Abstract
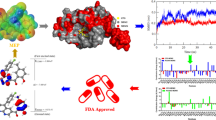
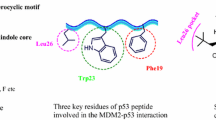
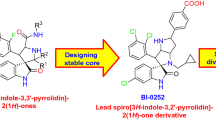
Spiro compounds have drawn ever-increasing attention in drug discovery because of its prevalence in natural products/drugs and unique 3D structural features. A large number of spiro compounds have been proved to possess diverse bioactivities, and some of them have advanced into clinical trials for the treatment of diseases. The interruption of MDM2–p53 protein-protein interactions has been highly pursued as an attractive therapeutic strategy for cancer therapy. A large number of small-molecule inhibitors have been identified based on the well-defined MDM2–p53 interactions. Currently, several small-molecule inhibitors such as SAR405838, APG-115, MK-8242, DS-3032b, NVP-CGM097, RG7112, RG7388, and AMG 232 are undergoing clinical assessment for cancer therapy. In this chapter, we focus on the identification of spirooxindole containing small-molecule inhibitors (SAR405838, APG-115, RG7388, RO8994, RO2468, and RO5353), strategies employed for optimizations, structure–activity relationship studies (SARs) as well as their biochemical profiles. The identification of these lead compounds makes spirooxindoles promising scaffolds in designing potent inhibitors targeting MDM2–p53 interactions. Based on the SARs and the co-crystal structures of p53–MDM2 complexes, we first tentatively propose the prolinamide-based ‘3+1’ model for designing potential MDM2 inhibitors.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Bieging KT, Mello SS, Attardi LD (2014) Unravelling mechanisms of p53-mediated tumour suppression. Nat Rev Cancer 14:359–370
Wu X, Bayle JH, Olson D et al (1993) The p53-mdm-2 autoregulatory feedback loop. Genes Dev 7:1126–1132
Harris SL, Levine AJ (2005) The p53 pathway: positive and negative feedback loops. Oncogene 24:2899–2908
Chen F, Wang W, El-Deiry WS (2010) Current strategies to target p53 in cancer. Biochem Pharmacol 80:724–730
Moll UM, Petrenko O (2003) The MDM2-p53 Interaction. Mol Cancer Res 1:1001–1008
Feki A, Irminger-Finger I (2004) Mutational spectrum of p53 mutations in primary breast and ovarian tumors. Crit Rev Oncol Hematol 52:103–116
Momand J, Jung D, Wilczynski S et al (1998) The MDM2 gene amplification database. Nucleic Acids Res 26:3453–3459
Forslund A, Zeng Z, Qin L-X et al (2008) MDM2 gene amplification is correlated to tumor progression but not to the presence of SNP309 or TP53 mutational status in primary colorectal cancers. Mol Cancer Res 6:205–211
Dickens MP, Fitzgerald R, Fischer PM (2010) Small-molecule inhibitors of MDM2 as new anticancer therapeutics. Semin Cancer Biol 20:10–18
Barakat K, Mane J, Friesen D et al (2010) Ensemble-based virtual screening reveals dual-inhibitors for the p53–MDM2/MDMX interactions. J Mol Graph Model 28:555–568
Shangary S, Wang S (2009) Small-molecule inhibitors of the MDM2-p53 protein-protein interaction to reactivate p53 function: a novel approach for cancer therapy. Annu Rev Pharmacol Toxicol 49:223–241
Tanimura S, Ohtsuka S, Mitsui K et al (1999) MDM2 interacts with MDMX through their RING finger domains. FEBS Lett 447:5–9
Zhao Y, Aguilar A, Bernard D et al (2015) Small-molecule inhibitors of the MDM2–p53 protein-protein interaction (MDM2 inhibitors) in clinical trials for cancer treatment. J Med Chem 58:1038–1052
Brown CJ, Cheok CF, Verma CS et al (2011) Reactivation of p53: from peptides to small molecules. Trends Pharmacol Sci 32:53–62
Saha T, Kar RK, Sa G (2015) Structural and sequential context of p53: a review of experimental and theoretical evidence. Prog Biophys Mol Biol 117:250–263
