Abstract
Transient, tissue-specific, embryonic progenitors are important cell populations in vertebrate development. In the course of respiratory system development, multipotent mesenchymal and epithelial progenitors drive the diversification of fates that results to the plethora of cell types that compose the airways and alveolar space of the adult lungs. Use of mouse genetic models, including lineage tracing and loss-of-function studies, has elucidated signaling pathways that guide proliferation and differentiation of embryonic lung progenitors as well as transcription factors that underlie lung progenitor identity. Furthermore, pluripotent stem cell-derived and ex vivo expanded respiratory progenitors offer novel, tractable, high-fidelity systems that allow for mechanistic studies of cell fate decisions and developmental processes. As our understanding of embryonic progenitor biology deepens, we move closer to the goal of in vitro lung organogenesis and resulting applications in developmental biology and medicine.
You have full access to this open access chapter, Download chapter PDF
Similar content being viewed by others
Keywords
- Embryonic lung progenitors
- Organoid culture
- Cell similarity scores
- Gene regulatory networks
- Cell date decisions
1 Introduction
Tissue-specific embryonic progenitors are important gateway cell populations at various stages of organ development [49]. Within the endoderm germ layer, a substantial body of work has deciphered the emergence of domain-specific primordial progenitors, including signaling requirements for tissue specification, and has uncovered key transcription factors (TFs), such as Nkx2-1 for the lung and thyroid domains, Pdx1 for the pancreatic domain, and Cdx2 for the intestinal domain, for tissue specification or development or both [30, 52, 55]. Most importantly, self-renewing, transplantable, multipotent lung progenitors are of value in lung regenerative medicine; mechanistic understanding of their cell fate decisions will be key for their successful clinical application.
In this chapter, we review the biology of multipotent embryonic progenitors of the respiratory system and recent efforts to employ such progenitors for the modeling and study of cell fate decisions in ex vivo organotypic models of human lung development (Fig. 4.1). We also assess efforts to derive de novo lung progenitors of different developmental stages from pluripotent stem cells (PSCs). As the areas of mammalian developmental and PSC biology continuously inform each other [96], there is an increasing need for computational models to evaluate and quantify the similarity between embryonic and PSC-derived progenitors. We critically appraise the nascent literature on cell similarity models, including their limitations and underlying assumptions.
2 Overview of Embryonic Lung Progenitors
2.1 Stage-Specific Epithelial Progenitors (Primordial, Distal Tip, Basal)
The respiratory system is specified within the anterior ventral foregut endoderm (the anterior part of the gut tube) at around embryonic day (E)9.0 (18–22 somites) in mice and 4 weeks of gestation in humans [48, 102]. During subsequent stages (embryonic, pseudoglandular, canalicular, saccular, and alveolar), developmental processes, such as tracheoesophageal septation, formation and elongation of the primary lung buds, branching morphogenesis, and alveologenesis, lead to the anatomically and functionally distinct compartments of the adult lung that perform the basic functions of air conduction, mucociliary clearance, and gas exchange. These processes are concurrent with and inseparable from diversification of cell fates driven by stage-specific embryonic progenitors. Genetic alteration or ablation of such progenitors can lead to profound developmental defects, such as lung agenesis and mispatterning of the lung proximal-distal axis.
2.1.1 Lung Primordial Progenitors
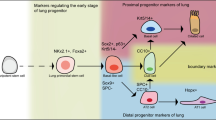
The lung epithelial primordial progenitors are the first epithelial cells within the developing gut tube to adopt a lung fate at around E9.0 in the mouse. The appearance of this rare, transient progenitor population is marked by expression of the homeodomain TF Nkx2-1 [48, 54]. Nkx2-1+ lung primordial progenitors are multipotent progenitors and give rise to the vast majority of lung epithelial lineages (both airway and alveolar), as shown by a lineage tracing study performed in our laboratory [48]. Although Nkx2-1 is not necessary for lung specification [55], it is the earliest marker of lung epithelial fate, and its expression has been used extensively to benchmark PSC-derived lung multipotent progenitors, as discussed below. Remarkably, ventx2, a Xenopus homeodomain TF, was recently shown to be expressed in the prospective respiratory domain at Nieuwkoop and Faber (NF)31/32 stage, earlier than Nkx2-1 (NF33/34), raising the tantalizing possibility that it is the earliest marker of respiratory (tracheal and lung) fate in Xenopus and potentially in humans [90].
Given the inaccessibility of the lung primordial population in humans, most information on the molecular pathways regulating its emergence and potential gene regulatory networks derives from mouse and Xenopus studies, both in vivo and ex vivo [15, 48, 89, 102]. Xenopus (Xenopus laevis, African clawed frog) has provided an excellent, complementary to mouse, model of foregut development, due to highly conserved endoderm formation, rapid development, and facile acquisition of large numbers of embryos, offering the possibility of medium-to-high-throughput loss-of-function studies [62, 89]. Mesoderm-derived Wnt signaling is indispensable for lung specification [32, 38], and Bmp signaling is necessary for correct spatial restriction of respiratory fated progenitors and tracheoesophageal separation [26]. Retinoic acid (RA) signaling confers lung competence to endodermal cells, which can then respond to Wnt and Bmp signals [87, 88]. The minimal signaling requirements for lung endodermal fate have also been demonstrated by pathway analysis between the transcriptomes of flow sorted lung primordial progenitors and prespecified endoderm, which demonstrated upregulation of Wnt and Tgf-β superfamily pathways in the former population [48].
In stark contrast to other Nkx2-1+ embryonic tissues, such as thyroid and forebrain [24, 97], little else is known regarding the gene regulatory program underlying the singular lung primordial identity. Apart from Nkx2-1, pan-endodermal TFs, such as Foxa2 [99], and TFs with known roles in later lung development, such as Irx2 and Foxp2 [36, 48], are also expressed in the lung primordium. Future work will certainly decipher TF interactions that uniquely define the lung primordial identity and are involved in early epithelial cell fate decisions in lung development.
2.1.2 Distal Tip Progenitors
Following respiratory system specification, its anterior-posterior axis is rapidly formed as witnessed by both morphological changes, such as trachea formation and elongation and emergence of primary lung buds, and mutually exclusive expression of the TFs Sox2 and Sox9 in the murine tracheal and lung domains [4]. During the ensuing pseudoglandular and canalicular stages, highly proliferating progenitors at the distal tips of the developing lung are essential for branching morphogenesis and formation of airway and alveolar epithelia [4, 81, 91]. Extensive work in mouse genetic models and, most recently, in human fetal organoids has shed light on the molecular profile of distal tip progenitors, their competence, and gene regulatory networks.
In mice, distal tip progenitors possess a distinct transcriptional profile with high expression of a constellation of TFs, such as Sox9, Id2, Nmyc, and Etv5 [4, 44, 61, 67, 77, 80, 91]; matrisomal genes such as Thbs1, Lama3, and Fbn2 [67, 81, 95]; and cell surface markers such as Clu [12, 81]. Interestingly, human distal tip progenitors also express SOX2, which is considered a proximal airway marker in murine development, during the pseudoglandular stage, but its distal expression is extinguished at the canalicular stage [22, 72, 78]. Lineage tracing studies have demonstrated that tip progenitors have a broad differentiation repertoire during the pseudoglandular stage (as early as E11.5), giving rise to major lung epithelial lineages (club, ciliated, neuroendocrine, and alveolar type I (AT1) and type II (AT2) cells) [91], while at later time points (E16.5) their differentiation potential is restricted to alveolar epithelial types. Nevertheless, late tip progenitors appear to possess some degree of plasticity since they can give rise to airway progeny when grafted in cultured E12.5 lung explants [61]. Further insights in the developmental potential of distal tip progenitors have been derived by combining RNA-sequencing (RNA-seq) with assay for transposase-accessible chromatin using sequencing (ATAC-seq) at two developmental time points, E11.5 and E16.5 [53]. Remarkably, almost 25% of open chromatin regions were specific to each time point, probably reflecting the time-dependent competence of tip progenitors. This integrated analysis identified increased expression and chromatin accessibility of PI3K signaling pathway components at the later point, and inhibition of this pathway at an early time point increased the percentage of SOX9+/NKX2-1+ lung epithelial progenitors.
Integration of several pathways, such as β-catenin, Fgf, and Bmp signaling, ensure the proliferation and maintenance of the distal tip progenitor identity, as has been demonstrated in numerous genetic mouse studies. β-catenin-dependent Wnt signaling, which is indispensable for lung specification, is also required to maintain distal tip progenitors, by controlling Sox9 and Nkx2-1 expression and acting downstream of Fgf/Kras signaling [81]. Precise modulation of β-catenin levels is critical for respiratory identity of tip progenitors, as either deletion of hyperactivation of β-catenin leads to the ectopic expression of gastrointestinal or liver genes within the developing respiratory epithelium [79, 81]. Glucocorticoid signaling, which is necessary for differentiation of AT2 cells [31, 105], also dictates, at late stages of respiratory development, the formation of proximal-distal boundary, namely, the bronchoalveolar duct junction (BADJ). Premature activation or deletion of glucocorticoid signaling leads to proximal or distal shift of the BADJ, respectively [4]. Glucocorticoid signaling and STAT3 signaling, activated by LIF and other IL6 family ligands, can act in concert to promote alveolar differentiation of distal tip progenitors from the canalicular stage [61].
