Abstract
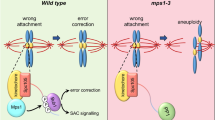
The cell cycle culminates in mitosis with the purpose of dividing the cell’s DNA content equally over two daughter cells. Error-free segregation relies on correct connections between chromosomes and spindle microtubules. Kinetochores are complex multi-protein assemblies that mediate these connections and are the platforms for attachment-error-correction and spindle assembly checkpoint signaling. Proper kinetochore function is therefore key in preventing aneuploidization. Mutations in genes encoding kinetochore proteins are associated with several severe developmental disorders associated with microcephaly, and kinetochore defects contribute to chromosomal instability in certain cancers. This chapter gives an overview of the processes necessary for faithful chromosome segregation and how kinetochore malfunction causes various human pathologies.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Siegel JJ, Amon A (2012) New insights into the troubles of aneuploidy. Annu Rev Cell Dev Biol 28:189–214
Ricke RM, van Deursen JM (2013) Aneuploidy in health, disease, and aging. J Cell Biol 201(1):11–21
Cheeseman IM (2014) The kinetochore. Cold Spring Harb Perspect Biol 6(7):a015826
Wendell KL, Wilson L, Jordan MA (1993) Mitotic block in HeLa cells by vinblastine: ultrastructural changes in kinetochore-microtubule attachment and in centrosomes. J Cell Sci 104(2):261–274
Carmena M, Wheelock M, Funabiki H, Earnshaw WC (2012) The chromosomal passenger complex (CPC): from easy rider to the godfather of mitosis. Nat Rev Mol Cell Biol 13(12):789–803
Primorac I, Musacchio A (2013) Panta rhei: the APC/C at steady state. J Cell Biol 201(2):177–189
Sacristan C, Kops GJPL (2015) Joined at the hip: kinetochores, microtubules, and spindle assembly checkpoint signaling. Trends Cell Biol 25(1):21–28
London N, Biggins S (2014) Signalling dynamics in the spindle checkpoint response. Nat Rev Mol Cell Biol 15(11):736–747
McKinley KL, Cheeseman IM (2016) The molecular basis for centromere identity and function. Nat Rev Mol Cell Biol 17(1):16–29
Varma D, Salmon ED (2012) The KMN protein network–chief conductors of the kinetochore orchestra. J Cell Sci 125(Pt 24):5927–5936
Caldas GV, DeLuca JG (2014) KNL1: bringing order to the kinetochore. Chromosoma 123(3):169–181
Hiruma Y, Sacristan C, Pachis ST, Adamopoulos A, Kuijt T, Ubbink M, von Castelmur E, Perrakis A, Kops GJPL (2015) CELL DIVISION CYCLE. Competition between MPS1 and microtubules at kinetochores regulates spindle checkpoint signaling. Science 348(6240):1264–1267
Ji Z, Gao H, Yu H, Cayirlioglu P, Kadow IG, Vosshall LB, Stornetta RL, West GH, Guyenet PG, Isacson O, Nishiyama A, Mulkey DK, Koschnitzky JE (2015) CELL DIVISION CYCLE. Kinetochore attachment sensed by competitive Mps1 and microtubule binding to Ndc80C. Science 348(6240):1260–1264
Vleugel M, Hoek T, Tromer E, Sliedrecht T, Groenewold V, Omerzu M, Kops GJPL (2015) Dissecting the roles of human BUB1 in the spindle assembly checkpoint. J Cell Sci 128:2975–2982
Zhang G, Lischetti T, Hayward DG, Nilsson J (2015) Distinct domains in Bub1 localize RZZ and BubR1 to kinetochores to regulate the checkpoint. Nat Commun 6:7162
Johnson VL, Scott MIF, Holt SV, Hussein D, Taylor SS (2004) Bub1 is required for kinetochore localization of BubR1, Cenp-E, Cenp-F and Mad2, and chromosome congression. J Cell Sci 117(Pt 8):1577–1589
Holt SV, Vergnolle MA, Hussein D, Wozniak MJ, Allan VJ, Taylor SS (2005) Silencing Cenp-F weakens centromeric cohesion, prevents chromosome alignment and activates the spindle checkpoint. J Cell Sci 118(Pt 20):4889–4900