Nero TL, Morton CJ, Holien JK et al (2014) Oncogenic protein interfaces: small molecules, big challenges. Nat Rev Cancer 14:248–262
Guo W, Wisniewski JA, Ji H (2014) Hot spot-based design of small-molecule inhibitors for protein–protein interactions. Bioorg Med Chem Lett 24:2546–2554
Kussie PH, Gorina S, Marechal V et al (1996) Structure of the MDM2 oncoprotein bound to the p53 tumor suppressor transactivation domain. Science 274:948–953
Vassilev LT, Vu BT, Graves B et al (2004) In vivo activation of the p53 pathway by small-molecule antagonists of MDM2. Science 303:844–848
Vassilev LT (2007) MDM2 inhibitors for cancer therapy. Trends Mol Med 13:23–31
Lv P-C, Sun J, Zhu H-L (2015) Recent advances of p53-MDM2 small molecule inhibitors (2011-Present). Curr Med Chem 22:618–626
Zak K, Pecak A, Rys B et al (2013) Mdm2 and MdmX inhibitors for the treatment of cancer: a patent review (2011–present). Expert Opin Ther Pat 23:425–448
Wang W, Hu Y (2012) Small molecule agents targeting the p53-MDM2 pathway for cancer therapy. Med Res Rev 32:1159–1196
Liu L, Bernard D, Wang S (2015) Case study: discovery of inhibitors of the MDM2–p53 protein-protein interaction. In: Meyerkord CL, Fu H (eds) Protein-protein interactions, vol 1278. Methods in molecular biology. Springer, New York, pp 567–585
Zhang B, Golding BT, Hardcastle IR (2015) Small-molecule MDM2-p53 inhibitors: recent advances. Future Med Chem 7:631–645
Hardcastle IR (2014) Targeting the MDM2–p53 protein-protein interaction: design, discovery, and development of novel anticancer agents. In: Neidle S (ed) Cancer drug design and discovery, 2nd edn. Academic Press, San Diego, pp 391–426
Popowicz GM, Dömling A, Holak TA (2011) The structure-based design of Mdm2/Mdmx–p53 inhibitors gets serious. Angew Chem Int Ed 50:2680–2688
Khoury K, Popowicz GM, Holak TA et al (2011) The p53-MDM2/MDMX axis—a chemotype perspective. MedChemComm 2:246–260
Lemos A, Leão M, Soares J et al (2016) Medicinal chemistry strategies to disrupt the p53–MDM2/MDMX interaction. Med Res Rev 36:789–844
Secchiero P, Bosco R, Celeghini C et al (2011) Recent advances in the therapeutic perspectives of Nutlin-3. Curr Pharm Design 17:569–577
Koblish HK, Zhao S, Franks CF et al (2006) Benzodiazepinedione inhibitors of the Hdm2:p53 complex suppress human tumor cell proliferation in vitro and sensitize tumors to doxorubicin in vivo. Mol Cancer Ther 5:160–169
Rew Y, Sun D, Yan X et al (2014) Discovery of AM-7209, a potent and selective 4-amidobenzoic acid inhibitor of the MDM2–p53 interaction. J Med Chem 57:10499–10511
Wang S, Sun W, Zhao Y et al (2014) SAR405838: an optimized inhibitor of MDM2–p53 interaction that induces complete and durable tumor regression. Cancer Res 74:5855–5865
Sun D, Li Z, Rew Y et al (2014) Discovery of AMG 232, a potent, selective, and orally bioavailable MDM2–p53 inhibitor in clinical development. J Med Chem 57:1454–1472
Canon J, Osgood T, Olson SH et al (2015) The MDM2 inhibitor AMG 232 demonstrates robust anti-tumor efficacy and potentiates the activity of p53-inducing cytotoxic agents. Mol Cancer Ther 14:649–658
Wang Y, Zhu J, Liu J et al (2014) Optimization beyond AMG 232: Discovery and SAR of sulfonamides on a piperidinone scaffold as potent inhibitors of the MDM2-p53 protein–protein interaction. Bioorg Med Chem Lett 24:3782–3785
Valat T, Masuya K, Baysang F et al (2014) Mechanistic study of NVP-CGM097: a potent, selective and species specific inhibitor of p53-Mdm2. Cancer Res 74:1798
