Meyer JN (2016) In vivo determination of mitochondrial function using luciferase-expressing caenorhabditis elegans: contribution of oxidative phosphorylation, glycolysis, and fatty acid oxidation to toxicant-induced dysfunction. Curr Protoc Toxicol 69:2581–25822
Boerries M et al (2007) Ca2+-dependent interaction of S100A1 with F1-ATPase leads to an increased ATP content in cardiomyocytes. Mol Cell Biol 27:4365–4373
Drew B, Leeuwenburgh C (2003) Method for measuring ATP production in isolated mitochondria: ATP production in brain and liver mitochondria of Fischer-344 rats with age and caloric restriction. Am J Physiol Regul Integr Comp Physiol 285:1259–1267
Vinkler C, Korenstein R (1982) Characterization of external electric field-driven ATP synthesis in chloroplasts. Proc Natl Acad Sci 79:3183–3187
Sun T et al (2000) Ralstonia solanacearum elicitor RipX induces defense reaction by suppressing the mitochondrial atpA gene in host plant. Int J Mol Sci 2020:21
Meighen EA (1991) Molecular biology of bacterial bioluminescence. Microbiol Rev 55:123–142
Nijvipakul S et al (2008) LuxG is a functioning flavin reductase for bacterial luminescence. J Bacteriol 190:1531–1538
Kalyabina VP, Esimbekova EN, Torgashina IG, Kopylova KV, Kratasyuk VA (2019) Principles for construction of bioluminescent enzyme biotests for analysis of complex media. Dokl Biochem Biophys 485:107–110
Zavilgelsky GB, Kotova VY, Mazhul’ MM, Manukhov IV (2002) Role of Hsp70 (DnaK–DnaJ–GrpE) and Hsp100 (ClpA and ClpB) chaperones in refolding and increased thermal stability of bacterial luciferases in Escherichia coli cells. Biochem 67:986–992
Bernardi P (2020) Mechanisms for Ca2+-dependent permeability transition in mitochondria. Proc Natl Acad Sci USA 117:2743–2744
Neginskaya MA et al (2019) ATP synthase c-subunit-deficient mitochondria have a small cyclosporine a-sensitive channel, but lack the permeability transition pore. Cell Rep 26:11-17.e2
Giorgio V et al (2013) Dimers of mitochondrial ATP synthase form the permeability transition pore. Proc Natl Acad Sci 110:5887–5892
Kokoszka JE et al (2004) The ADP/ATP translocator is not essential for the mitochondrial permeability transition pore. Nature 427:461–465
He J et al (2017) Persistence of the mitochondrial permeability transition in the absence of subunit c of human ATP synthase. Proc Natl Acad Sci 114:3409–3414
Alavian KN et al (2014) An uncoupling channel within the c-subunit ring of the F1FO ATP synthase is the mitochondrial permeability transition pore. Proc Natl Acad Sci 111:10580–10585
Bernardi P, Di Lisa F (2015) The mitochondrial permeability transition pore: molecular nature and role as a target in cardioprotection. J Mol Cell Cardiol 78:100–106
Zhou W, Marinelli F, Nief C, Faraldo-Gómez JD (2017) Atomistic simulations indicate the c-subunit ring of the F1Fo ATP synthase is not the mitochondrial permeability transition pore. Elife 6:e23781
Spikes TE, Montgomery MG, Walker JE (2020) Structure of the dimeric ATP synthase from bovine mitochondria. Proc Natl Acad Sci 117:23519–23526
Carraro M et al (2014) Channel formation by yeast F-ATP synthase and the role of dimerization in the mitochondrial permeability transition. J Biol Chem 289:15980–15985
Carraro M et al (2018) High-conductance channel formation in yeast mitochondria is mediated by F-ATP synthase e and g subunits. Cell Physiol Biochem 50:1840–1855
Urbani A et al (2019) Purified F-ATP synthase forms a Ca2+-dependent high-conductance channel matching the mitochondrial permeability transition pore. Nat Commun 10:1–11
Bonora M et al (2017) Mitochondrial permeability transition involves dissociation of F1FO ATP synthase dimers and C-ring conformation. EMBO Rep 18:1077–1089
Mnatsakanyan N et al (2019) A mitochondrial megachannel resides in monomeric F1FO ATP synthase. Nat Commun 10:1–11