Suijkerbuijk SJE, Vleugel M, Teixeira A, Kops GJPL (2012) Integration of kinase and phosphatase activities by BUBR1 ensures formation of stable kinetochore-microtubule attachments. Dev Cell 23(4):745–755
Akera T, Goto Y, Sato M, Yamamoto M, Watanabe Y (2015) Mad1 promotes chromosome congression by anchoring a kinesin motor to the kinetochore. Nat Cell Biol 17(9):1124–1133
Kawashima SA, Yamagishi Y, Honda T, Ishiguro K, Watanabe Y (2010) Phosphorylation of H2A by Bub1 prevents chromosomal instability through localizing shugoshin. Science 327(5962):172–177
Yamagishi Y, Honda T, Tanno Y, Watanabe Y (2010) Two histone marks establish the inner centromere and chromosome bi-orientation. Science 330(6001):239–243
Kops GJPL, Shah JV (2012) Connecting up and clearing out: how kinetochore attachment silences the spindle assembly checkpoint. Chromosoma 121(5):509–525
Funabiki H, Wynne DJ (2013) Making an effective switch at the kinetochore by phosphorylation and dephosphorylation. Chromosoma 122(3):135–158
Foley EA, Kapoor TM (2013) Microtubule attachment and spindle assembly checkpoint signalling at the kinetochore. Nat Rev Mol Cell Biol 14(1):25–37
Etemad B, Kops GJPL (2016) Attachment issues: kinetochore transformations and spindle checkpoint silencing. Curr Opin Cell Biol 39:101–108
Kops GJPL, Saurin AT, Meraldi P (2010) Finding the middle ground: how kinetochores power chromosome congression. Cell Mol Life Sci 67(13):2145–2161
Yao X, Abrieu A, Zheng Y, Sullivan KF, Cleveland DW (2000) CENP-E forms a link between attachment of spindle microtubules to kinetochores and the mitotic checkpoint. Nat Cell Biol 2:484–491
Bader JR, Vaughan KT (2010) Dynein at the kinetochore: timing, interactions and functions. Semin Cell Dev Biol 21(3):269–275
Thompson SL, Bakhoum SF, Compton DA (2010) Mechanisms of chromosomal instability. Curr Biol 20(6):R285
Al Alwan I, Khadora M, Amir I, Nasrat G, Omair A, Brown L, Al Dubayee M, Badri M (2014) Turner syndrome genotype and phenotype and their effect on presenting features and timing of diagnosis. Int J Health Sci (Qassim) 8(2):195–202
Knoch J, Kamenisch Y, Kubisch C, Berneburg M (2012) Rare hereditary diseases with defects in DNA-repair. Eur J Dermatol 22(4):443–455
Cucco F, Musio A (2016) Genome stability: what we have learned from cohesinopathies. Am J Med Genet C Semin Med Genet 172:171
Zakari M, Yuen K, Gerton JL (2015) Etiology and pathogenesis of the cohesinopathies. Wiley Interdiscip Rev Dev Biol 4:489
Thompson SL, Compton DA (2010) Proliferation of aneuploid human cells is limited by a p53-dependent mechanism. J Cell Biol 188(3):369–381
Janssen A, van der Burg M, Szuhai K, Kops GJPL, Medema RH (2011) Chromosome segregation errors as a cause of DNA damage and structural chromosome aberrations. Science 333(6051):1895–1898
Li M, Fang X, Baker DJ, Guo L, Gao X, Wei Z, Han S, van Deursen JM, Zhang P (2010) The ATM-p53 pathway suppresses aneuploidy-induced tumorigenesis. Proc Natl Acad Sci U S A 107(32):14188–14193
Stingele S, Stoehr G, Peplowska K, Cox J, Mann M, Storchova Z (2012) Global analysis of genome, transcriptome and proteome reveals the response to aneuploidy in human cells. Mol Syst Biol 8:608
Upender MB, Habermann JK, McShane LM, Korn EL, Barrett JC, Difilippantonio MJ, Ried T (2004) Chromosome transfer induced aneuploidy results in complex dysregulation of the cellular transcriptome in immortalized and cancer cells. Cancer Res 64(19):6941–6949
Dürrbaum M, Storchová Z (2015) Effects of aneuploidy on gene expression: implications for cancer. FEBS J 283:791–802