Gessier F, Kallen J, Jacoby E et al (2015) Discovery of dihydroisoquinolinone derivatives as novel inhibitors of the p53–MDM2 interaction with a distinct binding mode. Bioorg Med Chem Lett 25:3621–3625
Holzer P, Masuya K, Furet P et al (2015) Discovery of a dihydroisoquinolinone derivative (NVP-CGM097): a highly potent and selective MDM2 inhibitor undergoing phase 1 clinical trials in p53wt tumors. J Med Chem 58:6348–6358
Tovar C, Graves B, Packman K et al (2013) MDM2 small-molecule antagonist RG7112 activates p53 signaling and regresses human tumors in preclinical cancer models. Cancer Res 73:2587–2597
Vu B, Wovkulich P, Pizzolato G et al (2013) Discovery of RG7112: a small-molecule MDM2 inhibitor in clinical development. ACS Med Chem Lett 4:466–469
Ding Q, Zhang Z, Liu J-J et al (2013) Discovery of RG7388, a potent and selective p53–MDM2 inhibitor in clinical development. J Med Chem 56:5979–5983
Aguilar A, Lu J, Liu L et al (2017) Discovery of 4-((3′R,4′S,5′R)-6″-Chloro-4′-(3-chloro-2-fluorophenyl)-1′-ethyl-2″-oxodispiro[cyclohexane-1,2′-pyrrolidine-3′,3″-indoline]-5′-carboxamido)bicyclo[2.2.2]octane-1-carboxylic Acid (AA-115/APG-115): a potent and orally active Murine Double Minute 2 (MDM2) inhibitor in clinical development. J Med Chem 60:2819–2839
Over B, Wetzel S, Grütter C et al (2013) Natural-product-derived fragments for fragment-based ligand discovery. Nat Chem 5:21–28
Galliford CV, Scheidt KA (2007) Pyrrolidinyl-spirooxindole natural products as inspirations for the development of potential therapeutic agents. Angew Chem Int Ed 46:8748–8758
Yu B, Yu Z, Qi P-P et al (2015) Discovery of orally active anticancer candidate CFI-400945 derived from biologically promising spirooxindoles: Success and challenges. Eur J Med Chem 95:35–40
Yeung BKS, Zou B, Rottmann M et al (2010) Spirotetrahydro β-Carbolines (Spiroindolones): a new class of potent and orally efficacious compounds for the treatment of malaria. J Med Chem 53:5155–5164
Rottmann M, McNamara C, Yeung BKS et al (2010) Spiroindolones, a potent compound class for the treatment of malaria. Science 329:1175–1180
White NJ, Pukrittayakamee S, Phyo AP et al (2014) Spiroindolone KAE609 for falciparum and vivax malaria. N Engl J Med 371:403–410
van Pelt-Koops JC, Pett HE, Graumans W et al (2012) The spiroindolone drug candidate NITD609 potently inhibits gametocytogenesis and blocks Plasmodium falciparum transmission to anopheles mosquito vector. Antimicrob Agents Chemother 56:3544–3548
Yu B, Qi P-P, Shi X-J et al (2014) Discovery of novel steroidal pyran–oxindole hybrids as cytotoxic agents. Steroids 88:44–52
Yu B, Shi X-J, Qi P-P et al (2014) Design, synthesis and biological evaluation of novel steroidal spiro-oxindoles as potent antiproliferative agents. J Steroid Biochem Mol Biol 141:121–134
Yu B, Yu D-Q, Liu H-M (2015) Spirooxindoles: promising scaffolds for anticancer agents. Eur J Med Chem 97:673–698
Yu B, Sun X-N, Shi X-J et al (2015) Efficient synthesis of novel antiproliferative steroidal spirooxindoles via the [3+2] cycloaddition reactions of azomethine ylides. Steroids 102:92–100
Bin Y, Yi-Chao Z, Xiao-Jing S et al (2016) Natural product-derived spirooxindole fragments serve as privileged substructures for discovery of new anticancer agents. Anticancer Agents Med Chem 16:1315–1324
Zhang Y-L, Li Y-F, Wang J-W et al Multicomponent assembly of novel antiproliferative steroidal dihydropyridinyl spirooxindoles. Steroids 109:22–28
Shi X-J, Yu B, Wang J-W et al (2016) Structurally novel steroidal spirooxindole by241 potently inhibits tumor growth mainly through ROS-mediated mechanisms. Sci Rep 6:31607
