Abstract
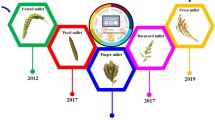
Nutritionally important cereals of small millets are used as food crop for poor people around the world. They are mainly cultivated in arid and semi-arid regions of Asian and African countries. Despite the nutritional benefits, small millets remain an underutilized crop and have received very little attention from researchers as well as farmers around the world. In the last few decades, very little research efforts have been made to study the features of small millets. More concentrated research efforts are needed to characterize germplasm resources, developing mapping population and identify quantitative trait loci (QTL) or genes which may help to improve the growth and production of small millets under biotic and abiotic stresses. The annotated genome sequences are currently available for foxtail millet and finger millet. Draft genome sequences are also available for proso millet and barnyard millet. The availability of whole and draft genome sequences of these millets can be used for further studies, such as SNP identification, next-generation sequencing-based allele discovery, association and linkage map construction, identification of candidate genes for agronomically important traits, and marker-assisted breeding programs. Two millets (little millet and kodo millet) have no genome sequences. Research is needed in the future to develop genome sequence for these two millets to improve the growth and production. In this chapter, we discuss about the details on genome sequences of small millets, how to mine genes from draft genome sequence of millets with finger millet as example, and comparative genomic studies of small millets. This chapter will help understand the current progress of research work related to comparative genomics approaches and mining genes across small millets.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
Abbreviations
- CISP:
-
Conserved intron scanning primer
- EST-SSR:
-
Expressed sequences tags-simple sequence repeat
- PHT1:
-
Phosphate transporter 1
- QTL:
-
Quantitative trait loci
- SSR:
-
Simple sequence repeat
- ZIP:
-
Zinc-regulated, iron-regulated transporter-like proteins
References
Alagarasan G, Dubey M, Aswathy KS, Chandel G (2017) Genome-wide identification of orthologous ZIP genes associated with zinc and iron translocation in Setaria italica. Front Plant Sci 8:775
Amadou I, Gbadamosi, Ole G (2011) Millet-based traditional processed foods and beverages-a review. Cereal Foods World 56:115–121
Annor GA, Tyl C, Marcone M, Ragaee S, Marti A (2017) Why do millets have slower starch and protein digestibility than other cereals? Trends Food SciTech 1:73–83
Arya L, Chauhan D, Yadav Y, Verma M (2014) Transferability of simple sequence repeat (SSR) markers developed in finger millet, and pearl millet to kodo millet and barnyard millet. In: Innovative approach in stem cell research, cancer biology and applied biotechnology, pp 60–64
Babu B, Rashmi C, Sood S (2018a) Cross transferability of finger millet and maize genomic SSR markers for genetic diversity and population structure analysis of barnyard millet. Indian J Genet Plant Breed 78:364–372
Babu BK, Joshi A, Sood S, Agrawal PK (2017) Identification of microsatellite markers for finger millet genomics application through cross transferability of rice genomic SSR markers. Indian J Genet 77:92–98
Babu BK, Sood S, Kumar D, Joshi A, Pattanayak A, Kant L, Upadhyaya HD (2018b) Cross-genera transferability of rice and finger millet genomic SSRs to barnyard millet (Echinochloa spp.). 3 Biotech 8:1–10
Bennetzen JL, Schmutz J, Wang H (2012) Reference genome sequence of the model plant Setaria. Nature Biotech 30:555–561
Bolser D, Staines DM, Pritchard E, Kersey P (2016) Ensembl plants: integrating tools for visualizing, mining, and analyzing plant genomics data. In: Edwards D (ed) Plant bioinformatics, Methods in molecular biology. Humana Press, New York, pp 115–140
