Abstract
Biomass is an important phenotypic trait in plant growth analysis. In this study, we established and compared 8 models for measuring aboveground biomass of 402 rice varieties. Partial least squares (PLS) regression and all subsets regression (ASR) were carried out to determine the effective predictors. Then, 6 models were developed based on support vector regression (SVR). The kernel function used in this study was radial basis function (RBF). Three different optimization methods, Genetic Algorithm (GA) K-fold Cross Validation (K-CV), and Particle Swarm Optimization (PSO), were applied to optimize the penalty error C and RBF \( \upgamma \). We also compared SVR models with models based on PLS regression and ASR. The result showed the model in combination of ASR, GA optimization and SVR outperformed other models with coefficient of determination (R2) of 0.85 for the 268 varieties in the training set and 0.79 for the 134 varieties in the testing set, respectively. This paper extends the application of SVR and intelligent algorithm in measurement of cereal biomass and has the potential of promoting the accuracy of biomass measurement for different varieties.
You have full access to this open access chapter, Download conference paper PDF
Similar content being viewed by others
Keywords
1 Introduction
Plant phenotyping is essential in the study of plant biology, plant functional genomic and plant breeding (Dhondt et al. 2013; Yang et al. 2013; Bolger et al. 2014). Yet plant phenotyping has become a new bottleneck in plant biology. In the recent 5 years, lots of efforts have been done on automatic phenotyping (Duan et al. 2011a, 2011b; Jiang et al. 2012; Huang et al. 2013). However, much work still needs to be done to fill the genotype-phenotype gap.
Biomass is an important phenotypic trait in functional plant biology and plant growth analysis (Honsdorf et al. 2014). Shoot dry weight (DW) is a popular measure of biomass in studying biomass of individual plants (Golzarian et al. 2011). In traditional measurement of DW, the shoot of the plant is cut off, oven-dried to constant weight and weighed by a balance. The low efficiency of the traditional method makes it almost impossible for investigation of a large population of plants. In addition, because the traditional measurement is destructive, continuous inspection of DW over time for an individual plant is infeasible.
Inference of biomass based on machine vision and image analysis allows for non-destructive, high-throughput and continuous measurement of a large quantity of samples. There are researches contributing to automatic measurement of plant biomass (Rajendran et al. 2009; Munns et al. 2010; Hairmansis et al. 2014). However, these researches were only satisfying for young plant (several weeks after sowing) of few varieties.
Based on the statistical learning theory, Support Vector Machine (SVM) is advantageous in robustness to high input space dimension and generalization capabilities (Vapnik, 1995). Support Vector Regression (SVR) is an extension of SVM for regression application and is especially useful in presence of outliers and non-linearities (Brereton and Lloyd 2010).
This study aims to establish a model for measuring aboveground biomass of different rice varieties based on SVR. To the best of our knowledge, no publication available use SVR for biomass measurement.
2 Materials and Methods
2.1 Plant Materials and Image Acquisition
402 rice plants (402 accessions with 1 replicate) were grown in the greenhouse. At late booting stage, all the plants were imaged with a rice automatic phenotyping platform (RAP) (Yang et al. 2014). A turntable rotated the plant and a charge-coupled device (CCD) camera (Stingray F-504C, Applied Vision Technologies, Germany) acquires images at 30° intervals. For each plant, 12 color images at different angles were taken. Simultaneously, a linear X-ray CT captured sinogram of the plant, from which section image were reconstructed and used to extract the tiller number (Yang et al. 2011). The images were saved in the computer for further processing. Next, the plants were harvested and manually measured for the shoot dry weight (DW).
2.2 Feature Extraction and Feature Selection
After image acquisition, the images were analyzed and 39 features, including tiller number, 8 texture features and 30 morphological features, were extracted for each plant. The 39 features included 33 features introduced in Yang et al. (Yang et al. 2014), differential boxing counting dimension (DBC), ratio of plant area to area of bounding rectangle (ABR), greenness area (A_G), yellowness area (A_Y), information fractal dimension (IFD), ratio of perimeter to area (PAR). The features were then used as the potential predictors for DW.
To determine the effective predictors, partial least squares (PLS) regression (Cho et al. 2007) and all subsets regression (ASR) were carried out (Montgomery et al. 2012). PLS regression was accomplished using Matlab 2012b. Prior to performing the PLS regression, the data were normalized so that the mean value and standard deviation of the data was zero and one, respectively. The leave-one-out cross-validation method was performed to determine the optimal number of PLS factors. ASR was done using SAS 9.3. The Cp criterion was used for selecting the best subset. The effective predictors were then used for model input.
