Abstract
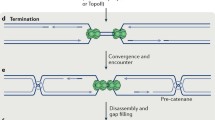
Termination of DNA replication forks takes place when two replication forks coming from neighbouring origins meet each other usually in the midpoint of the replicon. At this stage, the remaining fragments of DNA have to be unwound, all remaining DNA replicated and newly synthesised strands ligated to produce continuous sister chromatids. Finally, the replication machinery has to be taken off, chromatin re-assembled, and entwisted sister chromatids resolved topologically.
Over the last few decades, we have learned a lot about the assembly of the helicase and replisome and the initiation stage of DNA replication. We also know much more about the ability of forks to cope with replication stress. However, only within recent years we have gained the first glimpse of the mechanism of replication fork termination. In this chapter I will summarise the recent findings on replication termination, weigh this against the past literature and discuss relevant consequences and views for the future.
Similar content being viewed by others
References
Adham IM, Khulan J, Held T, Schmidt B, Meyer BI, Meinhardt A, Engel W (2008) Fas-associated factor (FAF1) is required for the early cleavage-stages of mouse embryo. Mol Hum Reprod 14:207–213
Ahuja AK, Jodkowska K, Teloni F, Bizard AH, Zellweger R, Herrador R, Ortega S, Hickson ID, Altmeyer M, Mendez J et al (2016) A short G1 phase imposes constitutive replication stress and fork remodelling in mouse embryonic stem cells. Nat Commun 7:10660
Ali N, Ismail IM, Khoder M, Shamy M, Alghamdi M, Costa M, Ali LN, Wang W, Eqani SA (2016) Polycyclic aromatic hydrocarbons (PAHs) in indoor dust samples from cities of Jeddah and Kuwait: levels, sources and non-dietary human exposure. Sci Total Environ 573:1607–1614
Balakrishnan L, Bambara RA (2013) Okazaki fragment metabolism. Cold Spring Harb Perspect Biol 5
Barkley LR, Song IY, Zou Y, Vaziri C (2009) Reduced expression of GINS complex members induces hallmarks of pre-malignancy in primary untransformed human cells. Cell Cycle 8:1577–1588
Bastia D, Zaman S (2014) Mechanism and physiological significance of programmed replication termination. Semin Cell Dev Biol 30:165–173
Baxter J, Diffley JF (2008) Topoisomerase II inactivation prevents the completion of DNA replication in budding yeast. Mol Cell 30:790–802
Been MD, Champoux JJ (1980) Breakage of single-stranded DNA by rat liver nicking-closing enzyme with the formation of a DNA-enzyme complex. Nucleic Acids Res 8:6129–6142
Bermejo R, Doksani Y, Capra T, Katou YM, Tanaka H, Shirahige K, Foiani M (2007) Top1- and Top2-mediated topological transitions at replication forks ensure fork progression and stability and prevent DNA damage checkpoint activation. Genes Dev 21:1921–1936
Blake D, Luke B, Kanellis P, Jorgensen P, Goh T, Penfold S, Breitkreutz BJ, Durocher D, Peter M, Tyers M (2006) The F-box protein Dia2 overcomes replication impedance to promote genome stability in Saccharomyces cerevisiae. Genetics 174:1709–1727
Brewer BJ, Fangman WL (1988) A replication fork barrier at the 3′ end of yeast ribosomal RNA genes. Cell 55:637–643
Brill SJ, DiNardo S, Voelkel-Meiman K, Sternglanz R (1987) Need for DNA topoisomerase activity as a swivel for DNA replication for transcription of ribosomal RNA. Nature 326:414–416