Dürrbaum M, Kuznetsova AY, Passerini V, Stingele S, Stoehr G, Storchová Z (2014) Unique features of the transcriptional response to model aneuploidy in human cells. BMC Genomics 15(1):139
Santaguida S, Amon A (2015) Short- and long-term effects of chromosome mis-segregation and aneuploidy. Nat Rev Mol Cell Biol 16(8):473–485
Crasta K, Ganem NJ, Dagher R, Lantermann AB, Ivanova EV, Pan Y, Nezi L, Protopopov A, Chowdhury D, Pellman D (2012) DNA breaks and chromosome pulverization from errors in mitosis. Nature 482(7383):53–58
Storchová Z, Kloosterman WP (2016) The genomic characteristics and cellular origin of chromothripsis. Curr Opin Cell Biol 40:106–113
Zhang C-Z, Spektor A, Cornils H, Francis JM, Jackson EK, Liu S, Meyerson M, Pellman D (2015) Chromothripsis from DNA damage in micronuclei. Nature 522(7555):179–184
De Pagter MS, Van Roosmalen MJ, Baas AF, Renkens I, Duran KJ, Van Binsbergen E, Tavakoli-Yaraki M, Hochstenbach R, Van Der Veken LT, Cuppen E, Kloosterman WP (2015) Chromothripsis in healthy individuals affects multiple protein-coding genes and can result in severe congenital abnormalities in offspring. Am J Hum Genet 96(4):651–656
Kloosterman WP, Guryev V, van Roosmalen M, Duran KJ, de Bruijn E, Bakker SCM, Letteboer T, van Nesselrooij B, Hochstenbach R, Poot M, Cuppen E (2011) Chromothripsis as a mechanism driving complex de novo structural rearrangements in the germline. Hum Mol Genet 20(10):1916–1924
Knouse KA, Wu J, Whittaker CA, Amon A (2014) Single cell sequencing reveals low levels of aneuploidy across mammalian tissues. Proc Natl Acad Sci U S A 111(37):13409–13414
McConnell MJ, Lindberg M, Brennand KJ, Piper JC, Voet T, Cowing-Zitron C, Shumilina S, Lasken RS, Vermeesch JR, Hall IM, Gage FH (2013) Mosaic copy number variation in human neurons. Science 342:632–638
Pfau SJ, Silberman RE, Knouse KA, Amon A (2016) Aneuploidy impairs hematopoietic stem cell fitness and is selected against in regenerating tissues in vivo. Genes Dev 30:1395–1408
Marthiens V, Rujano MA, Pennetier C, Tessier S, Paul-Gilloteaux P, Basto R (2013) Centrosome amplification causes microcephaly. Nat Cell Biol 15(7):731–740
Woods CG, Basto R (2014) Microcephaly. Curr Biol 24(23):R1109–R1111
Gogendeau D, Siudeja K, Gambarotto D, Pennetier C, Bardin AJ, Basto R (2015) Aneuploidy causes premature differentiation of neural and intestinal stem cells. Nat Commun 6:8894
Chetaille P, Preuss C, Burkhard S, Côté J-M, Houde C, Castilloux J, Piché J, Gosset N, Leclerc S, Wünnemann F, Thibeault M, Gagnon C, Galli A, Tuck E, Hickson GR, El Amine N, Boufaied I, Lemyre E, de Santa Barbara P, Faure S, Jonzon A, Cameron M, Dietz HC, Gallo-McFarlane E, Benson DW, Moreau C, Labuda D, Zhan SH, Shen Y, Jomphe M, Jones SJM, Bakkers J, Andelfinger G (2014) Mutations in SGOL1 cause a novel cohesinopathy affecting heart and gut rhythm. Nat Genet 46(11):1245–1249
Silió V, McAinsh AD, Millar JB (2015) KNL1-Bubs and RZZ provide two separable pathways for checkpoint activation at human kinetochores. Dev Cell 35(5):600–613
Kiyomitsu T, Obuse C, Yanagida M (2007) Human Blinkin/AF15q14 is required for chromosome alignment and the mitotic checkpoint through direct interaction with Bub1 and BubR1. Dev Cell 13(5):663–676
Vleugel M, Tromer E, Omerzu M, Groenewold V, Nijenhuis W, Snel B, Kops GJPL (2013) Arrayed BUB recruitment modules in the kinetochore scaffold KNL1 promote accurate chromosome segregation. J Cell Biol 203(6):943–955
Genin A, Desir J, Lambert N, Biervliet M, Van der Aa N, Pierquin G, Killian A, Tosi M, Urbina M, Lefort A, Libert F, Pirson I, Abramowicz M (2012) Kinetochore KMN network gene CASC5 mutated in primary microcephaly. Hum Mol Genet 21(24):5306–5317