2.3 Model Construction and Comparison
The 402 samples were randomly divided into two subsets at 2:1 ratio: 268 samples for training set and 134 samples for testing set. The training set and the testing set was applied for constructing model and evaluating the performance of the model, respectively.
6 models were developed based on support vector regression (SVR). The radial basis function (RBF) only needs to optimize one parameter (the value of \( \upgamma \)) and was adopted as the kernel function in this study. Penalty error C and RBF \( \upgamma \) were key to the performance of SVR (Brereton and Lloyd 2010). A larger C generates more significant misclassifications but meanwhile leads to a more complex boundary. And inappropriate RBF \( \upgamma \) may lead to overfitting. In this study, three different optimization methods, K-fold Cross Validation (K-CV, in this study 5-CV), Genetic Algorithm (GA) (Storn and Price 1997) and Particle Swarm Optimization (PSO) (Clerc and Kennedy 2002), were applied and compared to optimize C and \( \upgamma \). The fitness function for GA and PSO was set as the mean squared error under 5-CV in this study. Libsvm, a popular SVM software package for Matlab designed by professor Lin Chih-Jen was used to accomplish SVR in this study.
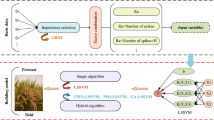
In comparison with SVM models, we also built models based on PLS regression and multiple linear regression (MLR). In total, 8 models were developed and compared in this study (Table 1). Figure 1 shows the flowchart of the model construction.
For model comparison, coefficient of determination (R2), mean absolute percentage error (MAPE, Eqs. 1-2) and standard deviation of the absolute percentage error (SAPE, Eq. 3) for training set and testing set were computed for each model.
where DW i.automatic represents the dry weight measured automatically using the method described, DW i.manual represents the dry weight measured manually, and n represents the number of samples.
3 Results and Discussion
After PLS regression, 4 PLS factors were selected. And a subset with 18 features was selected as the best subset using ASR. The selected predictors were used as input for the models.
Table 2 illustrates the comparison of the 8 models. Note that when using the best subset by the Cp criterion as independent variables for MLR modelling, the model suffered from multi-collinearity problem. So the following strategy was used to select the feature subset for MLR: (1) the subset that has the maximum R2 among all subsets with i \( i = 1,\,2,\, \cdots ,\,39 \) features was deemed as the best subset with i features, (2) the 39 best subsets were used as independent variables to build MLR models and the model was chosen as the optimal MLR model if it had the largest number of independent variables and did not present multi-collinearity. Finally, a model with 3 independent variables (exclude the constant) was chosen as the optimal MLR model.
As seen from the Table 2, the ASR-GA-SVR model outperformed other models, with R2 of 0.85, MAPE of 10.20 % and SAPE of 9.20 % for the training set and R2 of 0.79, MAPE of 12.44 % and SAPE of 9.79 %for the testing set, respectively. Consequently, the ASR-GA-SVR model was chosen as the optimal DW model. The SVR models were generally noticeably advantageous for the training set compared with PLS and ASR model. However, for the testing set, the performance of the PLS and ASR model were comparative to the SVR models. This was because the optimal C and γ were chosen to obtain the best performance (minimum mean squared error) for the training set but could not guarantee to get the best performance for the testing set under the optimal C and γ.
Figures 2, 3 and 4 show the performance of the final DW model (ASR-GA-SVR model), the PLS model and ASR model, respectively.
4 Conclusions
This paper presents 8 models based on SVR, PLS and ASR for measuring aboveground biomass of different rice varieties. The result showed the ASR-GA-SVR model outperformed other models with R2 of 0.85, MAPE of 10.20 % and SAPE of 9.20 % for the training set and R2 of 0.79, MAPE of 12.44 % and SAPE of 9.79 %for the testing set, respectively. The study extends the application of SVR and intelligent algorithm in the measurement of plant biomass. The method has the potential to promote the accuracy of biomass measurement for different varieties and thus contributes to automatic plant phenotyping.