Murine distal tip progenitors have a critical role in lung epithelial differentiation and morphogenesis. Information gleaned from developmental studies have propelled the development of lung-directed differentiation protocols from human and mouse PSCs, and ex vivo organoid culture of such human fetal progenitors has provided valuable insights in similarities and differences in respiratory development between the two species, as detailed below.
2.1.3 Airway Basal Cells
An important stem cell population in the adult respiratory system are P63+/KRT5+ airway basal cells that can self-renew and differentiate to a wide variety of proximal cell types, such as secretory, ciliated, tuft, ionocyte, and neuroendocrine [6, 93]. There is a clear species difference in basal cell distribution as basal cell-containing pseudostratified epithelium is limited in the trachea and mainstem bronchi in mice but extends more distally spanning several airway generations in humans [94]. Recent work has shed light to the developmental origins of this population [73, 112]. Intriguingly, dispersed P63+ cells are already present in the Nkx2-1GFP+ lung primordium (around E9.0–9.5) and can give rise to cells in both the airway and alveolar compartments [112]. Epithelial P63+ cells are rapidly restricted to the tracheal domain by E10.5 with concomitant lineage restriction to tracheal and proximal airways. Lineage tracing studies at later time points indicate that the pool of embryonic progenitors to E18.5 p63+Krt8- prebasal cells is established as early as E13.5–E14.5, as cells labeled at this time point comprise more than 80% of perinatal prebasal cells [112]. ITGB4hi multipotent progenitors have been described in early developing trachea (E14.5), but it is not clear to which extent these progenitors overlap with the P63+ progenitor pool [11]. In human fetal lungs, P63+/KRT5+ cells, considered bona fide basal cells, are present in trachea and mainstem bronchi by 12 weeks [73]. Interestingly, P63+/KRT5- in non-cartilaginous and small-cartilaginous airways have nuclear pSMAD staining, and they may be the human equivalent of basal cell progenitors described in the study of Yang et al. [112].
2.2 Stage-Specific Mesenchymal Progenitors
Mesenchymal-epithelial interactions are indispensable at every stage of lung development, from lung epithelial specification to branching morphogenesis. For example, different degrees of disruption of endodermally derived Shh signaling to splanchnic mesoderm lead to severe developmental effects, such as lung hypoplasia and tracheoesophageal fistula or even lung, tracheal, and esophageal agenesis [66, 74]. On the other hand, FGF10 expressed in the lung epithelium-adjacent mesoderm is important for bud induction [15], whereas, at later stages, mesenchymal FGF10 expression in the vicinity of distal tips is part of a complex signaling circuit that regulates tip outgrowth and proliferation [10, 63].
As the lung develops, the mesenchyme is also subject to a diversification of fates, leading to the various mesoderm-derived cell populations in the adult lung, such as vascular and airway smooth muscle cells, endothelial cells, pericytes, and pleura. Lineage tracing studies alone or in combination with new high-resolution genomic methods, such as single-cell RNA-seq (scRNA-seq), have allowed for the characterization of various mesenchymal progenitor populations and their developmental potential, as they have been comprehensively reviewed in Riccetti et al. [92]. For example, lineage tracing using different gene drivers has shown that several lung mesenchymal lineages, such as vascular and airway smooth muscle cells, proximal endothelial cells, and Pdgfrβ+ pericyte-like cells, originate from Wnt2+/Gli1+/Isl1+ cardiopulmonary progenitors [82]. Systems that employ Tbx4 lung-specific enhancers have also been described and used for lineage tracing (as Cre or rtTA drivers) of mesenchymal progenitor progeny at various windows during lung development [59, 115].
Zorn et al. has recently used scRNA-seq to describe signaling networks governing interactions between mesenchymal and epithelial domains in early foregut development (E8.5–E9.5) [36]. This study revealed several mesenchymal cell clusters corresponding to domain-specific mesenchymal populations that engage in signaling crosstalk with the corresponding epithelial populations. In the developing lung mesenchyme, there was expression of previously described lung mesenchymal markers, such as Wnt2 [32], Osr1 [86], Foxf1 [19, 84], and Tbx5 [13]. Nkx6-1, a known pancreatic epithelial TF, was also expressed in the respiratory mesenchymal domain (both by transcript and protein), and it was one of the key TFs on the cell-state trajectory from foregut lateral mesoderm to lung mesenchyme [36].
Similar transcriptomic characterization of the developing human lung mesenchyme was performed using fetal human tissue from various endodermal organs, including the lung, from 7 to 21 weeks postconception [113]. As predicted, human lung mesenchyme was quite different than intestinal and stomach mesenchyme, with WNT2 being one of the most differentially expressed genes. The TF PRRX1 marked chondrocyte-like cells and pericytes in the lung. Furthermore, distal tip mesenchymal progenitors were also identified using human lung fetal tissue from pseudoglandular to canalicular stages [43]. These progenitors were RSPO2+ and could be purified based on LIFR expression. The RSPO2+ progenitors were found to maintain the distal tip undifferentiated state by epithelial LGR5-mediated Wnt signaling [43].
3 Ex Vivo Culture of Multipotent Embryonic Lung Progenitors
As discussed in the previous section, use of mouse models has been instrumental in our understanding of lung embryonic progenitors, including their differentiation repertoire and signaling pathways that control their specification and self-renewal. Embryonic explant culture has provided another powerful tool to study early lung development, due to ease of manipulation of the soluble environment (including activators and inhibitors of developmental pathways) and molecular characterization of the explants [15, 16, 25, 61]. Recently, lung developmental studies have received considerable impetus by the ex vivo expansion and differentiation of fetal respiratory progenitors that are phenotypically and functionally similar to their in vivo counterparts. Ex vivo progenitor organoids combined with state-of-the-art technologies, such as multimodal single-cell profiling and clustered regularly interspaced short palindromic repeats (CRISPR)-mediated gene editing [50], are already enabling high-resolution studies of lung embryonic progenitor biology.
3.1 Ex Vivo Culture of Mouse Embryonic Progenitors
As embryonic progenitors are transient populations that exist within defined developmental windows, their capture and ex vivo expansion and multilineage differentiation offer a powerful, tractable system to study developmental questions and develop organoid platforms. While ex vivo culture of endodermal progenitors, such as pancreatic and hepatic progenitors, has been reported for some time now [35, 109], only recently has culture of respiratory progenitors been established. Nichane and coworkers were able to purify mouse Sox9+ distal tip progenitors using a Sox9 fluorescent reporter strain and develop defined culture conditions (lung progenitor medium, LPM) for their ex vivo expansion [77]. 3D culture in Matrigel, activation of Wnt signaling by GSK-3β inhibition (CHIR99021), inhibition of Tgf-β and p38 signaling, and addition of Fgf (FGF9, FGF10) ligands and EGF were necessary to maintain E12.5 tip progenitors in an undifferentiated, multipotent state, with high expression of known tip markers, such as Id2, Nmyc, Etv5, and Sox9. Importantly, ex vivo expanded progenitors were able to differentiate to airway and alveolar lineages both in vitro and after transplantation to the mouse lungs following naphthalene or bleomycin injury [77]. Interestingly, the independent and contemporaneous work of Spence et al. identified FGF7, CHIR99021, and RA (3F medium) as the minimal signaling requirements for ex vivo maintenance of distal tip progenitor cells from E12.5 mouse lungs [72]. A similar system established earlier in lung development (E11.5) was employed for a genetic screen of Sox9+ tip progenitor regulators [2]. This screen identified Aurkb, among others, as a regulator of tip progenitor cell cycle, and its relevance was confirmed by in vivo deletion in distal tip progenitors, leading to profound defects in branching morphogenesis.
In our work, we were able to isolate rare E9.0 lung primordial epithelial progenitors using the previously published Nkx2-1GFP reporter mouse [48, 68]. scRNA-seq of in vivo and ex vivo expanded lung primordial progenitors guided modifications of the distal tip expansion medium, thus allowing us to successfully culture primordial progenitors that proliferated and maintained Nkx2-1GFP expression for several passages [64]. This ex vivo system can be used to study cell fate decisions during the embryonic stage of lung development.
3.2 Ex Vivo Culture of Human Embryonic Progenitors
The need for ex vivo systems is even more salient with human fetal progenitors due to the relative inaccessibility of human fetal tissue and severe restrictions on related research in certain regions [3]. The possibility of culturing ex vivo human distal tip progenitors was first shown by Rawlins et al. [78]. Use of a cocktail of growth factors (EGF, FGF7, FGF10), Tgf-β (SB431542) and BMP (Noggin) inhibitors, and Wnt activators (CHIR99021) and potentiators (R-spondin 1) resulted in long-term culture of mesenchyme-depleted tip progenitors from 5- to 9-week fetal lungs. There was absolute requirement for activation of FGF and Wnt and inhibition of Tgf-β signaling for the self-renewal of the progenitors. The multilineage differentiation potential of the ex vivo expanded progenitors was convincingly demonstrated in a series of both in vitro (PneumaCult for bronchiolar differentiation, alveolar differentiation medium with expanded human lung fetal fibroblasts) and in vivo (kidney capsule co-transplantation with E13.5 mouse fetal cells) assays. Miller et al. showed that the 3F medium (FGF7, CHIR99021, RA) described for the expansion of the mouse distal tip progenitors, as well as the 4-factor medium (FG7, FGF10, CHR99021, RA), were sufficient for the ex vivo stable culture of 12-week dissected human lung distal tip progenitors [72]. The expanded progenitors had low expression of airway markers, such as P63, FOXJ1, and SCGB1A1, and high co-expression of tip markers, such as SOX9 and SOX2. Notably, expanded progenitors had significant transcriptional similarity with freshly isolated fetal lung tips. Although BMP4 removal resulted in higher SOX9 expression in Miller et al. [72] similarly to BMP inhibition in Nikolic et al. [78], the absence of Tgf-β inhibition requirement in the former protocol is intriguing, and it may be explained by subtle differences in the two systems due to different developmental times of the tissue of origin, including endogenous TGF-β levels.