Bruderer RM, Brasseur C, Meyer HH (2004) The AAA ATPase p97/VCP interacts with its alternative co-factors, Ufd1-Npl4 and p47, through a common bipartite binding mechanism. J Biol Chem 279:49609–49616
Buchsbaum S, Morris C, Bochard V, Jalinot P (2007) Human INT6 interacts with MCM7 and regulates its stability during S phase of the cell cycle. Oncogene 26:5132–5144
Burger J, Merlet J, Tavernier N, Richaudeau B, Arnold A, Ciosk R, Bowerman B, Pintard L (2013) CRL2(LRR-1) E3-ligase regulates proliferation and progression through meiosis in the Caenorhabditis elegans germline. PLoS Genet 9:e1003375
Burrell RA, McClelland SE, Endesfelder D, Groth P, Weller MC, Shaikh N, Domingo E, Kanu N, Dewhurst SM, Gronroos E et al (2013) Replication stress links structural and numerical cancer chromosomal instability. Nature 494:492–496
Chowdhury P, Lin GE, Liu K, Song Y, Lin FT, Lin WC (2014) Targeting TopBP1 at a convergent point of multiple oncogenic pathways for cancer therapy. Nat Commun 5:5476
Conti C, Sacca B, Herrick J, Lalou C, Pommier Y, Bensimon A (2007) Replication fork velocities at adjacent replication origins are coordinately modified during DNA replication in human cells. Mol Biol Cell 18:3059–3067
Costa A, Ilves I, Tamberg N, Petojevic T, Nogales E, Botchan MR, Berger JM (2011) The structural basis for MCM2-7 helicase activation by GINS and Cdc45. Nat Struct Mol Biol 18:471–477
Cuvier O, Stanojcic S, Lemaitre JM, Mechali M (2008) A topoisomerase II-dependent mechanism for resetting replicons at the S-M-phase transition. Genes Dev 22:860–865
Daigaku Y, Keszthelyi A, Muller CA, Miyabe I, Brooks T, Retkute R, Hubank M, Nieduszynski CA, Carr AM (2015) A global profile of replicative polymerase usage. Nat Struct Mol Biol 22:192–198
Dalgaard JZ, Klar AJ (2000) swi1 and swi3 perform imprinting, pausing, and termination of DNA replication in S. pombe. Cell 102:745–751
Dalgaard JZ, Eydmann T, Koulintchenko M, Sayrac S, Vengrova S, Yamada-Inagawa T (2009) Random and site-specific replication termination. Methods Mol Biol 521:35–53
Deichsel A, Mouysset J, Hoppe T (2009) The ubiquitin-selective chaperone CDC-48/p97, a new player in DNA replication. Cell Cycle 8:185–190
Devbhandari S, Jiang J, Kumar C, Whitehouse I, Remus D (2017) Chromatin constrains the initiation and elongation of DNA replication. Mol Cell 65:131–141
Dewar JM, Budzowska M, Walter JC (2015) The mechanism of DNA replication termination in vertebrates. Nature 525:345–350
Dewar JM, Low E, Mann M, Raschle M, Walter JC (2017) CRL2Lrr1 promotes unloading of the vertebrate replisome from chromatin during replication termination. Genes Dev 31:275–290
Dimude JU, Midgley-Smith SL, Stein M, Rudolph CJ (2016). Replication termination: containing fork fusion-mediated pathologies in Escherichia coli. Genes (Basel) 7
Downes CS, Clarke DJ, Mullinger AM, Gimenez-Abian JF, Creighton AM, Johnson RT (1994) A topoisomerase II-dependent G2 cycle checkpoint in mammalian cells. Nature 372:467–470
Duxin JP, Dewar JM, Yardimci H, Walter JC (2014) Repair of a DNA-protein crosslink by replication-coupled proteolysis. Cell 159:346–357