Bora P, Ragaee S, Marcone M (2019) Characterisation of several types of millets as functional food ingredients. Int J Food Sci Nutr 70:1–11
Ceasar SA, Hodge A, Baker A, Baldwin SA (2014) Phosphate concentration and arbuscular mycorrhizal colonisation influence the growth, yield and expression of twelve PHT1 family phosphate transporters in foxtail millet (Setaria italica). PLoS One 9(9):e108459
Ceasar SA, Maharajan T, Ajeesh Krishna TP, Ramakrishnan M, Victor Roch G, Satish L, Ignacimuthu S (2018) Finger millet [Eleusine coracana (L.) Gaertn.] improvement: current status and future interventions of whole genome sequence. Front. Plant Sci 9:1054
Conrad LJ, Khanday I, Johnson C, Guiderdoni E, An G, Vijayraghavan U, Sundaresan V (2014) The polycomb group gene EMF 2B is essential for maintenance of floral meristem determinacy in rice. Plant J 80:883–894
David RHA, Ramakrishnan M, Maharajan T, Barathi Kannan K, Babu GA, Daniel MA, Ignacimuthu S (2021) Mining QTL and genes for root traits and biochemical parameters under vegetative drought in south Indian genotypes of finger millet (Eleusine coracana (L.) Gaertn) by association mapping and in silico comparative genomics. Biocatalysis Agril Biotech 32:101935
Devi PB, Vijayabharathi R, Sathyabama S, Malleshi NG, Priyadarisini VB (2014) Health benefits of finger millet (Eleusine coracana L.) polyphenols and dietary fiber: a review. J Food Sci Tech 51:1021–1040
Dong Y, Wang YZ (2015) Seed shattering: from models to crops. Front Plant Sci 6:476
Dykes L, Rooney L (2007) Phenolic compounds in cereal grains and their health benefits. Cereal Foods World 52:105–111
Fu Z, Song J, Zhao J, Jameso PE (2019) Identification and expression of genes associated with the abscission layer controlling seed shattering in Lolium perenne. AoB Plants 11:ply076
Gong L, Cao W, Chi H, Wang J, Zhang H, Liu J, Sun B (2018) Whole cereal grains and potential health effects: involvement of the gut microbiota. Food Res Int 103:84–102
Goodstein DM, Shu S, Howson R, Neupane R, Hayes RD, Fazo J, Rokhsar DS (2012) Phytozome: a comparative platform for green plant genomics. Nucl acids Res 40:D1178–D1186
Gull A, Prasad K, Kumar P (2015) Optimization and functionality of millet supplemented pasta. Food Sci Tech 35:626–632
Guo L, Qiu J, Ye C, Jin G, Mao L, Zhang H, Fan L (2017) Echinochloa crus-galli genome analysis provides insight into its adaptation and invasiveness as a weed. Nature Comm 8:1–10
Gupta P, Naithani S, Tello Ruiz MK, Chougule K, D’Eustachio P, Fabregat A, Jaiswal P (2016) Gramene database: navigating plant comparative genomics resources. Curr Plant Biol 7:10–15
Haque M, Martinek P, Watanabe N, Kuboyama T (2011) Genetic mapping of gibberellic acid-sensitive genes for semi-dwarfism in durum wheat. Cereal Res Comm 39:171–178
Hasan M, Maheshwari C, Garg NK, Kumar M (2019) Millets: Nutri-cereals. Biotech Express 6:18–21
Hatakeyama M, Aluri S, Balachadran MT, Sivarajan SR, Patrignani A, Grüter S, Poveda L, Shimizu-Inatsugi R, Baeten J, Francoijs KJ (2017) Multiple hybrid de novo genome assembly of finger millet, an orphan allotetraploid crop. DNA Res 25:39–47
Hittalmani S, Mahesh H, Shirke MD, Biradar H, Uday G, Aruna Y, Lohithaswa H, Mohanrao A (2017) Genome and transcriptome sequence of finger millet (Eleusine coracana (L.) Gaertn.) provides insights into drought tolerance and nutraceutical properties. BMC Genomics 18:1–16
Krishna TA, Maharajan T, David RHA, Ramakrishnan M, Ceasar SA, Duraipandiyan V, Ignacimuthu S (2018) Microsatellite markers of finger millet (Eleusine coracana (L.) Gaertn) and foxtail millet (Setaria italica (L.) Beauv) provide resources for cross-genome transferability and genetic diversity analyses in other millets. Biocatalysis Agrl Biotech 16:493–501