References
Bolger, M., Weisshaar, B., Scholz, U., Stein, N., Usadel, B., Mayer, K.: Plant genome sequencing - applications for crop improvement. Plant Biotechnol. J. 8(1), 31–37 (2014)
Brereton, R., Lloyd, G.: Support vector machines for classification and regression. Analyst 135(2), 230–267 (2010)
Cho, M., Skidmore, A., Corsi, F., van Wieren, S., Sobhan, I.: Estimation of green grass/herb biomass from airborne hyperspectral imagery using spectral indices and partial least squares regression. Int. J. Appl. Earth Obs. Geoinf. 9, 414–424 (2007)
Clerc, M., Kennedy, J.: The particle swarm - explosion, stability, and convergence in a multidimensional complex space. IEEE Trans. Evol. Comput. 6(1), 58–73 (2002)
Dhondt, S., Wuyts, N., Inzé, D.: Cell to whole-plant phenotyping: the best is yet to come. Trends Plant Sci. 18(8), 428–439 (2013)
Duan, L., Yang, W., Huang, C., Liu, Q.: A novel machine-vision-based facility for the automatic evaluation of yield-related traits in rice. Plant Methods 7, 44 (2011a)
Duan, L., Yang, W., Bi, K., Chen, S., Luo, Q., Liu, Q.: Fast discrimination and counting of filled/unfilled rice spikelets based on two modal imaging. Comput. Electron. Agric. 75(1), 196–203 (2011b)
Golzarian, M., Frick, R., Rajendran, K., Berger, B., Roy, S., Tester, M., Lun, D.: Accurate inference of shoot biomass from high-throughput images of cereal plants. Plant Methods 7, 11 (2011)
Hairmansis, A., Berger, B., Tester, M., Roy, S.: Image-based phenotyping for non-destructive screening of different salinity tolerance traits in rice. Rice 7, 16 (2014)
Honsdorf, N., March, T., Berger, B., Tester, M., Pillen, K.: High-throughput phenotyping to detect drought tolerance QTL in wild barley introgression lines. PLoS ONE 9(5), e97047 (2014). doi:10.1371/journal.pone.0097047
Huang, C., Yang, W., Duan, L., Jiang, N., Chen, G., Xiong, L., Liu, Q.: Rice panicle length measuring system based on dual-camera imaging. Comput. Electron. Agric. 98, 158–165 (2013)
Jiang, N., Yang, W., Duan, L., Xu, X., Huang, C., Liu, Q.: Acceleration of CT reconstruction for wheat tiller inspection based on adaptive minimum enclosing rectangle. Comput. Electron. Agric. 85, 123–133 (2012)
Montgomery, D., Peck, E., Vining, G.: Introduction to Linear Regression Analysis. Wiley, Hoboken (2012)
Munns, R., James, R., Sirault, X., Furbank, R., Jones, H.: New phenotyping methods for screening wheat and barley for beneficial responses to water deficit. J. Exp. Bot. 61, 3499–3507 (2010)
Rajendran, K., Tester, M., Roy, S.: Quantifying the three main components of salinity tolerance in cereals. Plant Cell Environ. 32(3), 237–249 (2009)
Storn, R., Price, K.: Differential evolution – a simple and efficient heuristic for global optimization over continuous spaces. J. Global Optim. 11(4), 341–359 (1997)
Vapnik, V.: The Nature of Statistical Learning Theory. Springer, New York (1995)
Yang, W., Xu, X., Duan, L., Luo, Q., Chen, S., Zeng, S., Liu, Q.: High-throughput measurement of rice tillers using a conveyor equipped with X-ray computed tomography. Rev. Sci. Instrum. 82(2), 025102–025109 (2011)
Yang, W., Duan, L., Chen, G., Xiong, L., Liu, Q.: Plant phenomics and high-throughput phenotyping: accelerating rice functional genomics using multidisciplinary technologies. Curr. Opin. Plant Biol. 16, 180–187 (2013)
Yang, W., Guo, Z., Huang, C., Duan, L., Chen, G., Jiang, N., Fang, W., Feng, H., Xie, W., Lian, X., Wang, G., Luo, Q., Zhang, Q., Liu, Q., Xiong, L.: Combining high-throughput phenotyping and genome-wide association studies to reveal natural genetic variation in rice. Nat. Commun. 5, 5087 (2014)
Acknowledgment
This work was supported by the Fundamental Research Funds for the Central Universities (2662015QC006, 2662015QC016, 2013PY034, 2662014BQ036), the National High Technology Research and Development Program of China (2013AA102403), the National Natural Science Foundation of China (30921091, 31200274).
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2016 IFIP International Federation for Information Processing
About this paper
Cite this paper
Duan, L., Yang, W., Chen, G., Xiong, L., Huang, C. (2016). Accurate Inference of Rice Biomass Based on Support Vector Machine. In: Li, D., Li, Z. (eds) Computer and Computing Technologies in Agriculture IX. CCTA 2015. IFIP Advances in Information and Communication Technology, vol 478. Springer, Cham. https://doi.org/10.1007/978-3-319-48357-3_35
Download citation
DOI: https://doi.org/10.1007/978-3-319-48357-3_35
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-319-48356-6
Online ISBN: 978-3-319-48357-3
eBook Packages: Computer ScienceComputer Science (R0)