Abstract
Chromium(Cr) is a heavy metal and exerts a profusion of toxic effects in a plethora of organisms (bacteria, plants, fungi, archaebacteria, algae, and many more). Cr is increasingly being accumulated in the soil, freshwater, wastewater, etc. because of extensive anthropogenic activities. It is carcinogenic for humans and due to the alarming upsurge of Cr concentration in the environment, it is treated as a priority pollutant. Cr manifests in various valence states of which Cr(VI) (most toxic) and Cr(III) are the most stable. Cr when accumulated inside cells of an organism, leads to the production of reactive oxygen species (ROS) hampering a network of molecular, physiological, and metabolic processes. Several genetic studies have revealed that certain organisms contain specific genes conferring the potential to withstand as well as remediate high concentrations of Cr. These organisms may remove, accumulate or reduce Cr(VI) to the less toxic Cr(III) form. Recent years have shed light on the various genes involved in Cr tolerance. These genes have been exploited using genetic engineering (GE) tools to construct genetically modified organisms (GMOs) with the ability of Cr bioremediation. These GMOs can be allowed to grow in Cr-contaminated regions and ameliorate its toxic effects. This chapter summarizes the Cr toxicity in organisms, the significance of detoxification genes in Cr stress response, mechanisms of Cr tolerance and enzyme activity in Cr reduction and resistance. It also highlights various bacteria, plants, and other organisms such as phages, yeast, etc., utilized for bioremediation in Cr-contaminated regions.
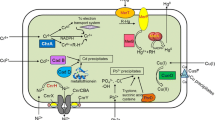
Graphical Abstract: Schematic representation of genetic engineering techniques used For Cr bioremediation.

Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Ackerley DF, Gonzalez CF, Keyhan M et al (2004a) Mechanism of chromate reduction by the Escherichia coli protein, NfsA, and the role of different chromate reductases in minimizing oxidative stress during chromate reduction. Environ Microbiol 6:851–860
Ackerley DF, Gonzalez CF, Park CH et al (2004b) Chromate-reducing properties of Soluble Flavoproteins from Pseudomonas putida and Escherichia coli. Appl Environ Microbiol 70:873–882
Akkurt Ş, Oğuz M, Alkan Uçkun, A (2022) Bioreduction and bioremoval of hexavalent chromium by genetically engineered strains (Escherichia coli MT2A and Escherichia coli MT3). World J Microbiol Biotechnol 38:45
Bennett LE, Burkhead JL, Hale KL et al (2000) Bioremediation and biodegradation analysis of transgenic Indian mustard plants for phytoremediation of metal-contaminated mine tailings. Hirschi
Dados A, Omirou M, Demetriou K et al (2015) Rapid remediation of soil heavily contaminated with hydrocarbons: a comparison of different approaches. Ann Microbiol 65:241–251
Daud MK, Mei L, Variath MT et al (2014) Chromium(VI) uptake and tolerance potential in cotton cultivars: effect on their root physiology, ultramorphology, and oxidative metabolism. Biomed Res Int
del Bubba M, Ancillotti C, Checchini L et al (2013) Chromium accumulation and changes in plant growth, selected phenolics and sugars of wild type and genetically modified Nicotiana langsdorffii. J Hazard Mater 262:394–403
Deng P, Tan X, Wu Y et al (2014) Cloning and sequence analysis demonstrate the chromate reduction ability of a novel chromate reductase gene from Serratia sp. Exp Ther Med 9:795–800
di Bona KR, Love S, Rhodes NR et al (2011) Chromium is not an essential trace element for mammals: Effects of a “low-chromium” diet. J Biol Inorg Chem 16:381–390
