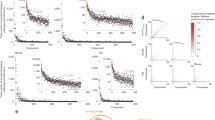
Abstract
The applications of information on copy number changes in cancer have been twofold. Recognizing that regions of copy number gain signalled the location of oncogenes and that, similarly, copy number loss signalled the location of tumour suppressor genes, has resulted in screening of the minimally defined regions for candidate genes involved in tumourigenesis. Once candidates emerged, other evidence of their role in tumours was sought, by functional assays for example, and a huge literature built up describing these gene classes. Even without knowledge of how the genes acted in the development of tumours, the second application has been to correlate the chromosomal abnormalities with various clinical parameters, again resulting in many thousands of publications, although to date the translation of laboratory observations into clinical practice is still not widespread.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Caspersson T, Zech L, Johansson C (1970) Differential binding of alkylating fluorochromes in human chromosomes. Exp Cell Res 60:315–319
Mitelman (ed) (1983) Catalog of chromosome aberrations in cancer. Wiley-Liss
George D, Powers V (1982) Amplified DNA sequences in Y1 mouse adrenal tumor cells: association with double minutes and localization to a homogeneously staining chromosomal region. Proc Natl Acad Sci U S A 79:1597–1601
Bishop J (1983) Cellular oncogenes and retroviruses. Annu Rev Biochem 52:301–354
Alitalo K, Schwab M, Lin C, Varmus H, Bishop J (1983) Homogeneously staining chromosomal regions contain amplified copies of an abundantly expressed cellular oncogene (c-myc) in malignant neuroendocrine cells from a human colon carcinoma. Proc Natl Acad Sci U S A 80:1707–1711
Johnson B, Brennan J, Ihde D, Gazdar A (1992) myc family DNA amplification in tumors and tumor cell lines from patients with small-cell lung cancer. J Natl Cancer Inst Monogr 13:39–43
Johnson B, Makuch R, Simmons A, Gazdar A, Burch D et al (1988) myc family DNA amplification in small cell lung cancer patients’ tumors and corresponding cell lines. Cancer Res 48:5163–5166
Ma Y, Wei S, Lin Y, Lung J, Chang T et al (2000) PIK3CA as an oncogene in cervical cancer. Oncogene 19:2739–2744
Qian J, Massion P (2008) Role of chromosome 3q amplification in lung cancer. J Thorac Oncol 3:212–215
McCaughan F, Pole J, Bankier A, Konfortov B, Carroll B et al (2010) Progressive 3q amplification consistently targets SOX2 in preinvasive squamous lung cancer. Am J Respir Crit Care Med 182:83–91
Bass A, Watanabe H, Mermel C, Yu S, Perner S et al (2009) SOX2 is an amplified lineage-survival oncogene in lung and esophageal squamous cell carcinomas. Nat Genet 41:1238–1242
Justilien V, Walsh M, Ali S, Thompson E, Murray N et al (2014) The PRKCI and SOX2 oncogenes are coamplified and cooperate to activate Hedgehog signaling in lung squamous cell carcinoma. Cancer Cell 10:139–151
Wang J, Qian J, Hoeksema M, Zou Y, Espinosa A et al (2013) Integrative genomics analysis identifies candidate drivers at 3q26-29 amplicon in squamous cell carcinoma of the lung. Clin Cancer Res 19:5580–5590
Toschi L, Finocchiaro G, Nguyen T, Skokan M, Giordano L et al (2014) Increased SOX2 gene copy number is associated with FGFR1 and PIK3CA gene gain in non-small cell lung cancer and predicts improved survival in early stage disease. PLoS ONE 9:e95303
Shen L, Huang X, Xie X, Su J, Yuan J et al (2014) High expression of SOX2 and OCT4 indicates radiation resistance and an independent negative prognosis in cervical squamous cell carcinoma. J Histochem Cytochem 62:499–509