Evrin C, Clarke P, Zech J, Lurz R, Sun J, Uhle S, Li H, Stillman B, Speck C (2009) A double-hexameric MCM2-7 complex is loaded onto origin DNA during licensing of eukaryotic DNA replication. Proc Natl Acad Sci U S A 106:20240–20245
Fachinetti D, Bermejo R, Cocito A, Minardi S, Katou Y, Kanoh Y, Shirahige K, Azvolinsky A, Zakian VA, Foiani M (2010) Replication termination at eukaryotic chromosomes is mediated by Top2 and occurs at genomic loci containing pausing elements. Mol Cell 39:595–605
Franz A, Orth M, Pirson PA, Sonneville R, Blow JJ, Gartner A, Stemmann O, Hoppe T (2011) CDC-48/p97 coordinates CDT-1 degradation with GINS chromatin dissociation to ensure faithful DNA replication. Mol Cell 44:85–96
Franz A, Pirson PA, Pilger D, Halder S, Achuthankutty D, Kashkar H, Ramadan K, Hoppe T (2016) Chromatin-associated degradation is defined by UBXN-3/FAF1 to safeguard DNA replication fork progression. Nat Commun 7:10612
Fu YV, Yardimci H, Long DT, Ho TV, Guainazzi A, Bermudez VP, Hurwitz J, van Oijen A, Scharer OD, Walter JC (2011) Selective bypass of a lagging strand roadblock by the eukaryotic replicative DNA helicase. Cell 146:931–941
Fullbright G, Rycenga HB, Gruber JD, Long DT (2016) p97 promotes a conserved mechanism of helicase unloading during DNA cross-link repair. Mol Cell Biol 36:2983–2994
Gaggioli V, Le Viet B, Germe T, Hyrien O (2013) DNA topoisomerase II alpha controls replication origin cluster licensing and firing time in Xenopus egg extracts. Nucleic Acids Res 41:7313–7331
Gambus A, Jones RC, Sanchez-Diaz A, Kanemaki M, van Deursen F, Edmondson RD, Labib K (2006) GINS maintains association of Cdc45 with MCM in replisome progression complexes at eukaryotic DNA replication forks. Nat Cell Biol 8:358–366
Gambus A, Khoudoli GA, Jones RC, Blow JJ (2011) MCM2-7 form double hexamers at licensed origins in Xenopus egg extract. J Biol Chem 286:11855–11864
Ganai RA, Zhang XP, Heyer WD, Johansson E (2016) Strand displacement synthesis by yeast DNA polymerase epsilon. Nucleic Acids Res 44:8229–8240
Garg P, Stith CM, Sabouri N, Johansson E, Burgers PM (2004) Idling by DNA polymerase delta maintains a ligatable nick during lagging-strand DNA replication. Genes Dev 18:2764–2773
Gasser R, Koller T, Sogo JM (1996) The stability of nucleosomes at the replication fork. J Mol Biol 258:224–239
Georgescu RE, Langston L, Yao NY, Yurieva O, Zhang D, Finkelstein J, Agarwal T, O'Donnell ME (2014) Mechanism of asymmetric polymerase assembly at the eukaryotic replication fork. Nat Struct Mol Biol 21:664–670
Georgescu RE, Schauer GD, Yao NY, Langston LD, Yurieva O, Zhang D, Finkelstein J, O’Donnell ME (2015) Reconstitution of a eukaryotic replisome reveals suppression mechanisms that define leading/lagging strand operation. Elife 4:e04988
Germe T, Hyrien O (2005) Topoisomerase II-DNA complexes trapped by ICRF-193 perturb chromatin structure. EMBO Rep 6:729–735
Gilbert DM (2010) Cell fate transitions and the replication timing decision point. J Cell Biol 191:899–903
Hanzelmann P, Buchberger A, Schindelin H (2011) Hierarchical binding of cofactors to the AAA ATPase p97. Structure 19:833–843
Hawkins M, Retkute R, Muller CA, Saner N, Tanaka TU, de Moura AP, Nieduszynski CA (2013) High-resolution replication profiles define the stochastic nature of genome replication initiation and termination. Cell Rep 5:1132–1141