Fajardo C, Martín C, Garrido E et al (2022) Copper and Chrkomium toxicity is mediated by oxidative stress in Caenorhabditis elegans: the use of nanoparticles as an immobilization strategy. Environ Toxicol Pharmacol 92
Flora SJ (2009) Bioscience, structural, chemical and biological aspects of antioxidants for strategies against metal and metalloid exposure. Oxid Med Cell Longev 2:191–206
Frederick TM, Taylor EA, Willis JL et al (2013) Chromate reduction is expedited by bacteria engineered to produce the compatible solute trehalose. Biotechnol Lett 35:1291–1296
Fuoco R, Bogani P, Capodaglio G et al (2013) Response to metal stress of Nicotiana langsdorffii plants wild-type and transgenic for the rat glucocorticoid receptor gene. J Plant Physiol 170:668–675
Gill SS, Tuteja N (2010) Reactive oxygen species and antioxidant machinery in abiotic stress tolerance in crop plants. Plant Physiol Biochem 48:909–930
Gu R, Gao J, Dong L et al (2020) Chromium metabolism characteristics of coexpression of ChrA and ChrT gene. Ecotoxicol Environ Saf 204
Huang D, Yu P, Ye M et al (2021) Enhanced mutualistic symbiosis between soil phages and bacteria with elevated chromium-induced environmental stress. Microbiome 9
Jaishankar M, Tseten T, Anbalagan N et al (2014) Toxicity, mechanism and health effects of some heavy metals. Interdiscip Toxicol 7:60–72
Jin TE, Kim IG, Kim WS, Suh SC, Kim BD, Rhim SL (2001) Expression of chromium (VI) reductase gene of heavy metal reducing bacteria in tobacco plants. J Plant Biotechnol 3:13–17
Johnson J, Schewel L, Graedel TE (2006) The contemporary anthropogenic chromium cycle. Environ Sci Technol 40:7060–7069
Kanagaraj G, Elango L (2019) Chromium and fluoride contamination in groundwater around leather tanning industries in southern India: Implications from stable isotopic ratio Δ53Cr/Δ52Cr, geochemical and geostatistical modelling. Chemosphere 220:943–953
Kao WC, Huang CC, Chang JS (2008) Biosorption of nickel, chromium and zinc by MerP-expressing recombinant Escherichia coli. J Hazard Mater 158:100–106
Kim YJ, Kim JH, Lee CE et al (2006) Expression of yeast transcriptional activator MSN1 promotes accumulation of chromium and sulfur by enhancing sulfate transporter level in plants. FEBS Lett 580:206–210
Kumar S, Asif MH, Chakrabarty D et al (2013) Expression of a rice Lambda class of glutathione S-transferase, OsGSTL2, in Arabidopsis provides tolerance to heavy metal and other abiotic stresses. J Hazard Mater 248–249:228–237
Li FH, Tang Q, Fan Y-Y et al (2020) Developing a population-state decision system for intelligently reprogramming extracellular electron transfer in Shewanella oneidensis
Li J, Tang Q, Li Y et al (2020) Rediverting electron flux with an engineered CRISPR-ddAsCpf1 system to enhance the pollutant degradation capacity of Shewanella oneidensis. Environ Sci Technol 54:3599–3608
Marques APGC, Rangel AOSS, Castro PML (2009) Remediation of heavy metal contaminated soils: phytoremediation as a potentially promising clean-up technology. Crit Rev Environ Sci Technol 39:622–654
Ngwenya N, Chirwa EMN (2011) Biological removal of cationic fission products from nuclear wastewater. Water Sci Technol 63:124–128
Peitzsch N, Eberz N, Nies DH (1998) Alcaligenes eutrophus as a bacterial chromate sensor the HEPES-buffered me-dium contained the following (per liter of H2O): 0.3 mM Na2KPO4, 0.2 mM K2HPO4, 50 mM HEPES buffer (pH 7.0), 2 g of NH4Cl, 0.2 g of MgSO47H2O, 10 mg of CaCl22H2O, and 5 mg of FeCl36H2O. Analytical-grade salts of CdCl
Reisinger S, Schiavon M, Terry N, Pilon-Smits EAH (2008) Heavy metal tolerance and accumulation in Indian mustard (Brassica juncea L.) expressing bacterial γ-glutamylcysteine synthetase or glutathione synthetase. Int J Phytoremediation 10:440–454
