Abstract
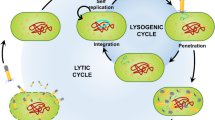
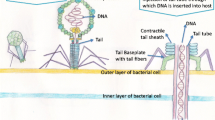
Bacteriophages (or simply ‘phages’) are viruses that replicate themselves within a host bacterial cell following infection of the host. For this project, environmental soil and water samples from around Singapore were collected to screen for phages, which have a myriad of purposes, due to their high specificity to their hosts. For instance, phages, coupled with genetic engineering of phage DNA, may provide an alternative to treating bacterial infections due to rapid replication times, and potential for non-intrusive therapy. Phages may also be used for diagnostics purposes, which allows scientists to potentially confirm the presence of a species of bacteria, and thus determine the need for the use of specific antibiotics. This research focuses on Staphylococcus aureus and Carbapanem-resistant Enterobacteriaceae, two of the species of the ESKAPE pathogens (Enterococcus faecium, S. aureus, Klebsiella pneumoniae, Acinetobacter baumannii, Pseudomonas aeruginosa, and Enterobacter species). The research aims to screen and isolate phages specific to these bacterial species (as they have high potential to achieve antibiotic resistance) and study them via purification, restriction enzyme digestion analysis and phenotypic characterisation. As a short summary, six distinct phages were found through the use of phenotypic and genotypic characterisation. Three of these phages targeted the Gram-positive S. aureus and Streptococcus pneumoniae, whereas the other three targeted Gram-negative K. pneumoniae. In particular, three of six studied phages targeted Carbapanem-resistant K. pneumoniae. The results of this study could provide insight into the species of phages present in Singapore, and whether they could be used for diagnostics and therapy Aslam (A Treatise on Electricity and Magnetism. Oxford, Clarendon, pp. 68–73, 2018 [1]), Dennehy (The Quarterly Review of Biology 85:109–109, 2010 [2])
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Aslam, B., Wang, W., Arshad, M. I., Khurshid, M., Muzammil, S., Rasool, M. H., Nisar, M. A., Alvi, R. F., Aslam, M. A., Qamar, M. U., Salamat, M. K. F., & Baloch, Z. (2018, October 10). Antibiotic resistance: A rundown of a global crisis. Infection and Drug Resistance. https://doi.org/10.2147/IDR.S173867 J. Clerk Maxwell, A Treatise on Electricity and Magnetism, 3rd ed., vol. 2. Oxford: Clarendon, 1892, pp. 68–73. https://doi.org/10.2147/IDR.S173867
Dennehy, J. J. (2010). Bacteriophage ecology: Population growth, evolution, and impact of bacterial viruses. The Quarterly Review of Biology, 85(1), 109–109. https://doi.org/10.1086/650260
Dedrick, R. M., Guerrero-Bustamante, C. A., Garlena, R. A., Russell, D. A., Ford, K., Harris, K., Gilmour, K. C., Soothill, J., Jacobs-Sera, D., Schooley, R. T., Hatfull, G. F., & Spencer, H. (2019). Engineered bacteriophages for treatment of a patient with a disseminated drug-resistant Mycobacterium Abscessus. Nature Medicine, 25(5), 730–733. https://doi.org/10.1038/s41591-019-0437-z
Singh, A., Poshtiban, S., & Evoy, S. (2013). Recent advances in bacteriophage based biosensors for food-borne pathogen detection. Sensors, 13(2), 1763–1786. https://doi.org/10.3390/s130201763
Lee, J. H., Domaille, D. W., & Cha, J. N. (2012). Amplified protein detection and identification through dna-conjugated m13 bacteriophage. ACS Nano, 6(6), 5621–5626. https://doi.org/10.1021/nn301565e
O’Sullivan, L., Buttimer, C., McAuliffe, O., Bolton, D., & Coffey, A. (2016). Bacteriophage-based tools: Recent advances and novel applications. F1000Research, 5, 2782. https://doi.org/10.12688/f1000research.9705.1
Synthetic phages with programmable specificity. (n.d.). ScienceDaily. Retrieved June 28, 2020, from https://www.sciencedaily.com/releases/2019/11/191104112838.htm
Merwe, R. G. van der, Helden, P. D. van, Warren, R. M., Sampson, S. L., & Pittius, N. C. G. van. (2014). Phage-based detection of bacterial pathogens. Analyst, 139(11), 2617–2626. https://doi.org/10.1039/C4AN00208C
Keen, E. C. (2012). Phage therapy: Concept to cure. Frontiers in Microbiology, 3. https://doi.org/10.3389/fmicb.2012.00238
Acknowledgements
We would sincerely like to thank Prof. Juan Pablo Bifani for offering us this opportunity, as well as for guiding us through the project. We would also like to thank Liew Jun Hao and Barry Choo for supervising and providing assistance during our research at the NUS Department of Microbiology and Immunology laboratories, as well as the laboratory staff and fellow researchers at the laboratories. Lastly, we would like to thank our research mentor, Dr. Tang Hock Chun from NUS High School of Mathematics and Science for his support throughout the entirety of this project.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2021 The Author(s), under exclusive license to Springer Nature Singapore Pte Ltd.
About this paper
Cite this paper
Tan, C.Y., Ong, J.H.M., Bifani, J.P. (2021). Screening and Characterisation of Novel Environmental Phages. In: Guo, H., Ren, H., Kim, N. (eds) IRC-SET 2020. Springer, Singapore. https://doi.org/10.1007/978-981-15-9472-4_28
Download citation
DOI: https://doi.org/10.1007/978-981-15-9472-4_28
Published:
Publisher Name: Springer, Singapore
Print ISBN: 978-981-15-9471-7
Online ISBN: 978-981-15-9472-4
eBook Packages: Physics and AstronomyPhysics and Astronomy (R0)