Szczepanski S, Hussain MS, Sur I, Altmüller J, Thiele H, Abdullah U, Waseem SS, Moawia A, Nürnberg G, Noegel AA, Baig SM, Nürnberg P (2016) A novel homozygous splicing mutation of CASC5 causes primary microcephaly in a large Pakistani family. Hum Genet 135(2):157–170
Saadi A, Verny F, Siquier-Pernet K, Bole-Feysot C, Nitschke P, Munnich A, Abada-Dendib M, Chaouch M, Abramowicz M, Colleaux L (2016) Refining the phenotype associated with CASC5 mutation. Neurogenetics 17(1):71–78
Mirzaa GM, Vitre B, Carpenter G, Abramowicz I, Gleeson JG, Paciorkowski AR, Cleveland DW, Dobyns WB, O’Driscoll M (2014) Mutations in CENPE define a novel kinetochore-centromeric mechanism for microcephalic primordial dwarfism. Hum Genet 133(8):1023–1039
Kapoor TM, Lampson MA, Hergert P, Cameron L, Cimini D, Salmon ED, McEwen BF, Khodjakov A (2006) Chromosomes can congress to the metaphase plate before biorientation. Science 311(5759):388–391
Yen TJ, Li G, Schaar BT, Szilak I, Cleveland DW (1992) CENP-E is a putative kinetochore motor that accumulates just before mitosis. Nature 359:536–539
Powles-Glover N (2014) Cilia and ciliopathies: classic examples linking phenotype and genotype-an overview. Reprod Toxicol 48:98–105
Waters AM, Asfahani R, Carroll P, Bicknell L, Lescai F, Bright A, Chanudet E, Brooks A, Christou-Savina S, Osman G, Walsh P, Bacchelli C, Chapgier A, Vernay B, Bader DM, Deshpande C, Sullivan MO’, Ocaka L, Stanescu H, Stewart HS, Hildebrandt F, Otto E, Johnson CA, Szymanska K, Katsanis N, Davis E, Kleta R, Hubank M, Doxsey S, Jackson A, Stupka E, Winey M, Beales PL (2015) The kinetochore protein, CENPF, is mutated in human ciliopathy and microcephaly phenotypes. J Med Genet 52(3):147–156
Filges I, Bruder E, Brandal K, Meier S, Undlien DE, Waage TR, Hoesli I, Schubach M, de Beer T, Sheng Y, Hoeller S, Schulzke S, Røsby O, Miny P, Tercanli S, Oppedal T, Meyer P, Selmer KK, Strømme P (2016) Strømme syndrome is a ciliary disorder caused by mutations in CENPF. Hum Mutat 37(4):359–363
Kanfer G, Courthéoux T, Peterka M, Meier S, Soste M, Melnik A, Reis K, Aspenström P, Peter M, Picotti P, Kornmann B (2015) Mitotic redistribution of the mitochondrial network by Miro and Cenp-F. Nat Commun 6:8015
Volkov VA, Grissom PM, Arzhanik VK, Zaytsev AV, Renganathan K, McClure-Begley T, Old WM, Ahn N, Richard McIntosh J (2015) Centromere protein F includes two sites that couple efficiently to depolymerizing microtubules. J Cell Biol 209(6):813–828
Vergnolle MAS, Taylor SS (2007) Cenp-F links kinetochores to Ndel1/Nde1/Lis1/dynein microtubule motor complexes. Curr Biol 17(13):1173–1179
Kim S, Zaghloul NA, Bubenshchikova E, Oh EC, Rankin S, Katsanis N, Obara T, Tsiokas L (2011) Nde1-mediated inhibition of ciliogenesis affects cell cycle re-entry. Nat Cell Biol 13(4):351–360
Bakircioglu M, Carvalho OP, Khurshid M, Cox JJ, Tuysuz B, Barak T, Yilmaz S, Caglayan O, Dincer A, Nicholas AK, Quarrell O, Springell K, Karbani G, Malik S, Gannon C, Sheridan E, Crosier M, Lisgo SN, Lindsay S, Bilguvar K, Gergely F, Gunel M, Woods CG (2011) The essential role of centrosomal NDE1 in human cerebral cortex neurogenesis. Am J Hum Genet 88(5):523–535
Guven A, Gunduz A, Bozoglu TM, Yalcinkaya C, Tolun A (2012) Novel NDE1 homozygous mutation resulting in microhydranencephaly and not microlyssencephaly. Neurogenetics 13(3):189–194
Alkuraya FS, Cai X, Emery C, Mochida GH, Al-Dosari MS, Felie JM, Hill RS, Barry BJ, Partlow JN, Gascon GG, Kentab A, Jan M, Shaheen R, Feng Y, Walsh CA (2011) Human mutations in NDE1 cause extreme microcephaly with lissencephaly. Am J Hum Genet 88(5):536–547