Yu B, Xing H, Yu D-Q et al (2016) Catalytic asymmetric synthesis of biologically important 3-hydroxyoxindoles: an update. Beilstein J Org Chem 12:1000–1039
Aguilar A, Sun W, Liu L et al (2014) Design of chemically stable, potent, and efficacious MDM2 inhibitors that exploit the retro-mannich ring-opening-cyclization reaction mechanism in spiro-oxindoles. J Med Chem 57:10486–10498
Ding K, Lu Y, Nikolovska-Coleska Z et al (2005) Structure-based design of potent non-peptide MDM2 inhibitors. J Am Chem Soc 127:10130–10131
Smart BE (2001) Fluorine substituent effects (on bioactivity). J Fluor Chem 109:3–11
Ding K, Lu Y, Nikolovska-Coleska Z et al (2006) Structure-based design of spiro-oxindoles as potent, specific small-molecule inhibitors of the MDM2–p53 interaction. J Med Chem 49:3432–3435
Lin J, Chen J, Elenbaas B et al (1994) Several hydrophobic amino acids in the p53 amino-terminal domain are required for transcriptional activation, binding to MDM-2 and the adenovirus 5 E1B 55-kD protein. Genes Dev 8:1235–1246
Picksley SM, Vojtesek B, Sparks A et al (1994) Immunochemical analysis of the interaction of p53 with MDM2;–fine mapping of the MDM2 binding site on p53 using synthetic peptides. Oncogene 9:2523–2529
Shangary S, Qin D, McEachern D et al (2008) Temporal activation of p53 by a specific MDM2 inhibitor is selectively toxic to tumors and leads to complete tumor growth inhibition. Proc Natl Acad Sci USA 105:3933–3938
Zhao Y, Liu L, Sun W et al (2013) Diastereomeric spirooxindoles as highly potent and efficacious MDM2 inhibitors. J Am Chem Soc 135:7223–7234
Zhao Y, Yu S, Sun W et al (2013) A Potent small-molecule inhibitor of the MDM2–p53 interaction (MI-888) achieved complete and durable tumor regression in mice. J Med Chem 56:5553–5561
Shu L, Li Z, Gu C et al (2013) Synthesis of a spiroindolinone pyrrolidinecarboxamide MDM2 antagonist. Org Proc Res Dev 17:247–256
Hoffman-Luca CG, Yang C-Y, Lu J et al (2015) Significant differences in the development of acquired resistance to the MDM2 inhibitor SAR405838 between in vitro and in vivo drug treatment. PLoS ONE 10:e0128807
Hoffman-Luca CG, Ziazadeh D, McEachern D et al (2015) Elucidation of acquired resistance to Bcl-2 and MDM2 inhibitors in acute leukemia in vitro and in vivo. Clin Cancer Res 21:2558–2568
Carry J-C, Garcia-Echeverria C (2013) Inhibitors of the p53/hdm2 protein–protein interaction—path to the clinic. Bioorg Med Chem Lett 23:2480–2485
Miyazaki M, Setoguchi M, Sugimoto Y et al (2012) Dispiropyrrolidine derivative. EP20120755073
Wang S, Sun W, Aguilar A et al (2014) Spiro-oxindole MDM2 antagonists. US 14/485,054
Phelps D, Bondra K, Seum S et al (2015) Inhibition of MDM2 by RG7388 confers hypersensitivity to X-radiation in xenograft models of childhood sarcoma. Pediatr Blood Cancer 62:1345–1352
Zhang Z, Ding Q, Liu J-J et al (2014) Discovery of potent and selective spiroindolinone MDM2 inhibitor, RO8994, for cancer therapy. Bioorg Med Chem 22:4001–4009
Zhang Z, Chu X-J, Liu J-J et al (2013) Discovery of potent and orally active p53-MDM2 inhibitors RO5353 and RO2468 for potential clinical development. ACS Med Chem Lett 5:124–127
Luk KC, So SS, Zhang J et al (2006) Oxindole derivatives. PCT/EP2006/063475
Ding Q, Jiang N, Yang S et al (2009) Spiroindolinone derivatives. US 12/272,870
Ding Q, Liu JJ, Zhang Z (2007) Spiroindolinone derivatives. PCT/EP2007/052038
Liu JJ, Tilley JW, Zhang Z (2009) 3,3-spiroindolinone derivatives. CN 200880016425
Liu JJ, Tilley JW, Zhang Z (2010) 3,3-spiroindolinone derivatives as anticancer agents. PCT/EP2010/051757
Liu JJ, Zhang Z (2008) Spiroindolinone derivatives. PCT/EP2008/055831
