Abstract
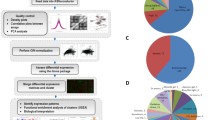
Synechococcus sp. PCC 7002 is a fast-growing cyanobacterium which flourishes in freshwater and marine environments, owing to its ability to tolerate high light intensity and a wide range of salinities. Harnessing the properties of cyanobacteria and understanding their metabolic efficiency has become an imperative goal in recent years owing to their potential to serve as biocatalysts for the production of renewable biofuels. To improve characterisation of metabolic networks, genome-scale models of metabolism can be integrated with multi-omic data to provide a more accurate representation of metabolic capability and refine phenotypic predictions. In this work, a heuristic pipeline is constructed for analysing a genome-scale metabolic model of Synechococcus sp. PCC 7002, which utilises flux balance analysis across multiple layers to observe flux response between conditions across four key pathways. Across various conditions, the detection of significant patterns and mechanisms to cope with fluctuations in light intensity and salinity provides insights into the maintenance of metabolic efficiency.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Hyduke, D., Schellenberger, J., Que, R., Fleming, R., Thiele, I., Orth, J., Feist, A., Zielinski, D., Bordbar, A., Lewis, N., Rahmanian, S., Kang, J., Palsson, B.Ø.: Cobra toolbox 2.0. Protoc. Exch., 1–35 (2011)
Ebrahim, A., Brunk, E., Tan, J., O’brien, E.J., Kim, D., Szubin, R., Lerman, J.A., Lechner, A., Sastry, A., Bordbar, A., Feist, A., Palsson, B.Ø.: Multi-omic data integration enables discovery of hidden biological regularities. Nat. Commun. 7, 13091 (2016)
Reed, J.L.: Shrinking the metabolic solution space using experimental datasets. PLoS Comput. Biol. 8(8), e1002662 (2012)
Ludwig, M., Bryant, D.A.: Synechococcus sp. strain PCC 7002 transcriptome: acclimation to temperature, salinity, oxidative stress, and mixotrophic growth conditions. Front. Microbiol. 3, 354 (2012)
Hendry, J.I., Prasannan, C.B., Joshi, A., Dasgupta, S., Wangikar, P.P.: Metabolic model of synechococcus sp. PCC 7002: prediction of flux distribution and network modification for enhanced biofuel production. Bioresour. Technol. 213, 190–197 (2016)
Ruffing, A.M., Jensen, T.J., Strickland, L.M.: Genetic tools for advancement of synechococcus sp. PCC 7002 as a cyanobacterial chassis. Microb. Cell Fact. 15(1), 190 (2016)
Rügen, M., Bockmayr, A., Steuer, R.: Elucidating temporal resource allocation and diurnal dynamics in phototrophic metabolism using conditional FBA. Sci. Rep. 5, 15247 (2015)
Reimers, A.-M., Knoop, H., Bockmayr, A., Steuer, R.: Evaluating the stoichiometric and energetic constraints of cyanobacterial diurnal growth. arXiv preprint arXiv:1610.06859 (2016)
Angione, C., Conway, M., Lió, P.: Multiplex methods provide effective integration of multi-omic data in genome-scale models. BMC Bioinform. 17(4), 257 (2016)
Angione, C., Lió, P.: Predictive analytics of environmental adaptability in multi-omic network models. Sci. Rep. 5, 15147 (2015)
Ludwig, M., Bryant, D.A.: Transcription profiling of the model cyanobacterium synechococcus sp. strain PCC 7002 by next-gen (SOLiD) sequencing of CDNA. Front. Microbiol. 2(41), 41 (2011)
Ludwig, M., Bryant, D.A.: Acclimation of the global transcriptome of the cyanobacterium synechococcus sp. strain PCC 7002 to nutrient limitations and different nitrogen sources. Front. Microbiol. 3, 145 (2012)
Yang, Y., Feng, J., Li, T., Ge, F., Zhao, J.: CyanOmics: an integrated database of omics for the model cyanobacterium synechococcus sp. PCC 7002. Database 2015, bau127 (2015)
Stevens, S.E., Porter, R.D.: Transformation in agmenellum quadruplicatum. Proc. Natl. Acad. Sci. 77(10), 6052–6056 (1980)
Rajaram, H., Chaurasia, A.K., Apte, S.K.: Cyanobacterial heat-shock response: role and regulation of molecular chaperones. Microbiology 160(4), 647–658 (2014)
Feist, A.M., Palsson, B.O.: The biomass objective function. Curr. Opin. Microbiol. 13(3), 344–349 (2010)
Vu, T.T., Hill, E.A., Kucek, L.A., Konopka, A.E., Beliaev, A.S., Reed, J.L.: Computational evaluation of synechococcus sp. PCC 7002 metabolism for chemical production. Biotechnol. J. 8(5), 619–630 (2013)
Xiong, Q., Feng, J., Li, S.T., Zhang, G.Y., Qiao, Z.X., Chen, Z., Wu, Y., Lin, Y., Li, T., Ge, F., Zhao, J.D.: Integrated transcriptomic and proteomic analysis of the global response of synechococcus to high light stress. Mol. Cell. Proteomics 14(4), 1038–1053 (2015)
Brunk, E., George, K.W., Alonso-Gutierrez, J., Thompson, M., Baidoo, E., Wang, G., Petzold, C.J., McCloskey, D., Monk, J., Yang, L., O’Brien, E.J., Batth, T.S., Martin, H.G., Feist, A., Adams, P.D., Keasling, J.D., Palsson, B.Ø., Lee, T.S.: Characterizing strain variation in engineered E. coli using a multi-omics-based workflow. Cell Syst. 2(5), 335–346 (2016)
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2017 Springer International Publishing AG
About this paper
Cite this paper
Vijayakumar, S., Angione, C. (2017). Multi-omic Data Integration Elucidates Synechococcus Adaptation Mechanisms to Fluctuations in Light Intensity and Salinity. In: Rojas, I., Ortuño, F. (eds) Bioinformatics and Biomedical Engineering. IWBBIO 2017. Lecture Notes in Computer Science(), vol 10208. Springer, Cham. https://doi.org/10.1007/978-3-319-56148-6_19
Download citation
DOI: https://doi.org/10.1007/978-3-319-56148-6_19
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-319-56147-9
Online ISBN: 978-3-319-56148-6
eBook Packages: Computer ScienceComputer Science (R0)