Huang C, Li G, Lennarz WJ (2012) Dynamic flexibility of the ATPase p97 is important for its interprotomer motion transmission. Proc Natl Acad Sci U S A 109:9792–9797
Hyrien O (2009) Topological analysis of plasmid DNA replication intermediates using two-dimensional agarose gels. Methods Mol Biol 521:139–167
Ilves I, Petojevic T, Pesavento JJ, Botchan MR (2010) Activation of the MCM2-7 helicase by association with Cdc45 and GINS proteins. Mol Cell 37:247–258
Ivessa AS, Zhou JQ, Zakian VA (2000) The Saccharomyces Pif1p DNA helicase and the highly related Rrm3p have opposite effects on replication fork progression in ribosomal DNA. Cell 100:479–489
Ivessa AS, Lenzmeier BA, Bessler JB, Goudsouzian LK, Schnakenberg SL, Zakian VA (2003) The Saccharomyces cerevisiae helicase Rrm3p facilitates replication past nonhistone protein-DNA complexes. Mol Cell 12:1525–1536
Jagannathan M, Nguyen T, Gallo D, Luthra N, Brown GW, Saridakis V, Frappier L (2014) A role for USP7 in DNA replication. Mol Cell Biol 34:132–145
Johnson RE, Klassen R, Prakash L, Prakash S (2015) A major role of DNA polymerase delta in replication of both the leading and lagging DNA strands. Mol Cell 59:163–175
Kang YH, Galal WC, Farina A, Tappin I, Hurwitz J (2012) Properties of the human Cdc45/Mcm2-7/GINS helicase complex and its action with DNA polymerase epsilon in rolling circle DNA synthesis. Proc Natl Acad Sci U S A 109:6042–6047
Koepp DM, Kile AC, Swaminathan S, Rodriguez-Rivera V (2006) The F-box protein Dia2 regulates DNA replication. Mol Biol Cell 17:1540–1548
Kohler C, Koalick D, Fabricius A, Parplys AC, Borgmann K, Pospiech H, Grosse F (2016) Cdc45 is limiting for replication initiation in humans. Cell Cycle 15:974–985
Kuhne C, Banks L (1998) E3-ubiquitin ligase/E6-AP links multicopy maintenance protein 7 to the ubiquitination pathway by a novel motif, the L2G box. J Biol Chem 273:34302–34309
Langston L, O’Donnell M (2017) Action of CMG with strand-specific DNA blocks supports an internal unwinding mode for the eukaryotic replicative helicase. Elife 6
Langston LD, Zhang D, Yurieva O, Georgescu RE, Finkelstein J, Yao NY, Indiani C, O’Donnell ME (2014) CMG helicase and DNA polymerase epsilon form a functional 15-subunit holoenzyme for eukaryotic leading-strand DNA replication. Proc Natl Acad Sci U S A 111:15390–15395
Lecona E, Fernandez-Capetillo O (2016) A SUMO and ubiquitin code coordinates protein traffic at replication factories. BioEssays 38:1209–1217
Lecona E, Rodriguez-Acebes S, Specks J, Lopez-Contreras AJ, Ruppen I, Murga M, Munoz J, Mendez J, Fernandez-Capetillo O (2016) USP7 is a SUMO deubiquitinase essential for DNA replication. Nat Struct Mol Biol 23:270–277
Lee JJ, Park JK, Jeong J, Jeon H, Yoon JB, Kim EE, Lee KJ (2013) Complex of Fas-associated factor 1 (FAF1) with valosin-containing protein (VCP)-Npl4-Ufd1 and polyubiquitinated proteins promotes endoplasmic reticulum-associated degradation (ERAD). J Biol Chem 288:6998–7011
Li G, Huang C, Zhao G, Lennarz WJ (2012) Interprotomer motion-transmission mechanism for the hexameric AAA ATPase p97. Proc Natl Acad Sci U S A 109:3737–3741
Lopez-Contreras AJ, Ruppen I, Nieto-Soler M, Murga M, Rodriguez-Acebes S, Remeseiro S, Rodrigo-Perez S, Rojas AM, Mendez J, Munoz J et al (2013) A proteomic characterization of factors enriched at nascent DNA molecules. Cell Rep 3:1105–1116