Dantas TJ, Carabalona A, Jun-Kit Hu D, Vallee RB (2016) Emerging roles for motor proteins in progenitor cell behavior and neuronal migration during brain development. Cytoskeleton 73:566–576
Isidor B, Küry S, Rosenfeld JA, Besnard T, Schmitt S, Joss S, Davies SJ, Roger Lebel R, Henderson A, Schaaf CP, Streff HE, Yang Y, Jain V, Chida N, Latypova X, Le Caignec C, Cogné B, Mercier S, Vincent M, Colin E, Bonneau D, Denommé AS, Parent P, Gilbert-Dussardier B, Odent S, Toutain A, Piton A, Dina C, Donnart A, Lindenbaum P, Charpentier E, Redon R, Iemura K, Ikeda M, Tanaka K, Bézieau S (2016) De novo truncating mutations in the kinetochore-microtubules attachment gene CHAMP1 cause syndromic intellectual disability. Hum Mutat 37(4):354–358
Tanaka AJ, Cho MT, Retterer K, Jones JR, Nowak C, Douglas J, Jiang Y-H, McConkie-Rosell A, Schaefer GB, Kaylor J, Rahman OA, Telegrafi A, Friedman B, Douglas G, Monaghan KG, Chung WK (2016) De novo pathogenic variants in CHAMP1 are associated with global developmental delay, intellectual disability, and dysmorphic facial features. Mol Case Stud 2(1):a000661
Hempel M, Cremer K, Ockeloen CW, Lichtenbelt KD, Herkert JC, Denecke J, Haack TB, Zink AM, Becker J, Wohlleber E, Johannsen J, Alhaddad B, Pfundt R, Fuchs S, Wieczorek D, Strom TM, Van Gassen KLI, Kleefstra T, Kubisch C, Engels H, Lessel D (2015) De Novo mutations in CHAMP1 cause intellectual disability with severe speech impairment. Am J Hum Genet 97(3):493–500
Itoh G, Kanno S, Uchida KSK, Chiba S, Sugino S, Watanabe K, Mizuno K, Yasui A, Hirota T, Tanaka K (2011) CAMP (C13orf8, ZNF828) is a novel regulator of kinetochore-microtubule attachment. EMBO J 30(1):130–144
Nozawa R-S, Nagao K, Masuda H-T, Iwasaki O, Hirota T, Nozaki N, Kimura H, Obuse C (2010) Human POGZ modulates dissociation of HP1alpha from mitotic chromosome arms through Aurora B activation. Nat Cell Biol 12(7):719–727
Tanno Y, Susumu H, Kawamura M, Sugimura H, Honda T, Watanabe Y (2015) The inner centromere-shugoshin network prevents chromosomal instability. Science 349(6253):1237–1240
Ye Y, Cho MT, Retterer K, Alexander N, Ben-Omran T, Al-Mureikhi M, Cristian I, Wheeler PG, Crain C, Zand D, Weinstein V, Vernon HJ, McClellan R, Krishnamurthy V, Vitazka P, Millan F, Chung WK (2015) De novo POGZ mutations are associated with neurodevelopmental disorders and microcephaly. Mol Case Stud 1(1):a000455
Jacquemont S, Bocéno M, Rival JM, Méchinaud F, David A (2002) High risk of malignancy in mosaic variegated aneuploidy syndrome. Am J Med Genet 109(1):17–21
García-Castillo H, Vásquez-Velásquez AI, Rivera H, Barros-Núñez P (2008) Clinical and genetic heterogeneity in patients with mosaic variegated aneuploidy: delineation of clinical subtypes. Am J Med Genet A 146A(13):1687–1695
Hanks S, Coleman K, Reid S, Plaja A, Firth H, Fitzpatrick D, Kidd A, Méhes K, Nash R, Robin N, Shannon N, Tolmie J, Swansbury J, Irrthum A, Douglas J, Rahman N (2004) Constitutional aneuploidy and cancer predisposition caused by biallelic mutations in BUB1B. Nat Genet 36(11):1159–1161
Snape K, Hanks S, Ruark E, Barros-Núñez P, Elliott A, Murray A, Lane AH, Shannon N, Callier P, Chitayat D, Clayton-Smith J, Fitzpatrick DR, Gisselsson D, Jacquemont S, Asakura-Hay K, Micale MA, Tolmie J, Turnpenny PD, Wright M, Douglas J, Rahman N (2011) Mutations in CEP57 cause mosaic variegated aneuploidy syndrome. Nat Genet 43(6):527–529
Miyamoto T, Porazinski S, Wang H, Borovina A, Ciruna B, Shimizu A, Kajii T, Kikuchi A, Furutani-Seiki M, Matsuura S (2011) Insufficiency of BUBR1, a mitotic spindle checkpoint regulator, causes impaired ciliogenesis in vertebrates. Hum Mol Genet 20(10):2058–2070