Establishing tractable systems of human fetal lung progenitors opens up the exciting possibility of ex vivo human development studies, as shown by recent work of Rawlins et al. and Spence et al. [18, 65, 100, 101]. Engineering a CRISPRi system in human distal tip progenitors, Sun and coworkers were able to perform loss-of-function studies and systematically screen the role of several TFs in self-renewal of SOX9+ progenitors [100]. For example, MYBL2 (proliferation-related gene) and CTNNB1 (Wnt signaling effector) guide RNAs were highly depleted in a 2-week culture of organoids, indicating a role of these genes in progenitor self-renewal. Knockdown of SOX9 resulted in downregulation of 455 common genes in two organoid lines, and further gain-of-function and genomic occupancy studies identified ETV4, ETV5, and NMYC and Wnt-related genes, such as LGR5 and CD44, as direct transcriptional SOX9 targets. Removal of Wnt activators from the tip medium resulted in SOX9 expression loss. Interestingly, genes suppressed by SOX9 included secretory cell-related genes, as well as non-lung (stomach, liver) endodermal genes, similarly to mouse respiratory development [81].
Human distal tip progenitors from a later (canalicular) stage were used to study alveolar differentiation due to their restricted differentiation potential [65]. CD36 was specifically expressed in these progenitors and along with CD44 marks distal tip progenitors from 16- to 21-week human fetal lungs. CD36/CD44 co-expression has also functional significance as CD36+CD44+ (LinPos) organoids had greater propensity to differentiate to alveolar cells. In this system, NKX2-1 and TFAP2C were found to regulate opposing fates, with the former TF promoting alveolar and the latter promoting airway (basal) fate, as shown by gain-of-function studies.
Distal tip organoids have been also used to interrogate lineage relationships, focusing on newly identified progenitors [18]. In scRNA-seq of the developing human lung, SCGB3A2/SFTPB/CFTR co-expression identified a progenitor cell, named fetal airway secretory (FAS) cell. FAS cells emerged during airway differentiation [72] of distal tip progenitors and were shown, following barcoding-mediated lineage tracing, to give rise to pulmonary neuroendocrine cells and a subset (C6+) of multiciliated cells.
Overall, culture of lung fetal progenitors has offered important insights in lung epithelial development and provided sophisticated systems that are being used to study the intricate relationships between progenitor competence, gene regulatory networks, and signaling pathways.
4 In Vitro Derivation of Multipotent Embryonic Lung Progenitors
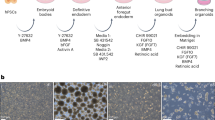
A large body of work reviewed recently [29, 104] has led to the development of directed differentiation protocols for the derivation of a variety of lung lineages from both mouse and human PSCs. It is now well established that derivation of lung-competent anterior foregut endoderm via dual Smad inhibition followed by Wnt and Bmp signaling can generate lung multipotent primordial-like progenitors in vitro [27, 33, 34, 39, 47, 48, 68, 75]. These progenitors can be highly purified by the use of either knock-in fluorescent reporters, such as eGFP and mCherry, or cell surface marker sorting algorithms, such as CPMhi and CD47hi/CD26lo [33, 39, 48, 68, 98, 110]. Interestingly, Cd26 (Dpp4) is one of the most downregulated genes upon in vivo lung specification in mice [48]. Wnt signaling is also important post-specification in vitro with timing of Wnt addition or withdrawal leading to different differentiation trajectories (airway or alveolar) [23, 51, 58, 71].
As PSC-based systems have been further refined, one major goal is to derive and stably expand in culture other important lung epithelial and mesenchymal progenitors. Earlier efforts had led to the long-term culture of complex PSC-derived organoids that initially contained tip progenitors and appeared to recapitulate the transcriptional profile of second-trimester human lung [17]. At the same time, there is an imperative need to develop computational methods to reliably benchmark such populations against their presumed in vivo counterparts or ex vivo expanded embryonic progenitors, as discussed in the following section. Creation of human or mouse developmental atlases at single-cell resolution [41, 73, 76, 113] can satisfy this need while also driving the study of developmental trajectories and cell fate decisions.
Hawkins and coworkers reported the derivation and isolation of airway basal-like cells (iBCs) from human PSCs using a bi-fluorescent (NKX2-1GFP; TP63tdTomato) reporter cell line [40]. These cells were expandable in airway media (PneumaCult-Ex) containing dual Smad inhibitors and rapidly acquired and maintained expression of the airway basal cell marker, NGFR [93]. Furthermore, their airway multilineage potential (ciliated, secretory, goblet) was demonstrated in both in vitro air-liquid interface (ALI) culture and in tracheal xenografts.
The derivation of stable distal tip-like progenitors from PSCs has also been reported [42, 45]. Refinement of a previously published protocol for the derivation of mouse lung primordial progenitors [48] and subsequent use of the LPM medium [77] led to the stable expansion of a progenitor population that maintained Nkx2-1mCherry expression and also expressed distal tip markers (Sox9, Id2) [45]. Remarkably, these PSC-derived progenitors were able to engraft (up to 15 weeks) in the distal lung, following transplantation into bleomycin-injured, syngeneic, immunocompetent recipients. The transplanted progenitors differentiated into AT1- and AT2-like cells that were highly similar to their endogenous counterparts as evidenced by protein expression of mature markers, such as PDPN and proSFTPC, and high similarity scores using a novel scRNA-seq-based computational method (single-cell type order parameters, scTOP) [45, 111]. Similarly, human PSC-derived lung progenitors adopted a bud tip fate in a previously described medium [72] and were expanded over several weeks as bud tip organoids (iBTOs) [42]. These progenitors express several markers of human pseudoglandular stage distal tip cells (SOX9, ETV5, ID2, HMGA2), had high similarity computational scores to culture-expanded primary human distal tip lung progenitors, and were able to differentiate to either alveolar or airway cells in respective media.
The value of PSC-based systems as a discovery/validation tool for hitherto unknown progenitor populations was evident in two publications [8, 70]. McCauley et al. used a human PSC reporter line to derive and isolate SCGB3A2+ early secretory progenitors. By using scRNA-seq, a Wnt-dependent subpopulation of these progenitors was found to co-express an AT2-like program, including genes such as SFTPC, NAPSA, and PGC, and even possess functional lamellar bodies. This in vitro engineered population was further validated in Basil et al. [8] and appears to have high similarity with the newly discovered SCGB3A2+ respiratory airway secretory (RAS) cells in human terminal bronchioles, which are progenitors to AT2 cells and whose alveolar differentiation is regulated by Notch and Wnt signals.
While lung epithelial progenitors are routinely derived from PSCs, derivation of multipotent lung mesenchymal progenitors has lagged behind due to the early developmental broad expression of several mesenchymal TFs and dearth of information on the signals specifying lung mesenchyme. Recently, several papers have reported efforts to derive functional multipotent lung mesenchyme from either human or mouse PSCs [5, 36, 56]. In a proof-of-principle study, Han and coworkers gleaned information from their study of coordinated epithelial-mesenchymal development within the early foregut endoderm to derive distinct subtypes of tissue-specific mesenchyme, including lung mesenchyme, from human PSCs [36]. Following derivation of lateral plate mesoderm, they used BMP4, FGF2, and RA with simultaneous inhibition of Wnt and Tgf-β signaling and activation of Shh signaling by purmorphamine (PMA), a Shh signaling agonist, to induce an anterior foregut splanchnic mesoderm fate. Further activation of BMP, Shh, and RA signaling with activation of Wnt at later stages led to the derivation of lunglike mesenchyme, based on co-expression of NKX6-1, TBX5, FOXF1, and WNT2 [57].
Similar work of Morimoto et al. identified β-catenin-dependent Wnt signaling as an important signal for early lung mesodermal Tbx4 expression in the tracheal domain [56]. Lateral plate mesoderm (Foxf1+) was treated with BMP4 and CHIR99021 to produce tracheal mesoderm from mouse PSCs, and the downstream differentiation to chondrocytes and smooth muscle cells of the latter was demonstrated with immunostaining and qPCR. Induction of tracheal mesoderm with chondrocyte and smooth muscle competence from human PSCs required further addition of PMA.
Alber and coworkers made heavy use of a Tbx4 enhancer Cre mouse PSC line (Tbx4-rtTA; TetO-Cre; mTmG; Tbx4-LER) to indelibly mark and purify PSC-derived lung-specific mesenchyme (iLM) [5, 115]. KDR+ lateral plate mesoderm (also Foxf1+) was treated with various combinations of Wnt, Bmp4, PMA, and RA, and it was determined that RA and PMA were the minimal signaling requirements for derivation of lung-specific mesenchyme, based on the percentage of Tbx4-LERGFP+ cells [5]. The functionality of the engineered iLM was demonstrated by the following: (a) enhanced lung epithelial differentiation in organoid culture with PSC-derived lung progenitors, (b) induction of lung fate (Nkx2-1+ cells) in non-lung epithelial progenitors, and (c) maintenance of primary AT2 cells in organoid coculture. Tbx4-LERGFP+ iLM was also shown to be multipotential and was able to differentiate to smooth muscle cells, adipocytes, and mesenchymal alveolar niche cells (MANCs) [114].