Simin Z, Lanlan D, Yuan HE, Hong X (2017) Characterization of chromate resistance in genetically engineered Escherichia coli expressing chromate ion transporter ChrA. J South Med Univ 37:1290–1295
Srivastava NK, Jha MK, Mall ID, Singh D (2010) Application of genetic engineering for chromium removal from industrial wastewater. Int J Environ Ecol Eng 4:633–638
Srivastava D, Verma G, Chauhan AS et al (2019) Rice (Oryza sativa L.) tau class glutathione S-transferase (OsGSTU30) overexpression in Arabidopsis thaliana modulates a regulatory network leading to heavy metal and drought stress tolerance. Metallomics 11:375–389
Sun GL, Reynolds EE, Belcher AM (2019) Designing yeast as plant-like hyperaccumulators for heavy metals. Nat Commun 10
Tahri Joutey N, Bahafid W, Sayel H et al (2014) Hexavalent chromium removal by a novel Serratia proteamaculans isolated from the bank of Sebou River (Morocco). Environ Sci Pollut Res 21:3060–3072
Tang R, Shen L, Yang L et al (2021) Killing two birds with one stone: biomineralized bacteria tolerate adverse environments and absorb hexavalent chromium. ACS Omega
Terzi H, Yıldız M (2015) Interactive effects of sulfur and chromium on antioxidative defense systems and BnMP1 gene expression in canola (Brassica napus L.) cultivars differing in Cr(VI) tolerance. Ecotoxicology 24:1171–1182
Thacker U, Parikh R, Shouche Y, Madamwar D (2006) Hexavalent chromium reduction by Providencia sp. Process Biochem 41:1332–1337
Tiwari P, Indoliya Y, Chauhan AS et al (2020) Over-expression of rice R1-type MYB transcription factor confers different abiotic stress tolerance in transgenic Arabidopsis. Ecotoxicol Environ Saf 206
Tripathi A, Indoliya Y, Tiwari M et al (2014) Transformed yeast (Schizosaccharomyces pombe) overexpressing rice Tau class glutathione S-transferase (OsGSTU30 and OsGSTU41) shows enhanced resistance to hexavalent chromium. Metallomics 6:1549–1557
Vandecasteele B, Meers E, Vervaeke P et al (2005) Growth and trace metal accumulation of two Salix clones on sediment-derived soils with increasing contamination levels. Chemosphere 58:995–1002
Velez PA, Talano MA, Paisio CE et al (2017) Synergistic effect of chickpea plants and Mesorhizobium as a natural system for chromium phytoremediation. Environ Technol (United Kingdom) 38:2164–2172
Verma JP, Jaiswal DK (2016) Book review: advances in biodegradation and bioremediation of industrial waste. Front Microbiol 6
Wang J, Zhao S, Ling Z et al (2021) Enhanced removal of trivalent chromium from leather wastewater using engineered bacteria immobilized on magnetic pellets. Sci Total Environ 775
Wangeline AL, Burkhead JL, Hale KL et al (1999) Overexpression of ATP Sulfurylase in Indian Mustard: effects on tolerance and accumulation of twelve metals
Zhou S, Dong L, Deng P et al (2017) Reducing capacity and enzyme activity of chromate reductase in a ChrT-engineered strain. Exp Ther Med 14:2361–2366
Author information
Authors and Affiliations
Corresponding authors
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2023 The Author(s), under exclusive license to Springer Nature Switzerland AG
About this chapter
Cite this chapter
Bose, R., Kulkarni, S., Lakkakula, J., Wagh, N.S. (2023). Genetic Engineering for Chromium Removal. In: Kumar, N., Walther, C., Gupta, D.K. (eds) Chromium in Plants and Environment. Environmental Science and Engineering. Springer, Cham. https://doi.org/10.1007/978-3-031-44029-8_7
Download citation
DOI: https://doi.org/10.1007/978-3-031-44029-8_7
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-031-44028-1
Online ISBN: 978-3-031-44029-8
eBook Packages: Earth and Environmental ScienceEarth and Environmental Science (R0)