Chu XJ, Ding Q, Jiang N et al (2012) Substituted Spiro[3H-Indole-3,6′(5′H)-[1H]Pyrrolo[1,2c]Imidazole-1′,2(1H,2′H)-diones. US 20120071499
Pfafferott G, Oberhammer H, Boggs JE et al (1985) Geometric structure and pseudorotational potential of pyrrolidine. An ab initio and electron diffraction study. J Am Chem Soc 107:2305–2309
Cremer D, Pople JA (1975) General definition of ring puckering coordinates. J Am Chem Soc 97:1354–1358
Madison V (1977) Flexibility of the pyrrolidine ring in proline peptides. Biopolymers 16:2671–2692
Ramachandran GN, Lakshminarayanan AV, Balasubramanian R et al (1970) Studies on the conformation of amino acids XII. Energy calculations on prolyl residue. Biochim Biophys Acta (BBA)—Protein Struct 221:165–181
DeTar DF, Luthra NP (1977) Conformations of proline. J Am Chem Soc 99:1232–1244
Bertamino A, Soprano M, Musella S et al (2013) Synthesis, in vitro, and in cell studies of a new series of [Indoline-3,2′-thiazolidine]-based p53 modulators. J Med Chem 56:5407–5421
Ivanenkov YA, Vasilevski SV, Beloglazkina EK et al (2015) Design, synthesis and biological evaluation of novel potent MDM2/p53 small-molecule inhibitors. Bioorg Med Chem Lett 25:404–409
Kumar A, Gupta G, Bishnoi AK et al (2015) Design and synthesis of new bioisosteres of spirooxindoles (MI-63/219) as anti-breast cancer agents. Bioorg Med Chem 23:839–848
Li Bo ZR, He Gu, Li Guo, Wei Huang (2013) Molecular docking, QSAR and molecular dynamics simulation on spiro-oxindoles as MDM2 inhibitors. Acta Chim Sinica 71:1396–1403
Ribeiro CJA, Amaral JD, Rodrigues CMP et al (2014) Synthesis and evaluation of spiroisoxazoline oxindoles as anticancer agents. Bioorg Med Chem 22:577–584
Huang W, Cai L, Chen C et al (2015) Computational analysis of spiro-oxindole inhibitors of the MDM2-p53 interaction: insights and selection of novel inhibitors. J Biomol Struct Dyn 34:1–11
Zhou R, Wu Q, Guo M et al (2015) Organocatalytic cascade reaction for the asymmetric synthesis of novel chroman-fused spirooxindoles that potently inhibit cancer cell proliferation. Chem Commun 51:13113–13116
Wang S, Jiang Y, Wu S et al (2016) Meeting organocatalysis with drug discovery: asymmetric synthesis of 3,3′-Spirooxindoles fused with tetrahydrothiopyrans as novel p53-MDM2 inhibitors. Org Lett 18:1028–1031
Patil SP (2013) FOLICation: engineering approved drugs as potential p53–MDM2 interaction inhibitors for cancer therapy. Med Hypotheses 81:1104–1107
Acknowledgements
This work was supported by the National Natural Science Foundation of China (No. 81430085, 21372206, and 81703326, National Key Research Program of Proteins (No. 2016YFA0501800), Key Research Program of Henan Province (No. 1611003110100), Scientific Program of Henan Province (No. 182102310123), China Postdoctoral Science Foundation (No. 2018M630840), Key Research Program of Higher Education of Henan Province(No.18B350009), and the Starting Grant of Zhengzhou University (No. 32210533).
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2018 Springer Nature Singapore Pte Ltd.
About this chapter
Cite this chapter
Yu, B., Liu, HM. (2018). The Development of New Spirooxindoles Targeting the p53–MDM2 Protein-Protein Interactions for Cancer Therapy. In: Sheng, C., Georg, G. (eds) Targeting Protein-Protein Interactions by Small Molecules. Springer, Singapore. https://doi.org/10.1007/978-981-13-0773-7_8
Download citation
DOI: https://doi.org/10.1007/978-981-13-0773-7_8
Published:
Publisher Name: Springer, Singapore
Print ISBN: 978-981-13-0772-0
Online ISBN: 978-981-13-0773-7
eBook Packages: Chemistry and Materials ScienceChemistry and Material Science (R0)