5 Progenitor Cell Similarity Models
A major challenge facing the field is to develop tractable computational methods for assessing cell similarity between engineered and natural cells. An ideal method would assess similarity along multiple phenotypic axes: transcriptomic, epigenetic, metabolomic, etc. [28, 37]. However, such multimodal comparisons remain uncommon due to the difficulty and expense of generating the required datasets. For this reason, the vast majority of cell similarity methods restrict themselves to analyzing transcriptomic measurements made using RNA-seq methods. RNA-seq can occur in bulk, where a multitude of cells in a tissue are sequenced simultaneously, or in single-cell form (scRNA-seq), where individual cells are sequenced. The latter, although more expensive and resource intensive, provides granular resolution of the gene expression of individual cells and has become standard for identifying cell types. For this reason, here, we focus on scRNA-seq-based computational methods for understanding cell similarity (compared and reviewed in Abdelaal et al. [1] and Xie et al. [108]).
Any scRNA-seq-based method for quantifying cell similarity must account for several technical challenges [46]. Single-cell data tends to be extremely sparse, with many low-expressing genes being measured as zero expression (a phenomenon often called “dropout” in the computational literature). The noisy nature of scRNA-seq data is exacerbated by the large amount of cell-to-cell technical variation. These problems become especially acute when comparing data across experiments and biological conditions, where batch effects can dominate variation in scRNA-seq data, making it difficult to assess cell similarity, especially in the context of closely related cell types [83]. This presents a major technical hurdle for utilizing the comprehensive cell atlases now being generated by the community [69, 103].
To address the noisy nature of scRNA-seq, numerous computational methods have been proposed (many of which are collected in the now indispensable Bioconductor package) [7]. To deal with the missing zeroes and data sparsity in scRNA-seq datasets, analysis pipelines often include methods for data imputation (see Dai et al. [20] for a recent comparison of imputation methods). Although data imputation methods vary in their technical details (model-dependent or model-free deep learning methods), they all make use of the expression profiles of similar cells to impute missing values. This process tends to homogenize expression profiles across similar cells, making it difficult to assess small differences in gene expression profiles between cells. Most importantly, for our purposes, this homogenization can lead to inflated cell similarity scores, giving rise to an overly optimistic assessments of similarity between engineered and natural cell types. This general phenomenon is also true of commonly used batch correction methods, which decrease variation across conditions or experiments and thus can conflate technical variability with biological variability [69]. For this reason, great care must be taken when using imputation and batch correction methods in analysis pipelines whose goal is comparing cell similarity.
Perhaps the biggest challenge for assessing cell similarity is the high-dimensional nature of scRNA-seq data, with RNA from thousands of genes being measured simultaneously in individual cells. High dimensionality requires the use of specialized methods for visualization and interpretation. A typical data analysis pipeline [7] for discovering and classifying cell types involves applying a dimensional reduction algorithm to summarize the information provided by thousands of genes down to just two dimensions (e.g., using a dimensional reduction method such as SPRING [106] or uniform manifold approximation and projection (UMAP) [9], followed by clustering, and the utilization of marker genes to assign clusters to cell types. Many analysis pipelines also restrict the features being analyzed to a smaller subset, such as highly variable genes. Finally, to assess cell similarity of the new cell, the expression profile of the new cell is compared to expression profiles of the clusters that are used as a proxy for cell type.
A powerful feature of this pipeline is that it allows for easy visualization and outputs biologically interpretable clusters. This allows experimentalists to quickly assess experiments, identify biologically interesting features, and develop novel biological hypothesis that can be tested in new experiments. Given the widespread adoption, it is worth highlighting a number of limitations of this analysis pipeline for assessing cell similarity. First, both the feature selection and the dimensional reduction depend on the datasets that are included in the analysis. The inclusion or exclusion of different datasets generally yields different results even for the same experiment. This dependence on choice of data is especially important when trying to assess cell similarity between closely related cell types or comparing natural and engineered cells. Second, dimensional reduction techniques are primarily useful for visualization and almost always fundamentally distort biological patterns that occur in the full high-dimensional space [14]. In general, axes and distances in a UMAP or SPRING plot have no natural biological interpretation, making it challenging to rigorously interpret biological experiments.
To address these limitations, the authors have developed a novel computational method for computing cell similarity at the single-cell level (single-cell type order parameters, scTOP) [45, 111]. scTOP utilizes insights developed from statistical physics-inspired epigenetic landscape models to assess cell similarity in a holistic manner [21, 48, 60, 85]. The inputs to scTOP are a “cell basis,” consisting of the transcriptomic profiles of cell types of interest (e.g., the expression profiles of lung cells in a cell atlas, such as the single-cell lung cell atlas) and the scRNA-seq measurements of the cell being analyzed (e.g., an engineered cell whose cell similarity we wish to compute). The output of scTOP is a set of cell similarity scores that characterize the cell similarity with each cell type in the “cell basis” (e.g., a score to be a club cell, AT1 cell, AT2 cell, etc.). An appealing feature of the scTOP algorithm is that it allows for biologically meaningful visualizations and sensitive and accurate classification of cell similarity using scRNA-seq data [45]. Importantly, the method does not require any dimensional reduction techniques, feature selection, produces visualizations with meaningful axes, and does not depend on what other data is being analyzed. One limitation of scTOP is that it requires experimentalists to specify cell types of interest ahead of time through the choice of cellular basis. For this reason, scTOP cannot be used to assess or discover novel cell types. Nonetheless, we have found scTOP to be an excellent tool for comparing engineered and natural cells, where the cells of interest are often known [111].
Assessing cell similarity of progenitor populations is an especially computationally challenging task [108]. Transcriptome profiles of developmental progenitor cell types are generally more similar to each other than their adult counterparts, highlighting the need to collect scRNA-seq data on multiple progenitor populations. Furthermore, during the course of development, progenitor cell populations often rapidly change their transcriptome profiles as they differentiate, making it difficult to accurately measure transcriptomes in vivo.
This has important implications for thinking about lung progenitors at various developmental stages. Even when in vivo progenitors can be stabilized and expanded ex vivo, their transcriptomes can be altered due to radical differences in in vivo and ex vivo environments [42]. Hence, it is important to decide how to benchmark in vitro engineered cells, and characterization of progenitor-based organoids should extend beyond similarity scores to encompass functional differentiation, global chromatin accessibility, and proteomics.
6 Conclusion
As our understanding of lung multipotent progenitor biology deepens, the range of potential applications increases (Fig. 4.2). Ex vivo or engineered PSC-derived lung organoids have already been deployed for modeling of cell fate decisions and production of cells of clinical relevance. Tissue interactions and morphogenetic processes have been long-standing areas of interest in developmental biology [107], and one could expect modern-day organoids to be increasingly used to similar ends. Other future applications include respiratory disease modeling and the study of lung-microbiome interactions, thus reducing the need for animal experimentation. Ultimately, lung progenitor-based organoids will become widely and routinely used tools in medicine and developmental biology.
Change history
11 August 2023
A correction has been published.
References
Abdelaal, T., Michielsen, L., Cats, D., Hoogduin, D., Mei, H.L., Reinders, M.J.T., and Mahfouz, A. (2019). A comparison of automatic cell identification methods for single-cell RNA sequencing data. Genome Biol. 20, 194. https://doi.org/10.1186/s13059-019-1795-z.
Ah-Cann, C., Wimmer, V.C., Weeden, C.E., Marceaux, C., Law, C.W., Galvis, L., Filby, C.E., Liu, J., Breslin, K., Willson, T., et al. (2021). A functional genetic screen identifies aurora kinase b as an essential regulator of Sox9-positive mouse embryonic lung progenitor cells. Development 148, dev199543. https://doi.org/10.1242/dev.199543.
Al Alam, D., Ballard, P.L., Le Saux, C.J., Ingram, J.L., Mariani, T.J., Moore, N.S., Weiss, D.J., and Yu, M. (2021). Human Fetal Tissue Regulation Impact on Pediatric and Adult Respiratory-related Research. Ann. Am. Thoracic Society 18, 204–208. https://doi.org/10.1513/AnnalsATS.202005-460PS.
Alanis, D.M., Chang, D.R., Akiyama, H., Krasnow, M.A., and Chen, J.C. (2014). Two nested developmental waves demarcate a compartment boundary in the mouse lung. Nat. Commun. 5, 3923. https://doi.org/10.1038/ncomms4923.
Alber, A.B., Marquez, H.A., Ma, L., Kwong, G., Thapa, B.R., Villacorta-Martin, C., Lindstrom-Vautrin, J., Bawa, P., Luo, Y., Ikonomou, L., et al. (2022). Directed differentiation of mouse pluripotent stem cells into functional lung-specific mesenchyme. bioRxiv, 2022.2008.2012.502651. https://doi.org/10.1101/2022.08.12.502651.
Alysandratos, K.D., Herriges, M.J., and Kotton, D.N. (2021). Epithelial Stem and Progenitor Cells in Lung Repair and Regeneration. In Annual Review of Physiology, Vol 83, M.T. Nelson, and K. Walsh, eds. (Annual Reviews), pp. 529–550. https://doi.org/10.1146/annurev-physiol-041520-092904.
Amezquita, R.A., Lun, A.T.L., Becht, E., Carey, V.J., Carpp, L.N., Geistlinger, L., Marini, F., Rue-Albrecht, K., Risso, D., Soneson, C., et al. (2020). Orchestrating single-cell analysis with Bioconductor. Nat. Methods 17, 137–145. https://doi.org/10.1038/s41592-019-0654-x.