Baker DJ, Jeganathan KB, Cameron JD, Thompson M, Juneja S, Kopecka A, Kumar R, Jenkins RB, de Groen PC, Roche P, van Deursen JM (2004) BubR1 insufficiency causes early onset of aging-associated phenotypes and infertility in mice. Nat Genet 36(7):744–749
Suijkerbuijk SJE, Vleugel M, Teixeira A, Kops GJPL (2012) Integration of kinase and phosphatase activities by BUBR1 ensures formation of stable kinetochore-microtubule attachments. Dev Cell 23(4):745–755
Izumi H, Matsumoto Y, Ikeuchi T, Saya H, Kajii T, Matsuura S (2009) BubR1 localizes to centrosomes and suppresses centrosome amplification via regulating Plk1 activity in interphase cells. Oncogene 28(31):2806–2820
Suijkerbuijk SJE, van Osch MHJ, Bos FL, Hanks S, Rahman N, Kops GJPL (2010) Molecular causes for BUBR1 dysfunction in the human cancer predisposition syndrome mosaic variegated aneuploidy. Cancer Res 70(12):4891–4900
Bossard C, Laurell H, Van den Berghe L, Meunier S, Zanibellato C, Prats H (2003) Translokin is an intracellular mediator of FGF-2 trafficking. Nat Cell Biol 5:433–439
Momotani K, Khromov AS, Miyake T, Stukenberg PT, Somlyo AV (2008) Cep57, a multidomain protein with unique microtubule and centrosomal localization domains. Biochem J 412(2):265–273
Zhou H, Wang T, Zheng T, Teng J, Chen J (2016) Cep57 is a Mis12-interacting kinetochore protein involved in kinetochore targeting of Mad1–Mad2. Nat Commun 7:10151
Snape K, Hanks S, Ruark E, Barros-Núñez P, Elliott A, Murray A, Lane AH, Shannon N, Callier P, Chitayat D, Clayton-Smith J, Fitzpatrick DR, Gisselsson D, Jacquemont S, Asakura-Hay K, Micale MA, Tolmie J, Turnpenny PD, Wright M, Douglas J, Rahman N (2011) Mutations in CEP57 cause mosaic variegated aneuploidy syndrome. Nat Genet 43(6):527–529
Beroukhim R, Mermel C, Porter D (2010) The landscape of somatic copy-number alteration across human cancers. Nature 463:899–905
Mitelman F, Johansson B, Mertens F (2016) Mitelman Database of Chromosome Aberrations and Gene Fusions in Cancer. [Online]. Available: http://cgap.nci.nih.gov/Chromosomes/Mitelman
Duijf PHG, Benezra R (2013) The cancer biology of whole-chromosome instability. Oncogene 32(40):4727–4736
Burrell RA, McClelland SE, Endesfelder D, Groth P, Weller M-C, Shaikh N, Domingo E, Kanu N, Dewhurst SM, Gronroos E, Chew SK, Rowan AJ, Schenk A, Sheffer M, Howell M, Kschischo M, Behrens A, Helleday T, Bartek J, Tomlinson IP, Swanton C (2013) Replication stress links structural and numerical cancer chromosomal instability. Nature 494(7438):492–496
Fukasawa K (2005) Centrosome amplification, chromosome instability and cancer development. Cancer Lett 230(1):6–19
Bakhoum SF, Thompson SL, Manning AL, Compton DA (2009) Genome stability is ensured by temporal control of kinetochore-microtubule dynamics. Nat Cell Biol 11(1):27–35
Kops GJPL, Weaver BAA, Cleveland DW (2005) On the road to cancer: aneuploidy and the mitotic checkpoint. Nat Rev Cancer 5(10):773–785
Bakhoum SF, Genovese G, Compton DA (2009) Deviant kinetochore microtubule dynamics underlie chromosomal instability. Curr Biol 19(22):1937–1942
Gordon DJ, Resio B, Pellman D (2012) Causes and consequences of aneuploidy in cancer. Nat Rev Genet 13(3):189–203
Bastians H (2015) Causes of chromosomal instability. In: Ghadimi B, Ried T (eds) Chromosomal instability in cancer cells, vol 200. Springer, Cham, pp 95–113
Simon JE, Bakker B, Foijer F (2015) CINcere modelling: what have mouse models for chromosome instability taught us? In: Chromosomal instability in cancer cells. Springer, Cham, pp 39–60
Tighe A, Johnson VL, Albertella M, Taylor SS (2001) Aneuploid colon cancer cells have a robust spindle checkpoint. EMBO Rep 2(7):609–614
