Abstract
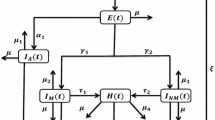
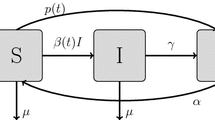
This study presents a family of stochastic models for the dynamics of influenza in a closed human population. We consider treatment for the disease in the form of vaccination and incorporate the periods of effectiveness of the vaccine and infectiousness for the individuals in the population. Our model is a SVIR model, with trinomial transition probabilities, where all individuals who recover from the disease acquire permanent natural immunity against the strain of the disease. A special SVIR model in the stochastic family based on correlated vaccination and infection probabilities at any instant is presented. The methods of maximum likelihood and expectation–maximization algorithm are applied to find estimates for the parameters of the chain. Moreover, estimators for some special epidemiological control parameters, such as the basic reproduction number, are computed. A numerical simulation example is presented to find the MLE of the parameters of the model.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
CDC Estimating Seasonal Influenza-Associated Deaths. https://www.cdc.gov/flu/about/disease/us_flu-related_deaths.htm
D. Iuliano, K.Roguski, H. Chang, D. Muscatello, R. Palekar, S. Tempia et al., Estimates of global seasonal influenza-associated respiratory mortality: a modelling study. Lancet 391(10127), 1285–1300, 31 Mar 2018
WHO Influenza, burden of disease. http://www.who.int/influenza/surveillance_monitoring/bod/en/
CDC Flu symptoms and complications. https://www.cdc.gov/flu/consumer/symptoms.htm/
CDC Types of Influenza Viruses. https://www.cdc.gov/flu/about/viruses/types.htm
CDC How Flu Spreads. https://www.cdc.gov/flu/about/disease/spread.htm
CDC Different types of flu vaccines. https://www.cdc.gov/flu/vaccines/index.htm
H.C. Tuckwell, R,J. Williams, Some properties of a simple stochastic epidemic model of SIR type. Math. Biosci. 208, 76–97 (2007)
D. Wanduku, Threshold conditions for a family of epidemic dynamic models for malaria with distributed delays in a non-random environment, Int. J. Biomath. 11(6) 1850085 (46 pages) (2018). https://doi.org/10.1142/S1793524518500857
P. Witbooi, G.E. Muller, G.J. Van Schalkwyk, Vaccination control in a stochastic SVIR epidemic model. Comput. Math. Methods Med. 2015, Article ID 271654, 9 pages (2015). https://doi.org/10.1155/2015/271654
D. Wanduku, Complete global analysis of a two-scale network SIRS epidemic dynamic model with distributed delay and random perturbation. Appl. Math. Comput. 294, 49–76 (2017)
D. Wanduku, G.S. Ladde, Global properties of a two-scale network stochastic delayed human epidemic dynamic model. Nonlinear Anal. Real World Appl. 13, 794–816 (2012)
F. Etbaigha, A. Willms, Z. Poljak, An SEIR model of influenza A virus infection and reinfection within a farrow-to-finish swine farm. PLOS ONE 13(9), e0202493. https://doi.org/10.1371/journal.pone.0202493
M.E. Alexander, C. Bowman, S.M. Moghadas et al., Vaccination model for transmission dynamics of influenza. SIAM J. Appl. Dyn. Syst. 3, 503–524 (2004)
D. Wanduku, The stochastic extinction and stability conditions for nonlinear malaria epidemics. Math. Biosci. Eng. 16, 3771–3806 (2019)
A. Scherer, A.R. McLean, Mathematical models of vaccination. Brit. Med. Bull. 62, 187–199 (2002)
H.S. Rodrigues, M.T.T. Monteiro, D.F.M. Torres, Vaccination models and optimal control strategies to dengue. Math. Biosci. 247, 1–12 (2014)
A. Lloyd, Introduction to Epidemiological Modeling: Basic Models and Their Properties 23 Jan. 2017
M. Li, J.R. Graef, L. Wang, J. Karsai, Global dynamics of a SEIR model with varying total population size. Math. Biosci. 160, 191–213 (1999)
C.M. Kribs-Zaleta, J.X. Velasco-Hernández, A simple vaccination model with multiple endemic states. J. Math. Biosci. 164, 183–201 (2000)