Basil, M.C., Cardenas-Diaz, F.L., Kathiriya, J.J., Morley, M.P., Carl, J., Brumwell, A.N., Katzen, J., Slovik, K.J., Babu, A., Zhou, S., et al. (2022). Human distal airways contain a multipotent secretory cell that can regenerate alveoli. Nature 604, 120–126. https://doi.org/10.1038/s41586-022-04552-0.
Becht, E., McInnes, L., Healy, J., Dutertre, C.A., Kwok, I.W.H., Ng, L.G., Ginhoux, F., and Newell, E.W. (2019). Dimensionality reduction for visualizing single-cell data using UMAP. Nat. Biotechnol. 37, 38–44. https://doi.org/10.1038/nbt.4314.
Bellusci, S., Grindley, J., Emoto, H., Itoh, N., and Hogan, B.L.M. (1997). Fibroblast growth factor 10(FGF10) and branching morphogenesis in the embryonic mouse lung. Development 124, 4867–4878.
Bilodeau, M., Shojaie, S., Ackerley, C., Post, M., and Rossant, J. (2014). Identification of a Proximal Progenitor Population from Murine Fetal Lungs with Clonogenic and Multilineage Differentiation Potential. Stem Cell Rep. 3, 634–649. https://doi.org/10.1016/j.stemcr.2014.07.010.
Chang, D.R., Alanis, D.M., Miller, R.K., Ji, H., Akiyama, H., McCrea, P.D., and Chen, J.C. (2013). Lung epithelial branching program antagonizes alveolar differentiation. Proc. Natl. Acad. Sci. U. S. A. 110, 18042–18051. https://doi.org/10.1073/pnas.1311760110.
Chapman, D.L., Garvey, N., Hancock, S., Alexiou, M., Agulnik, S.I., GibsonBrown, J.J., CebraThomas, J., Bollag, R.J., Silver, L.M., and Papaioannou, V.E. (1996). Expression of the T-box family genes, Tbx1-Tbx5, during early mouse development. Dev. Dyn. 206, 379–390.
Chari, T., Banerjee, J., and Pachter, L. (2021). The Specious Art of Single-Cell Genomics. bioRxiv, 2021.2008.2025.457696. https://doi.org/10.1101/2021.08.25.457696.
Chen, F., Cao, Y.X., Qian, J., Shao, F.Z., Niederreither, K., and Cardoso, W.V. (2010). A retinoic acid-dependent network in the foregut controls formation of the mouse lung primordium. J. Clin. Invest. 120, 2040–2048. https://doi.org/10.1172/jci40253.
Chen, F., and Cardoso, W.V. (2015). Culture of Mouse Embryonic Foregut Explants. In Tissue Morphogenesis: Methods and Protocols, C.M. Nelson, ed. (Humana Press Inc), pp. 163–169.
Chen, Y.W., Huang, S.X., de Carvalho, A., Ho, S.H., Islam, M.N., Volpi, S., Notarangelo, L.D., Ciancanelli, M., Casanova, J.L., Bhattacharya, J., et al. (2017). A three-dimensional model of human lung development and disease from pluripotent stem cells. Nat. Cell Biol. 19, 542–549. https://doi.org/10.1038/ncb3510.
Conchola, A.S., Frum, T., Xiao, Z., Hsu, P.P., Hein, R.F.C., Miller, A., Tsai, Y.-H., Wu, A., Kaur, K., Holloway, E.M., et al. (2022). Distinct airway progenitor cells drive epithelial heterogeneity in the developing human lung. bioRxiv, 2022.2006.2013.495813. https://doi.org/10.1101/2022.06.13.495813.
Costa, R.H., Kalinichenko, V.V., and Lim, L. (2001). Transcription factors in mouse lung development and function. Am. J. Physiol.-Lung Cell. Mol. Physiol. 280, L823–L838.
Dai, C.C., Jiang, Y., Yin, C.L., Su, R., Zeng, X.X., Zou, Q., Nakai, K., and Wei, L.Y. (2022). scIMC: a platform for benchmarking comparison and visualization analysis of scRNA-seq data imputation methods. Nucleic Acids Res. 50, 4877–4899. https://doi.org/10.1093/nar/gkac317.
Dame, K., Cincotta, S., Lang, A.H., Sanghrajka, R.M., Zhang, L.Y., Choi, J.Y., Kwok, L., Wilson, T., Kandula, M.M., Monti, S., et al. (2017). Thyroid Progenitors Are Robustly Derived from Embryonic Stem Cells through Transient, Developmental Stage-Specific Overexpression of Nkx2-1. Stem Cell Rep. 8, 216–225. https://doi.org/10.1016/j.stemcr.2016.12.024.
Danopoulos, S., Alonso, I., Thornton, M.E., Grubbs, B.H., Bellusci, S., Warburton, D., and Al Alam, D. (2018). Human lung branching morphogenesis is orchestrated by the spatiotemporal distribution of ACTA2, SOX2, and SOX9. Am. J. Physiol.-Lung Cell. Mol. Physiol. 314, L144-L149. https://doi.org/10.1152/ajplung.00379.2017.
de Carvalho, A., Strikoudis, A., Liu, H.Y., Chen, Y.W., Dantas, T.J., Vallee, R.B., Correia-Pinto, J., and Snoeck, H.W. (2019). Glycogen synthase kinase 3 induces multilineage maturation of human pluripotent stem cell-derived lung progenitors in 3D culture. Development 146, dev171652. https://doi.org/10.1242/dev.171652.
De Felice, M., and Di Lauro, R. (2011). Minireview: Intrinsic and Extrinsic Factors in Thyroid Gland Development: An Update. Endocrinology 152, 2948–2956. https://doi.org/10.1210/en.2011-0204.
Desai, T.J., Malpel, S., Flentke, G.R., Smith, S.M., and Cardoso, W.V. (2004). Retinoic acid selectively regulates Fgf10 expression and maintains cell identity in the prospective lung field of the developing foregut. Dev. Biol. 273, 402–415. https://doi.org/10.1016/j.ydbio.2004.04.039.
Domyan, E.T., Ferretti, E., Throckmorton, K., Mishina, Y., Nicolis, S.K., and Sun, X. (2011). Signaling through BMP receptors promotes respiratory identity in the foregut via repression of Sox2. Development 138, 971–981. https://doi.org/10.1242/dev.053694.
Dye, B.R., Hill, D.R., Ferguson, M.A.H., Tsai, Y.H., Nagy, M.S., Dyal, R., Wells, J.M., Mayhew, C.N., Nattiv, R., Klein, O.D., et al. (2015). In vitro generation of human pluripotent stem cell derived lung organoids. eLife 4, 25. https://doi.org/10.7554/eLife.05098.
Efremova, M., and Teichmann, S.A. (2020). Computational methods for single-cell omics across modalities. Nat. Methods 17, 14–17. https://doi.org/10.1038/s41592-019-0692-4.
Frum, T., and Spence, J.R. (2021). hPSC-derived organoids: models of human development and disease. J. Mol. Med. 99, 463–473. https://doi.org/10.1007/s00109-020-01969-w.
Gao, N., White, P., and Kaestner, K.H. (2009). Establishment of Intestinal Identity and Epithelial-Mesenchymal Signaling by Cdx2. Dev. Cell 16, 588–599. https://doi.org/10.1016/j.devcel.2009.02.010.
Gonzales, L.W., Guttentag, S.H., Wade, K.C., Postle, A.D., and Ballard, P.L. (2002). Differentiation of human pulmonary type II cells in vitro by glucocorticoid plus cAMP. Am. J. Physiol.-Lung Cell. Mol. Physiol. 283, L940-L951. https://doi.org/10.1152/ajplung.00127.2002.
Goss, A.M., Tian, Y., Tsukiyama, T., Cohen, E.D., Zhou, D., Lu, M.M., Yamaguchi, T.P., and Morrisey, E.E. (2009). Wnt2/2b and beta-Catenin Signaling Are Necessary and Sufficient to Specify Lung Progenitors in the Foregut. Dev. Cell 17, 290–298. https://doi.org/10.1016/j.devcel.2009.06.005.
Gotoh, S., Ito, I., Nagasaki, T., Yamamoto, Y., Konishi, S., Korogi, Y., Matsumoto, H., Muro, S., Hirai, T., Funato, M., et al. (2014). Generation of Alveolar Epithelial Spheroids via Isolated Progenitor Cells from Human Pluripotent Stem Cells. Stem Cell Rep. 3, 394–403. https://doi.org/10.1016/j.stemcr.2014.07.005.
Green, M.D., Chen, A., Nostro, M.C., d’Souza, S.L., Schaniel, C., Lemischka, I.R., Gouon-Evans, V., Keller, G., and Snoeck, H.W. (2011). Generation of anterior foregut endoderm from human embryonic and induced pluripotent stem cells. Nat. Biotechnol. 29, 267-U153. https://doi.org/10.1038/nbt.1788.
Greggio, C., De Franceschi, F., Figueiredo-Larsen, M., Gobaa, S., Ranga, A., Semb, H., Lutolf, M., and Grapin-Botton, A. (2013). Artificial three-dimensional niches deconstruct pancreas development in vitro. Development 140, 4452–4462. https://doi.org/10.1242/dev.096628.