Kim HS, Kyung HP, Kim SA, Wen J, Seung WP, Park B, Gham CW, Woo JH, Sung HN, Ho KK, Song SY (2005) Frequent mutations of human Mad2, but not Bub1, in gastric cancers cause defective mitotic spindle checkpoint. Mutat Res Fundam Mol Mech Mutagen 578(1–2):187–201
Suijkerbuijk SJE, Kops GJPL (2008) Preventing aneuploidy: the contribution of mitotic checkpoint proteins. Biochim Biophys Acta Rev Cancer 1786(1):24–31
Thiru P, Kern DM, McKinley KL, Monda JK, Rago F, Su K-C, Tsinman T, Yarar D, Bell GW, Cheeseman IM (2014) Kinetochore genes are coordinately up-regulated in human tumors as part of a FoxM1-related cell division program. Mol Biol Cell 25(13):1983–1994
Trivedi P, Stukenberg PT (2016) A centromere-signaling network underlies the coordination among mitotic events. Trends Biochem Sci 41(2):160–174
Hill VK, Kim J-S, Waldman T (2016) Cohesin mutations in human cancer. Biochim Biophys Acta Rev Cancer 1866(1):1–11
Solomon DA, Kim T, Diaz-Martinez LA, Fair J, Elkahloun AG, Harris BT, Toretsky JA, Rosenberg SA, Shukla N, Ladanyi M, Samuels Y, James CD, Yu H, Kim J-S, Waldman T (2011) Mutational inactivation of STAG2 causes aneuploidy in human cancer. Science 333(6045):1039–1043
Lawrence MS, Stojanov P, Mermel CH, Robinson JT, Garraway LA, Golub TR, Meyerson M, Gabriel SB, Lander ES, Getz G (2014) Discovery and saturation analysis of cancer genes across 21 tumour types. Nature 505(7484):495–501
Bakhoum SF, Silkworth WT, Nardi IK, Nicholson JM, Compton DA, Cimini D (2014) The mitotic origin of chromosomal instability. Curr Biol 24(4):R148
Kabeche L, Compton DA (2012) Checkpoint-independent stabilization of kinetochore-microtubule attachments by Mad2 in human cells. Curr Biol 22(7):638–644
Silkworth WT, Nardi IK, Scholl LM, Cimini D (2009) Multipolar spindle pole coalescence is a major source of kinetochore mis-attachment and chromosome mis-segregation in cancer cells. PLoS One 4(8):e6564
Ganem NJ, Godinho SA, Pellman D (2009) A mechanism linking extra centrosomes to chromosomal instability. Nature 460(7252):278–282
Ertych N, Stolz A, Stenzinger A, Weichert W, Kaulfuß S, Burfeind P, Aigner A, Wordeman L, Bastians H (2014) Increased microtubule assembly rates influence chromosomal instability in colorectal cancer cells. Nat Cell Biol 16(8):779–791
Wooster R, Bignell G, Lancaster J, Swift S, Seal S, Mangion J, Collins N, Gregory S, Gumbs C, Micklem G (1995) Identification of the breast cancer susceptibility gene BRCA2. Nature 378(6559):789–792
T. Breast Cancer Linkage Consortium (1999) Cancer risks in BRCA2 mutation carriers. JNCI J Natl Cancer Inst 91(15):1310–1316
Choi E, Park PG, Lee HO, Lee YK, Kang GH, Lee JW, Han W, Lee HC, Noh DY, Lekomtsev S, Lee H (2012) BRCA2 fine-tunes the spindle assembly checkpoint through reinforcement of BubR1 acetylation. Dev Cell 22(2):295–308
Park I, Lee HO, Choi E, Lee YK, Kwon MS, Min J, Park PG, Lee S, Kong YY, Gong G, Lee H (2013) Loss of BubR1 acetylation causes defects in spindle assembly checkpoint signaling and promotes tumor formation. J Cell Biol 202(2):295–309
Hernando E, Nahlé Z, Juan G, Diaz-Rodriguez E, Alaminos M, Hemann M, Michel L, Mittal V, Gerald W, Benezra R, Lowe SW, Cordon-Cardo C (2004) Rb inactivation promotes genomic instability by uncoupling cell cycle progression from mitotic control. Nature 430(7001):797–802
Sotillo R, Schvartzman J-M, Socci ND, Benezra R (2010) Mad2-induced chromosome instability leads to lung tumour relapse after oncogene withdrawal. Nature 464(7287):436–440
Schvartzman JM, Duijf PHG, Sotillo R, Coker C, Benezra R (2011) Mad2 is a critical mediator of the chromosome instability observed upon Rb and p53 pathway inhibition. Cancer Cell 19(6):701–714