Pinke G, Zhou L, Sazanov LA (2020) Cryo-EM structure of the entire mammalian F-type ATP synthase. Nat Struct Mol Biol 27:1077–1085
Amodeo GF et al (2021) C subunit of the ATP synthase is an amyloidogenic calcium dependent channel-forming peptide with possible implications in mitochondrial permeability transition. Sci Rep 11:1–10
Matthies D et al (2014) High-resolution structure and mechanism of an F/V-hybrid rotor ring in a Na+-coupled ATP synthase. Nat Commun 5:5286
Meier T, Matthey U, Henzen F, Dimroth P, Müller DJ (2001) The central plug in the reconstituted undecameric c cylinder of a bacterial ATP synthase consists of phospholipids. FEBS Lett 505:353–356
Seelert H, Dencher NA, Müller DJ (2003) Fourteen protomers compose the oligomer III of the proton-rotor in spinach chloroplast ATP synthase. J Mol Biol 333:337–344
Novitskaia O, Buslaev P, Gushchin I (2019) Assembly of spinach chloroplast ATP synthase rotor ring protein–lipid complex. Front Mol Biosci 6:135
Meier T et al (2009) Complete ion-coordination structure in the rotor ring of Na+-dependent F-ATP synthases. J Mol Biol 391:498–507
Fromme P, Gräber P, Boekema EJ (1987) Isolation and characterization of a supramolecular complex of subunit III of the ATP-synthase from chloroplasts. Zeitschrift fur Naturforsch Sect C J Biosci 42:1239–1245
Vollmar M, Schlieper D, Winn M, Büchner C, Groth G (2009) Structure of the c14 rotor ring of the proton translocating chloroplast ATP synthase. J Biol Chem 284:18228–18235
Symersky J et al (2012) Structure of the c10 ring of the yeast mitochondrial ATP synthase in the open conformation. Nat Struct Mol Biol 19:485–491
Preiss L et al (2014) The c-ring ion binding site of the ATP synthase from Bacillus pseudofirmus OF4 is adapted to alkaliphilic lifestyle. Mol Microbiol 92:973–984
Pogoryelov D et al (2010) Microscopic rotary mechanism of ion translocation in the Fo complex of ATP synthases. Nat Chem Biol 6:891–899
Spikes TE, Montgomery MG, Walker JE (2021) Interface mobility between monomers in dimeric bovine ATP synthase participates in the ultrastructure of inner mitochondrial membranes. Proc Natl Acad Sci 118
Tacconelli E et al (2018) Discovery, research, and development of new antibiotics: the WHO priority list of antibiotic-resistant bacteria and tuberculosis. Lancet Infect Dis 18:318–327
Gao W, Howden BP, Stinear TP (2018) Evolution of virulence in Enterococcus faecium, a hospital-adapted opportunistic pathogen. Curr Opin Microbiol 41:76–82
Li Y, Sun F, Zhang W (2019) Bedaquiline and delamanid in the treatment of multidrug-resistant tuberculosis: promising but challenging. Drug Dev Res 80:98–105
Kevric I, Cappel MA, Keeling JH (2015) New world and old world leishmania infections: a practical review. Dermatol Clin 33:579–593
Migchelsen SJ, Büscher P, Hoepelman AIM, Schallig HDFH, Adams ER (2011) Human African trypanosomiasis: a review of non-endemic cases in the past 20 years. Int J Infect Dis 15:e517–e524
Young KM et al (2019) Zoonotic Babesia: a scoping review of the global evidence. PLoS One 14:e0226781
Saadatnia G, Golkar M (2012) A review on human toxoplasmosis. Scand J Infect Dis. https://doi.org/10.3109/00365548.2012.693197
Schuster FL, Ramirez-Avila L (2008) Current world status of Balantidium coli. Clin Microbiol Rev 21:626–638
Fayer R, Ungar BLP (1986) Cryptosporidium spp. and Cryptosporidiosis. Microbiol Rev 50:458–483
Tuteja R (2007) Malaria—an overview. FEBS J 274:4670–4679
Clark C, Espinosa Cantellano M, Bhattacharya A (2000) Entamoeba Histolytica: an overview of the biology of the organism. Amebiasis. https://doi.org/10.1142/9781848160583_0001
El-Dib NA (2017) Entamoeba histolytica: an overview. Curr Trop Med Rep 4:11–20
Visvesvara GS (1983) Giardiasis: an overview. IMJ Ill Med J 164:34–39
