Abstract
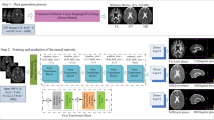
Diffusion Magnetic Resonance Imaging (dMRI) is a powerful non-invasive method for studying white matter tracts of the brain. However, accurate microstructure estimation with fiber orientation distribution (FOD) using existing computational methods requires a large number of diffusion measurements. In clinical settings, this is often not possible for neonates and fetuses because of increased acquisition times and subject movements. Therefore, methods that can estimate the FOD from reduced measurements are of high practical utility. Here, we exploited deep learning and trained a neural network to directly map dMRI data acquired with as low as six diffusion directions to FODs for neonates and fetuses. We trained the method using target FODs generated from densely-sampled multiple-shell data with the multi-shell multi-tissue constrained spherical deconvolution (MSMT-CSD). Detailed evaluations on independent newborns’ test data show that our method achieved estimation accuracy levels on par with the state-of-the-art methods while reducing the number of required measurements by more than an order of magnitude. Qualitative assessments on two out-of-distribution clinical datasets of fetuses and newborns show the consistency of the estimated FODs and hence the cross-site generalizability of the method.
D. Karimi and M. B. Cuadra—Equal contribution.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Aganj, I., et al.: Reconstruction of the orientation distribution function in single-and multiple-shell q-ball imaging within constant solid angle. Magn. Reson. Med. 64(2), 554–566 (2010)
Andescavage, N.N., et al.: Complex trajectories of brain development in the healthy human fetus. Cereb. Cortex 27(11), 5274–5283 (2017)
Bastiani, M., et al.: Automated processing pipeline for neonatal diffusion MRI in the developing human connectome project. Neuroimage 185, 750–763 (2019)
Chen, R., et al.: Deciphering the developmental order and microstructural patterns of early white matter pathways in a diffusion MRI based fetal brain atlas. Neuroimage 264, 119700 (2022)
Christiaens, D., et al.: Multi-shell shard reconstruction from scattered slice diffusion MRI data in the neonatal brain. ISMRM (Paris) (2018)
Counsell, S.J., Arichi, T., Arulkumaran, S., Rutherford, M.A.: Fetal and neonatal neuroimaging. Handb. Clin. Neurol. 162, 67–103 (2019)
Deprez, M., et al.: Higher order spherical harmonics reconstruction of fetal diffusion MRI with intensity correction. IEEE Trans. Med. Imaging 39(4), 1104–1113 (2019)
Descoteaux, M.: High angular resolution diffusion MRI: from local estimation to segmentation and tractography. Ph.D. thesis, Univ. Nice Sophia Antipolis (2008)
Dubois, J., et al.: The early development of brain white matter: a review of imaging studies in fetuses, newborns and infants. Neuroscience 276, 48–71 (2014)
Garyfallidis, E., et al.: Dipy, a library for the analysis of diffusion MRI data. Front. Neuroinform. 8, 8 (2014)
Gholipour, A., et al.: A normative spatiotemporal MRI atlas of the fetal brain for automatic segmentation and analysis of early brain growth. Sci. Rep. 7(1), 476 (2017)
Golkov, V., et al.: Q-space deep learning: twelve-fold shorter and model-free diffusion MRI scans. IEEE TMI 35(5), 1344–1351 (2016)
Hosseini, S., et al.: CTtrack: a CNN+ transformer-based framework for fiber orientation estimation & tractography. Neurosci. Inform. 2(4), 100099 (2022)
Hutter, J., et al.: Time-efficient and flexible design of optimized multishell HARDI diffusion. Magn. Reson. Med. 79(3), 1276–1292 (2018)
Jeurissen, B., et al.: Multi-tissue constrained spherical deconvolution for improved analysis of multi-shell diffusion MRI data. Neuroimage 103, 411–426 (2014)
Jeurissen, B., et al.: Diffusion MRI fiber tractography of the brain. NMR Biomed. 32(4), e3785 (2019)
Karimi, D., et al.: Deep learning-based parameter estimation in fetal diffusion-weighted MRI. Neuroimage 243, 118482 (2021)
Karimi, D., et al.: Learning to estimate the fiber orientation distribution function from diffusion-weighted MRI. Neuroimage 239, 118316 (2021)