Lucas I, Germe T, Chevrier-Miller M, Hyrien O (2001) Topoisomerase II can unlink replicating DNA by precatenane removal. EMBO J 20:6509–6519
Maculins T, Nkosi PJ, Nishikawa H, Labib K (2015) Tethering of SCF(Dia2) to the replisome promotes efficient ubiquitylation and disassembly of the CMG helicase. Curr Biol 25:2254–2259
Magnaghi P, D'Alessio R, Valsasina B, Avanzi N, Rizzi S, Asa D, Gasparri F, Cozzi L, Cucchi U, Orrenius C et al (2013) Covalent and allosteric inhibitors of the ATPase VCP/p97 induce cancer cell death. Nat Chem Biol 9:548–556
Maric M, Maculins T, De Piccoli G, Labib K (2014) Cdc48 and a ubiquitin ligase drive disassembly of the CMG helicase at the end of DNA replication. Science 346:1253596
McGranahan N, Burrell RA, Endesfelder D, Novelli MR, Swanton C (2012) Cancer chromosomal instability: therapeutic and diagnostic challenges. EMBO Rep 13:528–538
McGuffee SR, Smith DJ, Whitehouse I (2013) Quantitative, genome-wide analysis of eukaryotic replication initiation and termination. Mol Cell 50:123–135
Menges CW, Altomare DA, Testa JR (2009) FAS-associated factor 1 (FAF1): diverse functions and implications for oncogenesis. Cell Cycle 8:2528–2534
Merlet J, Burger J, Tavernier N, Richaudeau B, Gomes JE, Pintard L (2010) The CRL2LRR-1 ubiquitin ligase regulates cell cycle progression during C. elegans development. Development 137:3857–3866
Meyer H, Bug M, Bremer S (2012) Emerging functions of the VCP/p97 AAA-ATPase in the ubiquitin system. Nat Cell Biol 14:117–123
Montagnoli A, Moll J, Colotta F (2010) Targeting cell division cycle 7 kinase: a new approach for cancer therapy. Clin Cancer Res 16:4503–4508
Moreno SP, Bailey R, Campion N, Herron S, Gambus A (2014) Polyubiquitylation drives replisome disassembly at the termination of DNA replication. Science 346:477–481
Moreno A, Carrington JT, Albergante L, Al Mamun M, Haagensen EJ, Komseli ES, Gorgoulis VG, Newman TJ, Blow JJ (2016) Unreplicated DNA remaining from unperturbed S phases passes through mitosis for resolution in daughter cells. Proc Natl Acad Sci U S A 113:E5757–E5764
Morohashi H, Maculins T, Labib K (2009) The amino-terminal TPR domain of Dia2 tethers SCF(Dia2) to the replisome progression complex. Curr Biol 19:1943–1949
Mouysset J, Deichsel A, Moser S, Hoege C, Hyman AA, Gartner A, Hoppe T (2008) Cell cycle progression requires the CDC-48UFD-1/NPL-4 complex for efficient DNA replication. Proc Natl Acad Sci U S A 105:12879–12884
Moyer SE, Lewis PW, Botchan MR (2006) Isolation of the Cdc45/Mcm2-7/GINS (CMG) complex, a candidate for the eukaryotic DNA replication fork helicase. Proc Natl Acad Sci U S A 103:10236–10241
Muller JM, Deinhardt K, Rosewell I, Warren G, Shima DT (2007) Targeted deletion of p97 (VCP/CDC48) in mouse results in early embryonic lethality. Biochem Biophys Res Commun 354:459–465
Muramatsu S, Hirai K, Tak YS, Kamimura Y, Araki H (2010) CDK-dependent complex formation between replication proteins Dpb11, Sld2, Pol (epsilon), and GINS in budding yeast. Genes Dev 24:602–612
Neelsen KJ, Lopes M (2015) Replication fork reversal in eukaryotes: from dead end to dynamic response. Nat Rev Mol Cell Biol 16:207–220
Nishiyama A, Frappier L, Mechali M (2011) MCM-BP regulates unloading of the MCM2-7 helicase in late S phase. Genes Dev 25:165–175