D. Wanduku, G.S. Ladde, Fundamental properties of a two-scale network stochastic human epidemic dynamic model. Neural Parallel Sci. Comput. 19, 229–270 (2011)
M. Ferrante, E. Ferraris, C. Rovira, On a stochastic epidemic SEIHR model and its diffusion approximation. TEST 25, 482 (2016). https://doi.org/10.1007/s11749-015-0465-z
R. Yaesoubi, T. Cohen, Generalized Markov models of infectious disease spread: a novel framework for developing dynamic health policies. Eur. J. Oper. Res. 215(3) (2011)
M. Greenwood, On the statistical measure of infectiousness. J. Hyg. Camb. 31, 336 (1931)
H. Abbey, An examination of the Reed-Frost theory of epidemics. Hum. Biol. 24, 201 (1952)
J. Jacquez, A note on chain binomial models of epidemic spread: what is wrong with the Reed-Frost Model. Mathem. Biosci. 87(1), 73–82 (1987)
J. Gani, D. Jerwood, Markov chain methods in chain binomial epidemic models. Biometrics 27 (1971)
J. Lin, V. Andreasen, S.A. Levin, Dynamics of influenza A; the linear three strain model. Math. Biosci. 162, 33–51 (1999)
M. Ajelli, P. Poletti, A. Melegaro S. Merler, The role of different social contexts in shaping influenza transmission during the 2009 pandemic. Sci. Rep. 4, Article number: 7218 (2014)
L. Allen, An introduction to stochastic epidemic models, in Mathematical Epidemiology ed. by Brauer F., van den Driessche P., Wu J. Lecture Notes in Mathematics, vol. 1945 (Springer, Berlin, Heidelberg), pp. 81–130
M. Gupta, Y. Chen, Theory and use of the EM Algorithm. Found. Trends Sig. Process. 4(3) (2010)
J. Bilmes, A gentle tutorial of the EM algorithm and it’s Application to parameter estimation for Gausian mixture and hidden Markov models. Int. Comput. Sci. Inst. (1998)
G. Casella, R. Berger, Statistical Inference, 2 edn. (Duxbury, 2002)
K. Dietz, The estimation of the basic reproduction number for infectious diseases. Stat Methods Med Res. 2(1), 23–41 (1993)
J. Jones, Notes on \(R_0\) (Stanford University, Department of Anthropological Sciences, 2007)
P. Holme, N. Masuda, The basic reproduction number as a predictor for epidemic outbreaks in temporal networks. PLoS ONE 10(3) (2015)
E. Vergu, H. Busson, P. Ezanno, Impact of the infection period distribution on the epidemic spread in a metapopulation model. PLoS ONE 5(2) (2010)
CDC-FLUVIEW(1-17-2019). https://gis.cdc.gov/grasp/fluview/fluportaldashboard.html; https://www.cdc.gov/flu/weekly/weeklyarchives2017-2018/Week39.htm
FluVaxView 2017-18 Season. https://www.cdc.gov/flu/fluvaxview/coverage-1718estimates.htm
US census Bureau, 2018 National and State Population Estimates. https://www.census.gov/newsroom/press-kits/2018/pop-estimates-national-state.html
Acknowledgements
This work was completed during the graduate studies of Cameron Newman, Omotomilola Jegede and Mymuna Monem in the department of Mathematical Sciences of Georgia Southern University (GSU) in 2018–2019 academic year, supervised by Dr. Wanduku. Mymuna Monem was part of the general group discussions.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2020 Springer Nature Singapore Pte Ltd.
About this chapter
Cite this chapter
Wanduku, D., Newman, C., Jegede, O., Oluyede, B. (2020). Modeling the Stochastic Dynamics of Influenza Epidemics with Vaccination Control, and the Maximum Likelihood Estimation of Model Parameters. In: Dutta, H. (eds) Mathematical Modelling in Health, Social and Applied Sciences. Forum for Interdisciplinary Mathematics. Springer, Singapore. https://doi.org/10.1007/978-981-15-2286-4_2
Download citation
DOI: https://doi.org/10.1007/978-981-15-2286-4_2
Published:
Publisher Name: Springer, Singapore
Print ISBN: 978-981-15-2285-7
Online ISBN: 978-981-15-2286-4
eBook Packages: Mathematics and StatisticsMathematics and Statistics (R0)