Krishna TA, Maharajan T, Roch GV, Ramakrishnan M, Ceasar SA, Ignacimuthu S (2020) Hybridization and hybrid detection through molecular markers in finger millet [Eleusine coracana (L.) Gaertn.]. J Crop Improv 34:335–355
Kumari K, Muthamilarasan M, Misra G, Gupta S, Subramanian A, Parida SK, Prasad M (2013) Development of eSSR-markers in Setaria italica and their applicability in studying genetic diversity, cross-transferability and comparative mapping in millet and non-millet species. PLoS One 8:e67742
Li C, Zhou A, Sang T (2006) Rice domestication by reducing shattering. Sci 311:1936–1939
Lin Z, Li X, Shannon LM, Yeh CT, Wang ML, Bai G, Yu J (2012) Parallel domestication of the Shattering1 genes in cereals. Nat Genet 44:720–724
Maharajan T, Antony Ceasar S, Ajeesh Krishna TP, Ignacimuthu S (2021) Finger millet [Eleusine coracana (L.) Gaertn]: An orphan crop with a potential to alleviate the calcium deficiency in the semi-arid tropics of Asia and Africa. Front Sust Food Syst 5:684447
Maharajan T, Ceasar SA, Ajeesh Krishna TP, Ramakrishnan M, Duraipandiyan V, Naif Abdulla AD, Ignacimuthu S (2018) Utilization of molecular markers for improving the phosphorus efficiency in crop plants. Plant Breed 137:10–26
Maharajan T, Ceasar SA, Krishna TPA, Ignacimuthu S (2019) Phosphate supply influenced the growth, yield and expression of PHT1 family phosphate transporters in seven millets. Planta 250:1433–1448
Maharajan T, Roch GV, Ceasar SA (2021b) Recent advancements of molecular breeding and functional genomics for improving nitrogen-, phosphorus-and potassium-use efficiencies in wheat. In: Hossain MA et al (eds) Molecular breeding in wheat, maize and sorghum: strategies for improving abiotic stress tolerance and yield. CAB International, Wallingford, pp 170–196
Michaelraj P, Shanmugam A (2013) A study on millets based cultivation and consumption in India. Int J Marketing Finan Serv Manag Res 2:49–58
Muthamilarasan M, Prasad M (2021) Small millets for enduring food security amidst pandemics. Trends Plant Sci 26:33–40
Nambiar VS, Dhaduk J, Sareen N, Shahu T, Desai R (2011) Potential functional implications of pearl millet (Pennisetum glaucum) in health and disease. J Appl Pharmal Sci 10:62–67
Neelam Y, Kanchan C, Alka S, Alka G (2013) Evaluation of hypoglycemic properties of kodo millet based food products in healthy subjects. IOSR J Pharmacy 3:14–20
Ookawa T, Hobo T, Yano M, Murata K, Ando T, Miura H, Matsuoka M (2010) New approach for rice improvement using a pleiotropic QTL gene for lodging resistance and yield. Nat Commun 1:1–11
Pearce S, Saville R, Vaughan SP, Chandler PM, Wilhelm EP, Sparks CA, Thomas SG (2011) Molecular characterization of Rht-1 dwarfing genes in hexaploid wheat. Plant Physiol 157:1820–1831
Pudake RN, Mehta CM, Mohanta TK, Sharma S, Varma A, Sharma AK (2017) Expression of four phosphate transporter genes from finger millet (Eleusine coracana L.) in response to mycorrhizal colonization and pi stress. 3 Biotech 7:17
Puranik S, Sahu PP, Beynon S, Srivastava RK, Sehgal D, Ojulong H, Yadav R (2020) Genome-wide association mapping and comparative genomics identifies genomic regions governing grain nutritional traits in finger millet (Eleusine coracana L. Gaertn). Plants, People, Planet 2(6):649–662
Radhika G, Sathya RM, Ganesan A, Saroja R, Vijayalakshmi P, Sudha V, Mohan V (2011) Dietary profile of urban adult population in South India in the context of chronic disease epidemiology (CURES–68). Public Health Nutr 14:591–598
Rajput SG, Plyler-Harveson T, Santra DK (2014) Development and characterization of SSR markers in proso millet based on switchgrass genomics. Am J Plant Sci 5:175–186
Ramakrishna C, Singh S, Raghavendrarao S, Padaria JC, Mohanty S, Sharma TR, Solanke AU (2018) The membrane tethered transcription factor EcbZIP17 from finger millet promotes plant growth and enhances tolerance to abiotic stresses. Sci Rep 8:1–14