Ossareh-Nazari B, Katsiarimpa A, Merlet J, Pintard L (2016) RNAi-based suppressor screens reveal genetic interactions between the CRL2LRR-1 E3-ligase and the DNA replication Machinery in Caenorhabditis elegans. G3 (Bethesda) 6:3431–3442
Pavlov YI, Frahm C, Nick McElhinny SA, Niimi A, Suzuki M, Kunkel TA (2006) Evidence that errors made by DNA polymerase alpha are corrected by DNA polymerase delta. Curr Biol 16:202–207
Pelisch F, Sonneville R, Pourkarimi E, Agostinho A, Blow JJ, Gartner A, Hay RT (2014) Dynamic SUMO modification regulates mitotic chromosome assembly and cell cycle progression in Caenorhabditis elegans. Nat Commun 5:5485
Pellegrini L, Costa A (2016) New insights into the mechanism of DNA duplication by the eukaryotic replisome. Trends Biochem Sci 41:859–871
Petryk N, Kahli M, d'Aubenton-Carafa Y, Jaszczyszyn Y, Shen Y, Silvain M, Thermes C, Chen CL, Hyrien O (2016) Replication landscape of the human genome. Nat Commun 7:10208
Piano F, Schetter AJ, Morton DG, Gunsalus KC, Reinke V, Kim SK, Kemphues KJ (2002) Gene clustering based on RNAi phenotypes of ovary-enriched genes in C. elegans. Curr Biol 12:1959–1964
Picard F, Cadoret JC, Audit B, Arneodo A, Alberti A, Battail C, Duret L, Prioleau MN (2014) The spatiotemporal program of DNA replication is associated with specific combinations of chromatin marks in human cells. PLoS Genet 10:e1004282
Postow L, Crisona NJ, Peter BJ, Hardy CD, Cozzarelli NR (2001) Topological challenges to DNA replication: conformations at the fork. Proc Natl Acad Sci U S A 98:8219–8226
Reijns MA, Kemp H, Ding J, de Proce SM, Jackson AP, Taylor MS (2015) Lagging-strand replication shapes the mutational landscape of the genome. Nature 518:502–506
Remus D, Beuron F, Tolun G, Griffith JD, Morris EP, Diffley JF (2009) Concerted loading of Mcm2-7 double hexamers around DNA during DNA replication origin licensing. Cell 139:719–730
Reverdy C, Conrath S, Lopez R, Planquette C, Atmanene C, Collura V, Harpon J, Battaglia V, Vivat V, Sippl W et al (2012) Discovery of specific inhibitors of human USP7/HAUSP deubiquitinating enzyme. Chem Biol 19:467–477
Roca J, Ishida R, Berger JM, Andoh T, Wang JC (1994) Antitumor bisdioxopiperazines inhibit yeast DNA topoisomerase II by trapping the enzyme in the form of a closed protein clamp. Proc Natl Acad Sci U S A 91:1781–1785
Rudolph CJ, Upton AL, Stockum A, Nieduszynski CA, Lloyd RG (2013) Avoiding chromosome pathology when replication forks collide. Nature 500:608–611
Schalbetter SA, Mansoubi S, Chambers AL, Downs JA, Baxter J (2015) Fork rotation and DNA precatenation are restricted during DNA replication to prevent chromosomal instability. Proc Natl Acad Sci U S A 112:E4565–E4570
Scott DC, Rhee DY, Duda DM, Kelsall IR, Olszewski JL, Paulo JA, de Jong A, Ovaa H, Alpi AF, Harper JW et al (2016) Two distinct types of E3 ligases work in unison to regulate substrate ubiquitylation. Cell 166(1198–1214):e1124
Sengupta S, van Deursen F, de Piccoli G, Labib K (2013) Dpb2 integrates the leading-strand DNA polymerase into the eukaryotic replisome. Curr Biol 23:543–552
Simon AC, Sannino V, Costanzo V, Pellegrini L (2016) Structure of human Cdc45 and implications for CMG helicase function. Nat Commun 7:11638