Hansen M, Cavenee W (1988) Retinoblastoma and the progression of tumor genetics. Trends Genet 4:125–128
Knudson AG Jr (1971) Mutation and cancer: statistical study of retinoblastoma. Proc Natl Acad Sci U S A 68:820–823
White R (1992) Inherited cancer genes. Curr Opin Genet Dev 2:53–57
Levy D, Smith K, Beazer-Barclay Y, Hamilton S, Vogelstein et al (1994) Inactivation of both APC alleles in human and mouse tumors. Cancer Res 54:5953–5958
Smith S, Easton D, Evans D, Ponder B (1992) Allele losses in the region 17q12-21 in familial breast and ovarian cancer involve the wild-type chromosome. Nat Genet 2:128–131
Gudmundsson J, Johannesdottir G, Bergthorsson J, Arason A, Ingvarsson S et al (1995) Different tumor types from BRCA2 carriers show wild-type chromosome deletions on 13q12-q13. Cancer Res 55:4830–3832
Baker S, Fearon E, Nigro J, Hamilton S, Preisinger A et al (1989) Chromosome 17 deletions and p53 gene mutations in colorectal carcinomas. Science 244:217–221
Hollstein M, Sidransky D, Vogelstein B, Harris C (1991) p53 mutations in human cancers. Science 253:49–53
Devilee P, Cleton-Jansen A, Cornelisse C (2001) Ever since Knudson. Trends Genet 17:569–573
Kamb A, Gruis N, Weaver-Feldhaus J, Liu Q, Harshman K et al (1994) A cell cycle regulator potentially involved in genesis of many tumor types. Science 264:436–440
Li J, Yen C, Liaw D, Podsypanina K, Bose S, Wang SI et al (1997) PTEN, a putative protein tyrosine phosphatase gene mutated in human brain, breast, and prostate cancer. Science 275:1943–1947
Cox C, Bignell G, Greenman C, Stabenau A, Warren W et al (2005) A survey of homozygous deletions in human cancer genomes. Proc Natl Acad Sci U S A 102:4542–4547
Kok K, Naylor S, Buys C (1997) Deletions of the short arm of chromosome 3 in solid tumors and the search for suppressor genes. Adv Cancer Res 71:27–92
Burbee D, Forgacs E, Zöchbauer-Müller S, Shivakumar L, Fong K et al (2011) Epigenetic inactivation of RASSF1A in lung and breast cancers and malignant phenotype suppression. J Natl Cancer Inst 93:691–699
Berger A, Knudson A, Pandolfi P (2011) A continuum model for tumor suppression. Nature 476:163–169
Gatto F, Nookaew I, Nielsen J (2014) Chromosome 3p loss of heterozygosity is associated with a unique metabolic network in clear cell renal carcinoma. Proc Natl Acad Sci U S A 111:E866–E875
Watanabe H, Ma Q, Peng S, Adelmant G, Swain D et al (2014) SOX2 and p63 colocalize at genetic loci in squamous cell carcinomas. J Clin Invest 124:1636–1645
Booden MA, Ulka AS, Der CJ (2006) Cellular assays of oncogene transformation. In: Cellis JE (ed) Cell biology a laboratory handbook, published by Elsevier, pp 345–352
Brummelkamp T, Bernards R, Agami R (2002) A system for stable expression of short interfering RNAs in mammalian cells. Science 296:550–553
Engel AM, Schou G (2006) Assays of tumorigencity in nude mice. In: Cellis JE (ed) Cell biology a laboratory handbook, published by Elsevier, pp 353–357
Akavia U, Litvin O, Kim J, Sanchez-Garcia F, Kotliar D et al (2010) An integrated approach to uncover drivers of cancer. Cell 143:1005–1017
Santarius T, Shipley J, Brewer D, Stratton MR, Cooper C (2010) A census of amplified and overexpressed human cancer genes. Nat Rev Cancer 10:59–64
Sawey E, Chanrion M, Cai C, Wu G, Zhang J et al (2011) Identification of a therapeutic strategy targeting amplified FGF19 in liver cancer by Oncogenomic screening. Cancer Cell 19:347–358
Eifert C, Powers R (2012) From cancer genomes to oncogenic drivers, tumour dependencies and therapeutic targets. Nat Rev Cancer 12:572–578