Ramakrishnan M, Antony Ceasar S, Duraipandiyan V, Vinod KK, Kalpana K, Al-Dhabi NA, Ignacimuthu S (2016) Tracing QTLs for leaf blast resistance and agronomic performance of finger millet (Eleusine coracana (L.) Gaertn.) genotypes through association mapping and in silico comparative genomics analyses. PLoS One 11(7):e0159264
Ramakrishnan M, Ceasar SA, Vinod KK, Duraipandiyan V, Ajeesh Krishna TP, Upadhyaya HD, Ignacimuthu S (2017) Identification of putative QTLs for seedling stage phosphorus starvation response in finger millet (Eleusine coracana L. Gaertn.) by association mapping and cross species synteny analysis. PLoS One 12(8):e0183261
Roch GV, Maharajan T, Krishna TA, Ignacimuthu S, Ceasar SA (2020) Expression of PHT1 family transporter genes contributes for low phosphate stress tolerance in foxtail millet (Setaria italica) genotypes. Planta 252(6):1–9
Saleh AS, Zhang Q, Chen J, Shen Q (2013) Millet grains: nutritional quality, processing, and potential health benefits. Compr Rev Food Sci Food Safety 12:281–295
Shah L, Yahya M, Shah SMA, Nadeem M, Ali A, Ali A, Ma C (2019) Improving lodging resistance: using wheat and rice as classical examples. Int J Mol Sci 20:4211
Singh P, Raghuvanshi RS (2012) Finger millet for food and nutritional security. Afr J Food Sci 6:77–84
Tian B, Luan S, Zhang L, Liu Y, Zhang L, Li H (2018) Penalties in yield and yield associated traits caused by stem lodging at different developmental stages in summer and spring foxtail millet cultivars. Field Crops Res 217:104–112
Vinoth A, Ravindhran R (2017) Biofortification in millets: a sustainable approach for nutritional security. Front Plant Sci 8:29
Wang ML, Barkley NA, Yu JK, Dean RE, Newman ML, Sorrells ME, Pederson GA (2005) Transfer of simple sequence repeat (SSR) markers from major cereal crops to minor grass species for germplasm characterization and evaluation. Plant Genet Resour 3:45–57
Watanabe M (1999) Antioxidative phenolic compounds from Japanese barnyard millet (Echinochloa utilis) grains. J Agrl Food Chem 47:4500–4505
Wheeler T, Von Braun J (2013) Climate change impacts on global food security. Science 341:508–513
Xiao CL, Chen Y, Xie SQ et al (2017) MECAT: fast mapping, error correction, and de novo assembly for single-molecule sequencing reads. Nat Methods 14:1072–1074
Yano K, Ookawa T, Aya K, Ochiai Y, Hirasawa T, Ebitani T, Matsuoka M (2015) Isolation of a novel lodging resistance QTL gene involved in strigolactone signaling and its pyramiding with a QTL gene involved in another mechanism. Mol Plant 8:303–314
Yoon J, Cho LH, Kim SL, Choi H, Koh HJ, An G (2014) The BEL 1-type homeobox gene SH 5 induces seed shattering by enhancing abscission-zone development and inhibiting lignin biosynthesis. Plant J 79:717–728
Zhang G, Liu X, Quan Z, Cheng S, Xu X, Pan S, Wang J (2012) Genome sequence of foxtail millet (Setaria italica) provides insights into grass evolution and biofuel potential. Nat Biotech 30:549–554
Zhong J, Peng Z, Peng Q, Cai Q, Peng W, Chen M, Yao J (2018) Regulation of plant height in rice by the Polycomb group genes OsEMF2b, OsFIE2 and OsCLF. Plant Sci 267:157–167
Zou C, Li L, Miki D, Li D, Tang Q, Xiao L, Zhang H (2019) The genome of broomcorn millet. Nat Commun 10:1–11
Author information
Authors and Affiliations
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2022 The Author(s), under exclusive license to Springer Nature Singapore Pte Ltd.
About this chapter
Cite this chapter
Maharajan, T., Ceasar, S.A., Krishna, T.P.A., Ignacimuthu, S. (2022). Mining Genes and Markers Across Minor Millets Using Comparative Genomics Approaches. In: Pudake, R.N., Solanke, A.U., Sevanthi, A.M., Rajendrakumar, P. (eds) Omics of Climate Resilient Small Millets. Springer, Singapore. https://doi.org/10.1007/978-981-19-3907-5_9
Download citation
DOI: https://doi.org/10.1007/978-981-19-3907-5_9
Published:
Publisher Name: Springer, Singapore
Print ISBN: 978-981-19-3906-8
Online ISBN: 978-981-19-3907-5
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)