Abstract
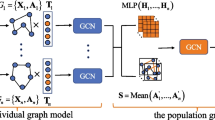
Analyzing connections between brain regions of interest (ROI) is vital to detect neurological disorders such as autism or schizophrenia. Recent advancements employ graph neural networks (GNNs) to utilize graph structures in brains, improving detection performances. Current methods use correlation measures between ROI’s blood-oxygen-level-dependent (BOLD) signals to generate the graph structure. Other methods use the training samples to learn the optimal graph structure through end-to-end learning. However, implementing those methods independently leads to some issues with noisy data for the correlation graphs and overfitting problems for the optimal graph. In this work, we proposed Bargrain (balanced graph structure for brains), which models two graph structures: filtered correlation matrix and optimal sample graph using graph convolution networks (GCNs). This approach aims to get advantages from both graphs and address the limitations of only relying on a single type of structure. Based on our extensive experiment, Bargrain outperforms state-of-the-art methods in classification tasks on brain disease datasets, as measured by average F1 scores.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
Notes
- 1.
The implementation of Bargrain: https://github.com/falihgoz/Bargrain.
- 2.
- 3.
http://fcon_1000.projects.nitrc.org/indi/ACPI/html/.
References
Chen, W., Wang, Y., Du, C., Jia, Z., Liu, F., Chen, R.: Balanced spatial-temporal graph structure learning for multivariate time series forecasting: a trade-off between efficiency and flexibility. In: ACML. PMLR (2023)
Cui, H., et al.: Braingb: a benchmark for brain network analysis with graph neural networks. IEEE Trans. Med. Imaging 42(2), 493–506 (2022)
Dadi, K., et al.: Benchmarking functional connectome-based predictive models for resting-state FMRI. Neuroimage 192, 115–134 (2019)
Di Martino, A., et al.: The autism brain imaging data exchange: towards a large-scale evaluation of the intrinsic brain architecture in autism. Mol. Psychiatry 19(6), 659–667 (2014)
ElGazzar, A., Thomas, R., Van Wingen, G.: Benchmarking graph neural networks for FMRI analysis. arXiv preprint arXiv:2211.08927 (2022)
Febrinanto, F.G.: Efficient graph learning for anomaly detection systems. In: Proceedings of the Sixteenth ACM International Conference on Web Search and Data Mining, pp. 1222–1223 (2023)
Febrinanto, F.G., Xia, F., Moore, K., Thapa, C., Aggarwal, C.: Graph lifelong learning: a survey. IEEE Comput. Intell. Mag. 18(1), 32–51 (2023)
Hutchison, R.M., et al.: Dynamic functional connectivity: promise, issues, and interpretations. Neuroimage 80, 360–378 (2013)
Jang, E., Gu, S., Poole, B.: Categorical reparameterization with gumbel-softmax. In: International Conference on Learning Representations (ICLR) (2017)
Kan, X., Cui, H., Lukemire, J., Guo, Y., Yang, C.: FBNetGen: task-aware GNN-based FMRI analysis via functional brain network generation. In: MIDL. PMLR (2022)
Kan, X., Dai, W., Cui, H., Zhang, Z., Guo, Y., Yang, C.: Brain network transformer. In: Advances in Neural Information Processing Systems (NeurIPS) (2022)
Kawahara, J., et al.: BrainNetCNN: convolutional neural networks for brain networks; towards predicting neurodevelopment. Neuroimage 146, 1038–1049 (2017)
Kazi, A., Cosmo, L., Ahmadi, S.A., Navab, N., Bronstein, M.M.: Differentiable graph module (DGM) for graph convolutional networks. IEEE Trans. Pattern Anal. Mach. Intell. 45(2), 1606–1617 (2022)
Li, X., et al.: BrainGNN: interpretable brain graph neural network for FMRI analysis. Med. Image Anal. 74, 102233 (2021)
Ren, J., Xia, F., Lee, I., Noori Hoshyar, A., Aggarwal, C.: Graph learning for anomaly analytics: algorithms, applications, and challenges. ACM Trans. Intell. Syst. Technol. 14(2), 1–29 (2023)
Shang, C., Chen, J., Bi, J.: Discrete graph structure learning for forecasting multiple time series. In: International Conference on Learning Representations (ICLR) (2020)
Welling, M., Kipf, T.N.: Semi-supervised classification with graph convolutional networks. In: International Conference on Learning Representations (ICLR) (2017)
Xia, F., et al.: CenGCN: centralized convolutional networks with vertex imbalance for scale-free graphs. IEEE Trans. Knowl. Data Eng. 35(5), 4555–4569 (2022)
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2023 The Author(s), under exclusive license to Springer Nature Singapore Pte Ltd.
About this paper
Cite this paper
Febrinanto, F.G., Liu, M., Xia, F. (2023). Balanced Graph Structure Information for Brain Disease Detection. In: Wu, S., Yang, W., Amin, M.B., Kang, BH., Xu, G. (eds) Knowledge Management and Acquisition for Intelligent Systems. PKAW 2023. Lecture Notes in Computer Science(), vol 14317. Springer, Singapore. https://doi.org/10.1007/978-981-99-7855-7_11
Download citation
DOI: https://doi.org/10.1007/978-981-99-7855-7_11
Published:
Publisher Name: Springer, Singapore
Print ISBN: 978-981-99-7854-0
Online ISBN: 978-981-99-7855-7
eBook Packages: Computer ScienceComputer Science (R0)