Guardavaccaro D, Frescas D, Dorrello NV, Peschiaroli A, Multani AS, Cardozo T, Lasorella A, Iavarone A, Chang S, Hernando E, Pagano M (2008) Control of chromosome stability by the beta-TrCP-REST-Mad2 axis. Nature 452(7185):365–369
Caldwell CM, Kaplan KB (2009) The role of APC in mitosis and in chromosome instability. Adv Exp Med Biol 656:51–64
Drost J, van Jaarsveld RH, Ponsioen B, Zimberlin C, van Boxtel R, Buijs A, Sachs N, Overmeer RM, Offerhaus GJ, Begthel H, Korving J, van de Wetering M, Schwank G, Logtenberg M, Cuppen E, Snippert HJ, Medema JP, Kops GJPL, Clevers H (2015) Sequential cancer mutations in cultured human intestinal stem cells. Nature 521(7550):43–47
Morris-Rosendahl DJ, Kaindl AM (2015) What next-generation sequencing (NGS) technology has enabled us to learn about primary autosomal recessive microcephaly (MCPH). Mol Cell Probes 29(5):271–281
Childs BG, Durik M, Baker DJ, van Deursen JM (2015) Cellular senescence in aging and age-related disease: from mechanisms to therapy. Nat Med 21(12):1424–1435
Foijer F, Draviam VM, Sorger PK (2008) Studying chromosome instability in the mouse. Biochim Biophys Acta Rev Cancer 1786(1):73–82
Jamieson CR, Govaerts C, Abramowicz MJ (1999) Primary autosomal recessive microcephaly: homozygosity mapping of MCPH4 to chromosome 15. Am J Hum Genet 65(5):1465–1469
Stromme P, Dahl E, Flage T, Stene-Johansen H (1993) Apple peel intestinal atresia in siblings with ocular anomalies and microcephaly. Clin Genet 44(4):208–210
Kavaslar GN, Onengut S, Derman O, Kaya A, Tolun A (2000) The novel genetic disorder microhydranencephaly maps to chromosome 16p13.3-12.1. Am J Hum Genet 66(5):1705–1709
Rauch A, Wieczorek D, Graf E, Wieland T, Endele S, Schwarzmayr T, Albrecht B, Bartholdi D, Beygo J, Di Donato N, Dufke A, Cremer K, Hempel M, Horn D, Hoyer J, Joset P, Röpke A, Moog U, Riess A, Thiel CT, Tzschach A, Wiesener A, Wohlleber E, Zweier C, Ekici AB, Zink AM, Rump A, Meisinger C, Grallert H, Sticht H, Schenck A, Engels H, Rappold G, Schröck E, Wieacker P, Riess O, Meitinger T, Reis A, Strom TM (2012) Range of genetic mutations associated with severe non-syndromic sporadic intellectual disability: an exome sequencing study. Lancet 380(9854):1674–1682
Ochiai H, Miyamoto T, Kanai A, Hosoba K, Sakuma T, Kudo Y, Asami K, Ogawa A, Watanabe A, Kajii T, Yamamoto T, Matsuura S (2013) TALEN-mediated single-base-pair editing identification of an intergenic mutation upstream of BUB1B as causative of PCS (MVA) syndrome. Proc Natl Acad Sci U S A 111:1461
Pinson L, Mannini L, Willems M, Cucco F, Sirvent N, Frebourg T, Quarantotti V, Collet C, Schneider A, Sarda P, Geneviève D, Puechberty J, Lefort G, Musio A (2014) CEP57 mutation in a girl with mosaic variegated aneuploidy syndrome. Am J Med Genet A 164(1):177–181
Chan GKT, Schaar BT, Yen TJ (1998) Characterization of the kinetochore binding domain of CENP-E reveals interactions with the kinetochore proteins CENP-F and hBUBR1. J Cell Biol 143(1):49–63
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2017 Springer International Publishing AG
About this chapter
Cite this chapter
de Wolf, B., Kops, G.J.P.L. (2017). Kinetochore Malfunction in Human Pathologies. In: Gotta, M., Meraldi, P. (eds) Cell Division Machinery and Disease. Advances in Experimental Medicine and Biology, vol 1002. Springer, Cham. https://doi.org/10.1007/978-3-319-57127-0_4
Download citation
DOI: https://doi.org/10.1007/978-3-319-57127-0_4
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-319-57125-6
Online ISBN: 978-3-319-57127-0
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)