Han, L., Chaturvedi, P., Kishimoto, K., Koike, H., Nasr, T., Iwasawa, K., Giesbrecht, K., Witcher, P.C., Eicher, A., Haines, L., et al. (2020). Single cell transcriptomics identifies a signaling network coordinating endoderm and mesoderm diversification during foregut organogenesis. Nat. Commun. 11, 4158. https://doi.org/10.1038/s41467-020-17968-x.
Hao, Y., Hao, S., Andersen-Nissen, E., Mauck, W.M., 3rd, Zheng, S., Butler, A., Lee, M.J., Wilk, A.J., Darby, C., Zager, M., et al. (2021). Integrated analysis of multimodal single-cell data. Cell 184, 3573–3587.e3529. https://doi.org/10.1016/j.cell.2021.04.048.
Harris-Johnson, K.S., Domyan, E.T., Vezina, C.M., and Sun, X. (2009). beta-Catenin promotes respiratory progenitor identity in mouse foregut. Proc. Natl. Acad. Sci. U. S. A. 106, 16287–16292. https://doi.org/10.1073/pnas.0902274106.
Hawkins, F., Kramer, P., Jacob, A., Driver, I., Thomas, D.C., McCauley, K.B., Skvir, N., Crane, A.M., Kurmann, A.A., Hollenberg, A.N., et al. (2017). Prospective isolation of NKX2-1-expressing human lung progenitors derived from pluripotent stem cells. J. Clin. Invest. 127, 2277–2294. https://doi.org/10.1172/jci89950.
Hawkins, F.J., Suzuki, S., Beermann, M.L., Barilla, C., Wang, R.B., Villacorta-Martin, C., Berical, A., Jean, J.C., Le Suer, J., Matte, T., et al. (2021). Derivation of Airway Basal Stem Cells from Human Pluripotent Stem Cells. Cell Stem Cell 28, 79–95. https://doi.org/10.1016/j.stem.2020.09.017.
He, P., Lim, K., Sun, D.W., Pett, J.P., Jeng, Q., Polanski, K., Dong, Z.Q., Bolt, L., Richardson, L., Mamanova, L., et al. (2022). A human fetal lung cell atlas uncovers proximal-distal gradients of differentiation and key regulators of epithelial fates. Cell, 185(25), 4841–4860. https://doi.org/10.1016/j.cell.2022.11.005.
Hein, R.F.C., Conchola, A.S., Fine, A.S., Xiao, Z., Frum, T., Brastrom, L.K., Akinwale, M.A., Childs, C.J., Tsai, Y.-H., Holloway, E.M., et al. (2022a). Stable iPSC-derived NKX2-1+ lung bud tip progenitor organoids give rise to airway and alveolar cell types. Development 149, dev200693. https://doi.org/10.1242/dev.200693.
Hein, R.F.C., Wu, J.H., Holloway, E.M., Frum, T., Conchola, A.S., Tsai, Y.-H., Wu, A., Fine, A.S., Miller, A.J., Szenker-Ravi, E., et al. (2022b). R-SPONDIN2+ mesenchymal cells form the bud tip progenitor niche during human lung development. Dev. Cell 57, 598–1614. https://doi.org/10.1016/j.devcel.2022.05.010.
Herriges, J.C., Yi, L., Hines, E.A., Harvey, J.F., Xu, G.L., Gray, P.A., Ma, Q.F., and Sun, X. (2012). Genome-scale study of transcription factor expression in the branching mouse lung. Dev. Dyn. 241, 1432–1453. https://doi.org/10.1002/dvdy.23823.
Herriges, M.J., Yampolskaya, M., Thapa, B.R., Lindstrom-Vautrin, J., Wang, F., Na, C.-L., Ma, L., Montminy, M.M., Huang, J., Villacorta-Martin, C., et al. (2022). Durable alveolar engraftment of PSC-derived lung epithelial cells into immunocompetent mice. bioRxiv, 2022.2007.2026.501591. https://doi.org/10.1101/2022.07.26.501591.
Hicks, S.C., Townes, F.W., Teng, M.X., and Irizarry, R.A. (2018). Missing data and technical variability in single-cell RNA-sequencing experiments. Biostatistics 19, 562–578. https://doi.org/10.1093/biostatistics/kxx053.
Huang, S.X.L., Islam, M.N., O’Neill, J., Hu, Z., Yang, Y.G., Chen, Y.W., Mumau, M., Green, M.D., Vunjak-Novakovic, G., Bhattacharya, J., and Snoeck, H.W. (2014). Efficient generation of lung and airway epithelial cells from human pluripotent stem cells. Nat. Biotechnol. 32, 84–91. https://doi.org/10.1038/nbt.2754.
Ikonomou, L., Herriges, M.J., Lewandowski, S.L., Marsland, R., Villacorta-Martin, C., Caballero, I.S., Frank, D.B., Sanghrajka, R.M., Dame, K., Kandula, M.M., et al. (2020a). The in vivo genetic program of murine primordial lung epithelial progenitors. Nat. Commun. 11, 635. https://doi.org/10.1038/s41467-020-14348-3.
Ikonomou, L., and Kotton, D.N. (2015). Derivation of Endodermal Progenitors From Pluripotent Stem Cells. J. Cell. Physiol. 230, 246–258. https://doi.org/10.1002/jcp.24771.
Ikonomou, L., Wagner, D.E., Gilpin, S.E., Weiss, D.J., and Ryan, A.L. (2020b). Technological advances in study of lung regenerative medicine: perspective from the 2019 Vermont lung stem cell conference. Cytotherapy 22, 519–520. https://doi.org/10.1016/j.jcyt.2020.04.001.
Jacob, A., Morley, M., Hawkins, F., McCauley, K.B., Jean, J.C., Heins, H., Na, C.L., Weaver, T.E., Vedaie, M., Hurley, K., et al. (2017). Differentiation of Human Pluripotent Stem Cells into Functional Lung Alveolar Epithelial Cells. Cell Stem Cell 21, 472–488. https://doi.org/10.1016/j.stem.2017.08.014.
Jonsson, J., Carlsson, L., Edlund, T., and Edlund, H. (1994). Insulin-Promoter-Factor-1 Is Required for Pancreas Development in Mice. Nature 371, 606–609.
Khattar, D., Fernandes, S., Snowball, J., Guo, M., Gillen, M.C., Jain, S.S., Sinner, D., Zacharias, W., and Swarr, D.T. (2022). PI3K signaling specifies proximal-distal fate by driving a developmental gene regulatory network in SOX9+ mouse lung progenitors. eLife 11, e67954. https://doi.org/10.7554/eLife.67954.
Kimura, S., Hara, Y., Pineau, T., FernandezSalguero, P., Fox, C.H., Ward, J.M., and Gonzalez, F.J. (1996). The T/ebp null mouse thyroid-specific enhancer-binding protein is essential for the organogenesis of the thyroid, lung, ventral forebrain, and pituitary. Genes Dev. 10, 60–69.
Kimura, S., Ward, J.M., and Minoo, P. (1999). Thyroid-specific enhancer-binding protein thyroid transcription factor 1 is not required for the initial specification of the thyroid and lung primordia. Biochimie 81, 321–327.
Kishimoto, K., Furukawa, K.T., Luz-Madrigal, A., Yamaoka, A., Matsuoka, C., Habu, M., Alev, C., Zorn, A.M., and Morimoto, M. (2020). Bidirectional Wnt signaling between endoderm and mesoderm confers tracheal identity in mouse and human cells. Nat. Commun. 11, 4159. https://doi.org/10.1038/s41467-020-17969-w.
Kishimoto, K., Iwasawa, K., Sorel, A., Ferran-Heredia, C., Han, L., Morimoto, M., Wells, J.M., Takebe, T., and Zorn, A.M. (2022). Directed differentiation of human pluripotent stem cells into diverse organ-specific mesenchyme of the digestive and respiratory systems. Nat. Protoc. 17, 2699–2719. https://doi.org/10.1038/s41596-022-00733-3.
Konishi, S., Gotoh, S., Tateishi, K., Yamamoto, Y., Korogi, Y., Nagasaki, T., Matsumoto, H., Muro, S., Hirai, T., Ito, I., et al. (2016). Directed Induction of Functional Multi-ciliated Cells in Proximal Airway Epithelial Spheroids from Human Pluripotent Stem Cells. Stem Cell Rep. 6, 18–25. https://doi.org/10.1016/j.stemcr.2015.11.010.
Kumar, M.E., Bogard, P.E., Espinoza, F.H., Menke, D.B., Kingsley, D.M., and Krasnow, M.A. (2014). Defining a mesenchymal progenitor niche at single-cell resolution. Science 346, 1258810, 1258810. https://doi.org/10.1126/science.1258810.
Lang, A.H., Li, H., Collins, J.J., and Mehta, P. (2014). Epigenetic Landscapes Explain Partially Reprogrammed Cells and Identify Key Reprogramming Genes. PLoS Comput. Biol. 10, e1003734. https://doi.org/10.1371/journal.pcbi.1003734.
Laresgoiti, U., Nikolic, M.Z., Rao, C., Brady, J.L., Richardson, R.V., Batchen, E.J., Chapman, K.E., and Rawlins, E.L. (2016). Lung epithelial tip progenitors integrate glucocorticoid- and STAT3-mediated signals to control progeny fate. Development 143, 3686–3699. https://doi.org/10.1242/dev.134023.
Lasser, M., Pratt, B., Monahan, C., Kim, S.W., and Lowery, L.A. (2019). The Many Faces of Xenopus: Xenopus laevis as a Model System to Study Wolf Hirschhorn Syndrome. Front. Physiol. 10, 12, 817. https://doi.org/10.3389/fphys.2019.00817.