Skoufias DA, Lacroix FB, Andreassen PR, Wilson L, Margolis RL (2004) Inhibition of DNA decatenation, but not DNA damage, arrests cells at metaphase. Mol Cell 15:977–990
Smith DJ, Whitehouse I (2012) Intrinsic coupling of lagging-strand synthesis to chromatin assembly. Nature 483:434–438
Sonneville R, Moreno SP, Knebel A, Johnson C, Hastie CJ, Gartner A, Gambus A, Labib K (2017) CUL-2LRR-1 and UBXN-3 drive replisome disassembly during DNA replication termination and mitosis. Nat Cell Biol 19(5):468–479
Starostina NG, Simpliciano JM, McGuirk MA, Kipreos ET (2010) CRL2(LRR-1) targets a CDK inhibitor for cell cycle control in C. elegans and actin-based motility regulation in human cells. Dev Cell 19:753–764
Sun J, Shi Y, Georgescu RE, Yuan Z, Chait BT, Li H, O’Donnell ME (2015) The architecture of a eukaryotic replisome. Nat Struct Mol Biol 22:976–982
Sundin O, Varshavsky A (1980) Terminal stages of SV40 DNA replication proceed via multiply intertwined catenated dimers. Cell 21:103–114
Tonddast-Navaei S, Stan G (2013) Mechanism of transient binding and release of substrate protein during the allosteric cycle of the p97 nanomachine. J Am Chem Soc 135:14627–14636
Uhlmann F (2009) A matter of choice: the establishment of sister chromatid cohesion. EMBO Rep 10:1095–1102
Waga S, Masuda T, Takisawa H, Sugino A (2001) DNA polymerase epsilon is required for coordinated and efficient chromosomal DNA replication in Xenopus egg extracts. Proc Natl Acad Sci U S A 98:4978–4983
Wang JC (2002) Cellular roles of DNA topoisomerases: a molecular perspective. Nat Rev Mol Cell Biol 3:430–440
Wei L, Zhao X (2016) A new MCM modification cycle regulates DNA replication initiation. Nat Struct Mol Biol 23:209–216
Wendel BM, Courcelle CT, Courcelle J (2014) Completion of DNA replication in Escherichia coli. Proc Natl Acad Sci U S A 111:16454–16459
Wojcik C, Yano M, DeMartino GN (2004) RNA interference of valosin-containing protein (VCP/p97) reveals multiple cellular roles linked to ubiquitin/proteasome-dependent proteolysis. J Cell Sci 117:281–292
Wong PG, Winter SL, Zaika E, Cao TV, Oguz U, Koomen JM, Hamlin JL, Alexandrow MG (2011) Cdc45 limits replicon usage from a low density of preRCs in mammalian cells. PLoS One 6:e17533
Yardimci H, Loveland AB, Habuchi S, van Oijen AM, Walter JC (2010) Uncoupling of sister replisomes during eukaryotic DNA replication. Mol Cell 40:834–840
Yeeles JT, Janska A, Early A, Diffley JF (2017) How the eukaryotic replisome achieves rapid and efficient DNA replication. Mol Cell 65:105–116
Yeung HO, Kloppsteck P, Niwa H, Isaacson RL, Matthews S, Zhang X, Freemont PS (2008) Insights into adaptor binding to the AAA protein p97. Biochem Soc Trans 36:62–67
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2017 Springer Nature Singapore Pte Ltd.
About this chapter
Cite this chapter
Gambus, A. (2017). Termination of Eukaryotic Replication Forks. In: Masai, H., Foiani, M. (eds) DNA Replication. Advances in Experimental Medicine and Biology, vol 1042. Springer, Singapore. https://doi.org/10.1007/978-981-10-6955-0_8
Download citation
DOI: https://doi.org/10.1007/978-981-10-6955-0_8
Published:
Publisher Name: Springer, Singapore
Print ISBN: 978-981-10-6954-3
Online ISBN: 978-981-10-6955-0
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)