Kebiri, H., et al.: Slice estimation in diffusion MRI of neonatal and fetal brains in image and spherical harmonics domains using autoencoders. In: Cetin-Karayumak, S., et al. (eds.) CDMRI 2022. LNCS, vol. 13722, pp. 3–13. Springer, Cham (2022). https://doi.org/10.1007/978-3-031-21206-2_1
Khan, S., et al.: Fetal brain growth portrayed by a spatiotemporal diffusion tensor MRI atlas computed from in utero images. Neuroimage 185, 593–608 (2019)
Kingma, D.P., Ba, J.: Adam: A method for stochastic optimization. arXiv preprint arXiv:1412.6980 (2014)
Koppers, S., Merhof, D.: Direct estimation of fiber orientations using deep learning in diffusion imaging. In: Wang, L., Adeli, E., Wang, Q., Shi, Y., Suk, H.-I. (eds.) MLMI 2016. LNCS, vol. 10019, pp. 53–60. Springer, Cham (2016). https://doi.org/10.1007/978-3-319-47157-0_7
Le Bihan, D., et al.: Diffusion tensor imaging: concepts and applications. J. Magn. Reson. Imaging 13(4), 534–546 (2001)
Li, H., et al.: SuperDTI: ultrafast DTI and fiber tractography with deep learning. Magn. Reson. Med. 86(6), 3334–3347 (2021)
Lin, Z., et al.: Fast learning of fiber orientation distribution function for MR tractography using convolutional neural network. Med. Phys. 46(7), 3101–3116 (2019)
Raffelt, D., et al.: Apparent fibre density: a novel measure for the analysis of diffusion-weighted magnetic resonance images. Neuroimage 59(4), 3976–3994 (2012)
Rokem, A., et al.: Evaluating the accuracy of diffusion MRI models in white matter. PLoS ONE 10(4), e0123272 (2015)
Schilling, K.G., et al.: Histological validation of diffusion MRI fiber orientation distributions and dispersion. Neuroimage 165, 200–221 (2018)
Schilling, K.G., et al.: Prevalence of white matter pathways coming into a single white matter voxel orientation: the bottleneck issue in tractography. Hum. Brain Mapp. 43(4), 1196–1213 (2022)
Shen, K., et al.: A spatio-temporal atlas of neonatal diffusion MRI based on kernel ridge regression. In: International Symposium on Biomedical Imaging (ISBI) (2017)
Skare, S., et al.: Condition number as a measure of noise performance of diffusion tensor data acquisition schemes with MRI. J. Magn. Reson. 147(2), 340–352 (2000)
Tournier, J.D., et al.: Direct estimation of the fiber orientation density function from diffusion-weighted MRI data using spherical deconvolution. Neuroimage 23(3), 1176–1185 (2004)
Tustison, N.J., Avants, B.B., Cook, P.A., Zheng, Y., et al.: N4ITK: improved N3 bias correction. IEEE Trans. Med. Imaging 29(6), 1310–1320 (2010)
Veraart, J., Fieremans, E., Novikov, D.S.: Diffusion MRI noise mapping using random matrix theory. Magn. Reson. Med. 76(5), 1582–1593 (2016)
Acknowledgment
This work was supported by the Swiss National Science Foundation (project 205321-182602). We acknowledge the CIBM Center for Biomedical Imaging, a Swiss research center of excellence founded and supported by CHUV, UNIL, EPFL, UNIGE, HUG and the Leenaards and Jeantet Foundations. This research was also partly supported by the US National Institutes of Health (NIH) under awards R01NS106030 and R01EB032366; by the Office of the Director of the NIH under award S10OD0250111; and by NVIDIA Corporation; and utilized an NVIDIA RTX A6000 GPU.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
1 Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Copyright information
© 2023 The Author(s), under exclusive license to Springer Nature Switzerland AG
About this paper
Cite this paper
Kebiri, H., Gholipour, A., Lin, R., Vasung, L., Karimi, D., Bach Cuadra, M. (2023). Robust Estimation of the Microstructure of the Early Developing Brain Using Deep Learning. In: Greenspan, H., et al. Medical Image Computing and Computer Assisted Intervention – MICCAI 2023. MICCAI 2023. Lecture Notes in Computer Science, vol 14226. Springer, Cham. https://doi.org/10.1007/978-3-031-43990-2_28
Download citation
DOI: https://doi.org/10.1007/978-3-031-43990-2_28
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-031-43989-6
Online ISBN: 978-3-031-43990-2
eBook Packages: Computer ScienceComputer Science (R0)