Lebeche, D., Malpel, S., and Cardoso, W.V. (1999). Fibroblast growth factor interactions in the developing lung. Mech. Dev. 86, 125–136.
Lewandowski, S., Villacorta-Martin, C., Kotton, D.N., and Ikonomou, L. (2020). Single Cell Analysis and Novel Ex Vivo Culture of Lung Primordial Progenitors. Am. J. Respir. Crit. Care Med. 201, 1.
Lim, K., Tang, W., Sun, D., He, P., Teichmann, S.A., Marioni, J.C., Meyer, K.B., and Rawlins, E.L. (2021). Acquisition of alveolar fate and differentiation competence by human fetal lung epithelial progenitor cells. bioRxiv, 2021.2006.2030.450501. https://doi.org/10.1101/2021.06.30.450501.
Litingtung, Y., Lei, L., Westphal, H., and Chiang, C. (1998). Sonic hedgehog is essential to foregut development. Nature Genet. 20, 58–61.
Liu, Y., and Hogan, B.L. (2002). Differential gene expression in the distal tip endoderm of the embryonic mouse lung. Gene Expr Patterns 2, 229–233.
Longmire, T.A., Ikonomou, L., Hawkins, F., Christodoulou, C., Cao, Y.X., Jean, J.C., Kwok, L.W., Mou, H.M., Rajagopal, J., Shen, S.S., et al. (2012). Efficient Derivation of Purified Lung and Thyroid Progenitors from Embryonic Stem Cells. Cell Stem Cell 10, 398–411. https://doi.org/10.1016/j.stem.2012.01.019.
Luecken, M.D., Buttner, M., Chaichoompu, K., Danese, A., Interlandi, M., Mueller, M.F., Strobl, D.C., Zappia, L., Dugas, M., Colome-Tatche, M., and Theis, F.J. (2022). Benchmarking atlas-level data integration in single-cell genomics. Nat. Methods 19, 41–50. https://doi.org/10.1038/s41592-021-01336-8.
McCauley, K.B., Alysandratos, K.D., Jacob, A., Hawkins, F., Caballero, I.S., Vedaie, M., Yang, W.L., Slovik, K.J., Morley, M., Carraro, G., et al. (2018). Single-Cell Transcriptomic Profiling of Pluripotent Stem Cell-Derived SCGB3A2+Airway Epithelium. Stem Cell Rep. 10, 1579–1595. https://doi.org/10.1016/j.stemcr.2018.03.013.
McCauley, K.B., Hawkins, F., Serra, M., Thomas, D.C., Jacob, A., and Kotton, D.N. (2017). Efficient Derivation of Functional Human Airway Epithelium from Pluripotent Stem Cells via Temporal Regulation of Wnt Signaling. Cell Stem Cell 20, 844–857. https://doi.org/10.1016/j.stem.2017.03.001.
Miller, A.J., Hill, D.R., Nagy, M.S., Aoki, Y., Dye, B.R., Chin, A.M., Huang, S., Zhu, F., White, E.S., Lama, V., and Spence, J.R. (2018). In Vitro Induction and In Vivo Engraftment of Lung Bud Tip Progenitor Cells Derived from Human Pluripotent Stem Cells. Stem Cell Rep. 10, 101–119. https://doi.org/10.1016/j.stemcr.2017.11.012.
Miller, A.J., Yu, Q.H., Czerwinski, M., Tsai, Y.H., Conway, R.F., Wu, A., Holloway, E.M., Walker, T., Glass, I.A., Treutlein, B., et al. (2020). In Vitro and In Vivo Development of the Human Airway at Single-Cell Resolution. Dev. Cell 53, 117–128. https://doi.org/10.1016/j.devcel.2020.01.033.
Motoyama, J., Liu, J., Mo, R., Ding, Q., Post, M., and Hui, C.C. (1998). Essential function of Gli2 and Gli3 in the formation of lung, trachea and oesophagus. Nature Genet. 20, 54–57.
Mou, H.M., Zhao, R., Sherwood, R., Ahfeldt, T., Lapey, A., Wain, J., Sicilian, L., Izvolsky, K., Musunuru, K., Cowan, C., and Rajagopal, J. (2012). Generation of Multipotent Lung and Airway Progenitors from Mouse ESCs and Patient-Specific Cystic Fibrosis iPSCs. Cell Stem Cell 10, 385–397. https://doi.org/10.1016/j.stem.2012.01.018.
Negretti, N.M., Plosa, E.J., Benjamin, J.T., Schuler, B.A., Habermann, A.C., Jetter, C.S., Gulleman, P., Bunn, C., Hackett, A.N., Ransom, M., et al. (2021). A single-cell atlas of mouse lung development. Development 148, dev199512. https://doi.org/10.1242/dev.199512.
Nichane, M., Javed, A., Sivakamasundari, V., Ganesan, M., Ang, L.T., Kraus, P., Lufkin, T., Loh, K.M., and Lim, B. (2017). Isolation and 3D expansion of multipotent Sox9(+) mouse lung progenitors. Nat. Methods 14, 1205–1212. https://doi.org/10.1038/nmeth.4498.
Nikolic, M.Z., Caritg, O., Jeng, Q., Johnson, J.A., Sun, D.W., Howell, K.J., Brady, J.L., Laresgoiti, U., Allen, G., Butler, R., et al. (2017). Human embryonic lung epithelial tips are multipotent progenitors that can be expanded in vitro as long-term selfrenewing organoids. eLife 6, e26575. https://doi.org/10.7554/eLife.26575.001.
Okubo, T., and Hogan, B.L. (2004). Hyperactive Wnt signaling changes the developmental potential of embryonic lung endoderm. J Biol 3, 11. https://doi.org/10.1186/jbiol3.
Okubo, T., Knoepfler, P.S., Eisenman, R.N., and Hogan, B.L.M. (2005). Nmyc plays an essential role during lung development as a dosage-sensitive regulator of progenitor cell proliferation and differentiation. Development 132, 1363–1374. https://doi.org/10.1242/dev.01678.
Ostrin, E.J., Little, D.R., Gerner-Mauro, K.N., Sumner, E.A., Rios-Corzo, R., Ambrosio, E., Holt, S.E., Forcioli-Conti, N., Akiyama, H., Hanash, S.M., et al. (2018). beta-Catenin maintains lung epithelial progenitors after lung specification. Development 145, 13, dev160788. https://doi.org/10.1242/dev.160788.
Peng, T., Tian, Y., Boogerd, C.J., Lu, M.M., Kadzik, R.S., Stewart, K.M., Evans, S.M., and Morrisey, E.E. (2013). Coordination of heart and lung co-development by a multipotent cardiopulmonary progenitor. Nature 500, 589–592. https://doi.org/10.1038/nature12358.
Peng, Y.R., Shekhar, K., Yan, W.J., Herrmann, D., Sappington, A., Bryman, G.S., van Zyl, T., Do, M.T.H., Regev, A., and Sanes, J.R. (2019). Molecular Classification and Comparative Taxonomics of Foveal and Peripheral Cells in Primate Retina. Cell 176, 1222–1237. https://doi.org/10.1016/j.cell.2019.01.004.
Peterson, R.S., Lim, L., Ye, H.G., Zhou, H.P., Overdier, D.G., and Costa, R.H. (1997). The winged helix transcriptional activator HFH-8 is expressed in the mesoderm of the primitive streak stage of mouse embryos and its cellular derivatives. Mech. Dev. 69, 53–69. https://doi.org/10.1016/s0925-4773(97)00153-6.
Pusuluri, S.T., Lang, A.H., Mehta, P., and Castillo, H.E. (2018). Cellular reprogramming dynamics follow a simple 1D reaction coordinate. Phys. Biol. 15, 016001. https://doi.org/10.1088/1478-3975/aa90e0.
Rankin, S.A., Gallas, A.L., Neto, A., Gomez-Skarmeta, J.L., and Zorn, A.M. (2012). Suppression of Bmp4 signaling by the zinc-finger repressors Osr1 and Osr2 is required for Wnt/beta-catenin-mediated lung specification in Xenopus. Development 139, 3010–3020. https://doi.org/10.1242/dev.078220.
Rankin, S.A., Han, L., McCracken, K.W., Kenny, A.P., Anglin, C.T., Grigg, E.A., Crawford, C.M., Wells, J.M., Shannon, J.M., and Zorn, A.M. (2016). A Retinoic Acid-Hedgehog Cascade Coordinates Mesoderm-Inducing Signals and Endoderm Competence during Lung Specification. Cell Reports 16, 66–78. https://doi.org/10.1016/j.celrep.2016.05.060.
Rankin, S.A., McCracken, K.W., Luedeke, D.M., Han, L., Wells, J.M., Shannon, J.M., and Zorn, A.M. (2018). Timing is everything: Reiterative Wnt, BMP and RA signaling regulate developmental competence during endoderm organogenesis. Dev. Biol. 434, 121–132. https://doi.org/10.1016/j.ydbio.2017.11.018.
Rankin, S.A., and Zorn, A.M. (2014). Gene regulatory networks Governing lung specification. J. Cell. Biochem. 115, 1343–1350. https://doi.org/10.1002/jcb.24810.
Rankin, S.A., and Zorn, A.M. (2022). The homeodomain transcription factor Ventx2 regulates respiratory progenitor cell number and differentiation timing during Xenopus lung development. Dev. Growth Diff. 64, 347–361. https://doi.org/10.1111/dgd.12807.