Docampo R, Ryan CM, de Miguel N, Johnson PJ (2011) Trichomonas vaginalis: current understanding of host-parasite interactions. Essays Biochem 51:161–175
Jumper J et al (2021) Highly accurate protein structure prediction with AlphaFold. Nature 596:583–589
Tunyasuvunakool K et al (2021) Highly accurate protein structure prediction for the human proteome. Nature 596:590–596
Baek M et al (2021) Accurate prediction of protein structures and interactions using a three-track neural network. Science 373:871–876
Mullard A (2021) What does AlphaFold mean for drug discovery? Nat Rev Drug Discov 20:725–727
Cui Y et al (2020) Recent progress in the use of mitochondrial membrane permeability transition pore in mitochondrial dysfunction-related disease therapies. Mol Cell Biochem 476:493–506
Acknowledgements
AVV greatly acknowledges the Council for Grants of the President of the Russian Federation for state support of young Russian scientists and for state support of leading scientific schools of the Russian Federation (Scholarship of the President of the Russian Federation, Order of the Ministry of Education and Science of the Russian Federation of January 26, 2021 No. 54 on the appointment of a scholarship for 2021-2023). SDO acknowledges funding from the Foundation of Promoting Innovations for financial support in the framework of the program UMNIK. VIG acknowledges funding from Frankfurt: Cluster of Excellence Frankfurt Macromolecular Complexes (to E.B.) by the Max Planck Society (to E.B.) and by the Commissariat à l’Energie Atomique et aux Energies Alternatives (Institut de Biologie Structurale) – Helmholtz-Gemeinschaft Deutscher Forschungszentren (Forschungszentrum Jülich) Special Terms and Conditions 5.1 specific agreement.
Funding
The work is supported by RFBR 19-29-12022. The work is supported by the Ministry of Science and Higher Education of the Russian Federation (075-00337-20-03/FSMG-2020-0003; 075–00958-21–05, project # 730000F.99.1.BV10AA00006).
Author information
Authors and Affiliations
Contributions
AVV designed, conceived and wrote the manuscript. SDO strongly contributed to the section “ATP synthases as potential therapeutic targets”, edited the text of the manuscript. NAB strongly contributed to the section “Mitochondrial permeability transition pore (mPTP)”. VNU strongly contributed to the section “Intrinsically disordered regions in ATP synthases”, contributed to all sections and edited the text of the manuscript. VIB contributed to the section “Outlook”, edited the text of the manuscript. MFY contributed to the section “Similarity and diversity of ATP synthases”. IVM contributed to the section “Validation of ATP synthase functionality”, edited the text of the manuscript. AVR organized funding acquisition, edited the text of the manuscript. ADV contributed to the section “Small-molecule cofactors of the c-ring”, edited the text of the manuscript. NSI contributed to the section “Intrinsically disordered regions in ATP synthases”, edited the text of the manuscript. AIK contributed to the section “High-resolution structural studies”. NAD contributed to all sections and edited the text of the manuscript. VIG supervised the project, strongly contributed to all sections and edited the text of the manuscript. All authors have read and agreed to the published version of the manuscript.
Corresponding author
Ethics declarations
Conflict of interests
The authors declare that they have no conflict of interest.
Ethics approval and consent to participate
Not applicable.
Consent for publication
Not applicable.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Vlasov, A.V., Osipov, S.D., Bondarev, N.A. et al. ATP synthase FOF1 structure, function, and structure-based drug design. Cell. Mol. Life Sci. 79, 179 (2022). https://doi.org/10.1007/s00018-022-04153-0
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00018-022-04153-0