Soucek L, Evan G (2010) The ups and downs of Myc biology. Curr Opin Genet Dev 20:91–95
Schwab M, Alitalo K, Klempnauer KH, Varmus HE, Bishop J et al (1983) Amplified DNA with limited homology to myc cellular oncogene is shared by human neuroblastoma cell lines and a neuroblastoma tumour. Nature 305:245–248
Brodeur G, Seeger R, Schwab M, Varmus H, Bishop J (1984) Amplification of N-myc in untreated human neuroblastomas correlates with advanced disease stage. Science 224:1121–1124
Slamon D, Clark G, Wong S, Levin W, Ullrich A et al (1987) Human breast cancer: correlation of relapse and survival with amplification of the HER-2/neu oncogene. Science 235:177–182
Cobleigh M, Vogel C, Tripathy D, Robert N, Scholl S et al (1999) Multinational study of the efficacy and safety of humanized anti-HER2 monoclonal antibody in women who have HER2-overexpressing metastatic breast cancer that has progressed after chemotherapy for metastatic disease. J Clin Oncol 17:2639–2648
Verma S, Miles D, Gianni L, Krop IE, Welslau M et al (2012) Trastuzumab emtansine for HER2-positive advanced breast cancer. N Engl J Med 367:1783–1791
Rakha E, Ellis O (2014) Breast cancer: updated guideline recommendations for HER2 testing. Nat Rev Clin Oncol 11:8–9
Myllykangas S, Himberg J, Böhling T, Nagy B, Hollmén J et al (2006) DNA copy number amplification profiling of human neoplasms. Oncogene 25:7324–7332
Myllykangas S, Böhling T, Knuutila S (2007) Specificity, selection and significance of gene amplifications in cancer. Semin Cancer Biol 17:42–55
Scheinin I, Myllykangas S, Borze I, Böhling T, Knuutila S et al (2008) CanGEM: mining gene copy number changes in cancer. Nucleic Acids Res 36:D830–D835
Crockford A, Jamal-Hanjani M, Hicks J, Swanton C (2014) Implications of intratumour heterogeneity for treatment stratification. Pathology 232:264–273
Poste G (2012) Biospecimens, biomarkers, and burgeoning data: the imperative for more rigorous research standards. Trends Mol Med 18:717–722
Mehta S, Shelling A, Muthukaruppan A, Lasham A, Blenkiron C et al (2010) Predictive and prognostic molecular markers for cancer medicine. Ther Adv Med Oncol 2:2125–2148
Chen HY, Yu SL, Chen CH, Chang GC, Chen CY et al (2007) A five-gene signature and clinical outcome in non-small-cell lung cancer. N Engl J Med 356:11–20
Hicks J, Krasnitz A, Lakshmi B, Navin NE, Riggs M et al (2006) Novel patterns of genome rearrangement and their association with survival in breast cancer. Genome Res 16:1465–1479
Campbell P, Stephens P, Pleasance E, O’Meara S, Li H et al (2008) Identification of somatically acquired rearrangements in cancer using genome-wide massively parallel paired-end sequencing. Nat Genet 40:722–729
Wood H, Belvedere O, Conway C, Daly C, Chalkley R et al (2010) Using next generation sequencing for high resolution multiplex analysis of copy number variation from nanogram quantities of DNA from formalin fixed paraffin embedded specimens. Nucl Acid Res 38:e151
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2015 Springer International Publishing Switzerland
About this chapter
Cite this chapter
Wood, H., Rabbitts, P. (2015). Copy Number Changes in Carcinomas: Applications. In: Rowley, J., Le Beau, M., Rabbitts, T. (eds) Chromosomal Translocations and Genome Rearrangements in Cancer. Springer, Cham. https://doi.org/10.1007/978-3-319-19983-2_6
Download citation
DOI: https://doi.org/10.1007/978-3-319-19983-2_6
Publisher Name: Springer, Cham
Print ISBN: 978-3-319-19982-5
Online ISBN: 978-3-319-19983-2
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)