Rawlins, E.L., Clark, C.P., Xue, Y., and Hogan, B.L.M. (2009). The Id2(+) distal tip lung epithelium contains individual multipotent embryonic progenitor cells. Development 136, 3741–3745. https://doi.org/10.1242/dev.037317.
Riccetti, M., Gokey, J.J., Aronow, B., and Perl, A.K.T. (2020). The elephant in the lung: Integrating lineage-tracing, molecular markers, and single cell sequencing data to identify distinct fibroblast populations during lung development and regeneration. Matrix Biol. 91–92, 51–74. https://doi.org/10.1016/j.matbio.2020.05.002.
Rock, J.R., Onaitis, M.W., Rawlins, E.L., Lu, Y., Clark, C.P., Xue, Y., Randell, S.H., and Hogan, B.L.M. (2009). Basal cells as stem cells of the mouse trachea and human airway epithelium. Proc. Natl. Acad. Sci. U. S. A. 106, 12771–12775. https://doi.org/10.1073/pnas.0906850106.
Rock, J.R., Randell, S.H., and Hogan, B.L.M. (2010). Airway basal stem cells: a perspective on their roles in epithelial homeostasis and remodeling. Dis. Model. Mech. 3, 545–556. https://doi.org/10.1242/dmm.006031.
Rockich, B.E., Hrycaj, S.M., Shih, H.P., Nagy, M.S., Ferguson, M.A.H., Kopp, J.L., Sander, M., Wellik, D.M., and Spence, J.R. (2013). Sox9 plays multiple roles in the lung epithelium during branching morphogenesis. Proc. Natl. Acad. Sci. U. S. A. 110, E4456-E4464. https://doi.org/10.1073/pnas.1311847110.
Rossant, J. (2011). The Impact of Developmental Biology on Pluripotent Stem Cell Research: Successes and Challenges. Dev. Cell 21, 20–23. https://doi.org/10.1016/j.devcel.2011.06.010.
Sandberg, M., Flandin, P., Silberberg, S., Su-Feher, L., Price, J.D., Hu, J.S., Kim, C., Visel, A., Nord, A.S., and Rubenstein, J.L.R. (2016). Transcriptional Networks Controlled by NKX2-1 in the Development of Forebrain GABAergic Neurons. Neuron 91, 1260–1275. https://doi.org/10.1016/j.neuron.2016.08.020.
Serra, M., Alysandratos, K.D., Hawkins, F., McCauley, K.B., Jacob, A., Choi, J., Caballero, I.S., Vedaie, M., Kurmann, A.A., Ikonomou, L., et al. (2017). Pluripotent stem cell differentiation reveals distinct developmental pathways regulating lung- versus thyroid-lineage specification. Development 144, 3879–3893. https://doi.org/10.1242/dev.150193.
Sherwood, R.I., Chen, T.Y.A., and Melton, D.A. (2009). Transcriptional Dynamics of Endodermal Organ Formation. Dev. Dyn. 238, 29–42. https://doi.org/10.1002/dvdy.21810.
Sun, D., Llora Batlle, O., van den Ameele, J., Thomas, J.C., He, P., Lim, K., Tang, W., Xu, C., Meyer, K.B., Teichmann, S.A., et al. (2022). SOX9 maintains human foetal lung tip progenitor state by enhancing WNT and RTK signalling. EMBO Journal, 41, e111338. https://doi.org/10.15252/embj.2022111338.
Sun, D.W., Evans, L., Perrone, F., Sokleva, V., Lim, K., Rezakhani, S., Lutolf, M., Zilbauer, M., and Rawlins, E.L. (2021). A functional genetic toolbox for human tissue-derived organoids. eLife 10, e67886. https://doi.org/10.7554/eLife.67886; https://doi.org/10.7554/eLife.67886.sa1; https://doi.org/10.7554/eLife.67886.sa2.
Swarr, D.T., and Morrisey, E.E. (2015). Lung Endoderm Morphogenesis: Gasping for Form and Function. In Annual Review of Cell and Developmental Biology, Vol 31, R. Schekman, ed. (Annual Reviews), pp. 553–573. https://doi.org/10.1146/annurev-cellbio-100814-125249.
Travaglini, K.J., Nabhan, A.N., Penland, L., Sinha, R., Gillich, A., Sit, R.V., Chang, S., Conley, S.D., Mori, Y., Seita, J., et al. (2020). A molecular cell atlas of the human lung from single-cell RNA sequencing. Nature 587, 619–625. https://doi.org/10.1038/s41586-020-2922-4.
Varghese, B., Ling, Z.H., and Ren, X. (2022). Reconstructing the pulmonary niche with stem cells: a lung story. Stem Cell Res. Ther. 13, 161. https://doi.org/10.1186/s13287-022-02830-2.
Wade, K.C., Guttentag, S.H., Gonzales, L.W., Maschhoff, K.L., Gonzales, J., Kolla, V., Singhal, S., and Ballard, P.L. (2006). Gene induction during differentiation of human pulmonary type II cells in vitro. Am. J. Respir. Cell Mol. Biol. 34, 727–737. https://doi.org/10.1165/rcmb.2004-0389OC.
Weinreb, C., Wolock, S., and Klein, A.M. (2018). SPRING: a kinetic interface for visualizing high dimensional single-cell expression data. Bioinformatics 34, 1246–1248. https://doi.org/10.1093/bioinformatics/btx792.
Wessells, N.K. (1973). Tissue interactions in development (Addison-Wesley Pub. Co.).
Xie, B.B., Jiang, Q., Mora, A., and Li, X.R. (2021). Automatic cell type identification methods for single-cell RNA sequencing. Comp. Struct. Biotechnol. J.. 19, 5874–5887. https://doi.org/10.1016/j.csbj.2021.10.027.
Xiong, A.M., Austin, T.W., Lagasse, E., Uchida, N., Tamaki, S., Bordier, B.B., Weissman, I.L., Glenn, J.S., and Millan, M.T. (2008). Isolation of human fetal liver progenitors and their enhanced proliferation by three-dimensional coculture with endothelial cells. Tissue Eng. Part A 14, 995–1006. https://doi.org/10.1089/ten.tea.2007.0087.
Yamamoto, Y., Gotoh, S., Korogi, Y., Seki, M., Konishi, S., Ikeo, S., Sone, N., Nagasaki, T., Matsumoto, H., Muro, S., et al. (2017). Long-term expansion of alveolar stem cells derived from human iPS cells in organoids. Nat. Methods 14, 1097–1106. https://doi.org/10.1038/nmeth.4448.
Yampolskaya, M., Herriges, M., Ikonomou, L., Kotton, D., & Mehta, P. (2023). scTOP: physics-inspired order parameters for cellular identification and visualization. bioRxiv, 2023.2001.2025.525581. https://doi.org/10.1101/2023.01.25.525581.
Yang, Y., Riccio, P., Schotsaert, M., Mori, M., Lu, J.N., Lee, D.K., Garcia-Sastre, A., Xu, J.M., and Cardoso, W.V. (2018). Spatial-Temporal Lineage Restrictions of Embryonic p63(+) Progenitors Establish Distinct Stem Cell Pools in Adult Airways. Dev. Cell 44, 752–761. https://doi.org/10.1016/j.devcel.2018.03.001.
Yu, Q.H., Kilik, U., Holloway, E.M., Tsai, Y.H., Harmel, C., Wu, A., Wu, J.H., Czerwinski, M., Childs, C.J., He, Z.S., et al. (2021). Charting human development using a multi-endodermal organ atlas and organoid models. Cell 184, 3281–3298.e3222. https://doi.org/10.1016/j.cell.2021.04.028.
Zepp, J.A., Zacharias, W.J., Frank, D.B., Cavanaugh, C.A., Zhou, S., Morley, M.P., and Morrisey, E.E. (2017). Distinct Mesenchymal Lineages and Niches Promote Epithelial Self-Renewal and Myofibrogenesis in the Lung. Cell 170, 1134–1148.e10. https://doi.org/10.1016/j.cell.2017.07.034.
Zhang, W.M., Menke, D.B., Jiang, M.S., Chen, H., Warburton, D., Turcatel, G., Lu, C.H., Xu, W., Luo, Y.F., and Shi, W. (2013). Spatial-temporal targeting of lung-specific mesenchyme by a Tbx4 enhancer. BMC Biol. 11, 111. https://doi.org/10.1186/1741-7007-11-111.
Acknowledgments
We apologize to those authors whose work we could not cite due to space limitations. L.I. was supported by the National Institutes of Health funding (R56 DE028545-01 and R01 HL158965-01) and the University at Buffalo (UB) Research Foundation Start-up Funds. M.Y. and P.M. were supported by the National Institutes of Health NIGMS funding (2R35 GM119461). Figures were created with Biorender.com.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Open Access This chapter is licensed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license and indicate if changes were made.
The images or other third party material in this chapter are included in the chapter's Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the chapter's Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder.
Copyright information
© 2023 The Author(s)
About this chapter
Cite this chapter
Ikonomou, L., Yampolskaya, M., Mehta, P. (2023). Multipotent Embryonic Lung Progenitors: Foundational Units of In Vitro and In Vivo Lung Organogenesis. In: Magin, C.M. (eds) Engineering Translational Models of Lung Homeostasis and Disease. Advances in Experimental Medicine and Biology, vol 1413. Springer, Cham. https://doi.org/10.1007/978-3-031-26625-6_4
Download citation
DOI: https://doi.org/10.1007/978-3-031-26625-6_4
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-031-26624-9
Online ISBN: 978-3-031-26625-6
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)