Abstract
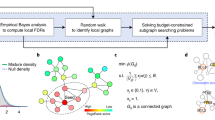
A standard paradigm in computational biology is to use interaction networks to analyze high-throughput biological data. Two common approaches for leveraging interaction networks are: (1) network ranking, where one ranks vertices in the network according to both vertex scores and network topology; (2) altered subnetwork identification, where one identifies one or more subnetworks in an interaction network using both vertex scores and network topology. The dominant approach in network ranking is network propagation which smooths vertex scores over the network using a random walk or diffusion process, thus utilizing the global structure of the network. For altered subnetwork identification, existing algorithms either restrict solutions to subnetworks in subnetwork families with simple topological constraints, such as connected subnetworks, or utilize ad hoc heuristics that lack a rigorous statistical foundation. In this work, we unify the network propagation and altered subnetwork approaches. We derive a subnetwork family which we call the propagation family that approximates the subnetworks ranked highly by network propagation. We introduce NetMix2, a principled algorithm for identifying altered subnetworks from a wide range of subnetwork families, including the propagation family, thus combining the advantages of the network propagation and altered subnetwork approaches. We show that NetMix2 outperforms network propagation on data simulated using the propagation family. Furthermore, NetMix2 outperforms other methods at recovering known disease genes in pan-cancer somatic mutation data and in genome-wide association data from multiple human diseases. NetMix2 is publicly available at https://github.com/raphael-group/netmix2.
U. Chitra and T. Y. Park—Contributed equally to the manuscript.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
Notes
- 1.
A related problem is the identification of altered subnetworks according to network topology alone. Many of the leading methods for this problem were benchmarked in a recent DREAM competition [18].
References
Addario-Berry, L., Broutin, N., Devroye, L., Lugosi, G.: On combinatorial testing problems. Ann. Stat. 38(5), 3063–3092 (2010)
Arias-Castro, E., Candès, E.J., Durand, A.: Detection of an anomalous cluster in a network. Ann. Stat. 39(1), 278–304 (2011)
Arias-Castro, E., Candès, E.J., Helgason, H., Zeitouni, O.: Searching for a trail of evidence in a maze. Ann. Stat. 36(4), 1726–1757 (2008)
Arias-Castro, E., Donoho, D.L., Huo, X.: Adaptive multiscale detection of filamentary structures in a background of uniform random points. Ann. Stat. 34(1), 326–349 (2006)
Azencott, C.A., Grimm, D., Sugiyama, M., Kawahara, Y., Borgwardt, K.M.: Efficient network-guided multi-locus association mapping with graph cuts. Bioinformatics 29(13), i171–i179 (2013)
Bailey, M.H., et al.: Comprehensive characterization of cancer driver genes and mutations. Cell 173(2), 371–385 (2018)
Barel, G., Herwig, R.: NetCore: a network propagation approach using node coreness. Nucleic Acids Res. 48(17), e98–e98 (2020)
Battaglia, S., Maguire, O., Campbell, M.J.: Transcription factor co-repressors in cancer biology: roles and targeting. Int. J. Cancer 126(11), 2511–2519 (2010)
Berger, B., Peng, J., Singh, M.: Computational solutions for omics data. Nature Rev. Genet. 14(5), 333–346 (2013)
Cadena, J., Chen, F., Vullikanti, A.: Near-optimal and practical algorithms for graph scan statistics with connectivity constraints. ACM Trans. Knowl. Discov. Data 13(2), 20:1-20:33 (2019)
Cai, T.T., Jin, J., Low, M.G.: Estimation and confidence sets for sparse normal mixtures. Ann. Stat. 35(6), 2421–2449 (2007)
Califano, A., Butte, A.J., Friend, S., Ideker, T., Schadt, E.: Leveraging models of cell regulation and GWAS data in integrative network-based association studies. Nat. Genet. 44(8), 841–847 (2012)
Cao, M., et al.: Going the distance for protein function prediction: a new distance metric for protein interaction networks. PLoS One 8(10), 1–12 (2013)
Chakravarty, D., et al.: OncoKB: a precision oncology knowledge base. JCO Precis. Oncol. 1, 1–16 (2017)
Chasman, D., Siahpirani, A.F., Roy, S.: Network-based approaches for analysis of complex biological systems. Curr. Opin. Biotech. 39, 157–166 (2016)
Chitra, U., Ding, K., Lee, J.C., Raphael, B.J.: Quantifying and reducing bias in maximum likelihood estimation of structured anomalies. In: Proceedings of the 38th International Conference on Machine Learning, pp. 1908–1919. PMLR, 18–24 July 2021
Cho, D.Y., Kim, Y.A., Przytycka, T.M.: Chapter 5: network biology approach to complex diseases. PLoS Comput. Biol. 8(12), 1–11 (2012)
Choobdar, S., et al.: Assessment of network module identification across complex diseases. Nat. Methods 16(9), 843–852 (2019)
Chua, H.N., Sung, W.K., Wong, L.: Exploiting indirect neighbours and topological weight to predict protein function from protein-protein interactions. Bioinformatics 22(13), 1623–1630 (2006)
modENCODE Consortium, Roy, S., Ernst, J., Kharchenko, P.V., Kheradpour, P., et al.: Identification of functional elements and regulatory circuits by drosophila modencode. Science 330(6012), 1787–1797 (2010)
Cornish, A.J., Markowetz, F.: SANTA: Quantifying the functional content of molecular networks. PLoS Comput. Biol. 10(9), e1003808 (2014)
Cowen, L., Devkota, K., Hu, X., Murphy, J.M., Wu, K.: Diffusion state distances: Multitemporal analysis, fast algorithms, and applications to biological networks. SIAM J. Math. Data Sci. 3(1), 142–170 (2021)
Cowen, L., Ideker, T., Raphael, B.J., Sharan, R.: Network propagation: a universal amplifier of genetic associations. Nat. Rev. Genet. 18(9), 551–562 (2017)
Creixell, P., et al.: Pathway and network analysis of cancer genomes. Nat. Methods 12(7), 615–621 (2015)
de la Fuente, A.: From ‘differential expression’ to ‘differential networking’ - identification of dysfunctional regulatory networks in diseases. Trends Genet. 26(7), 326–333 (2010)
Deng, M., Zhang, K., Mehta, S., Chen, T., Sun, F.: Prediction of protein function using protein-protein interaction data. J. Comput. Biol. 10(6), 947–960 (2003)
Dimitrakopoulos, C.M., Beerenwinkel, N.: Computational approaches for the identification of cancer genes and pathways. WIREs Syst. Biol. Med. 9(1), e1364 (2017)
Dittrich, M.T., Klau, G., Rosenwald, A., Dandekar, T., Muller, T.: Identifying functional modules in protein-protein interaction networks: an integrated exact approach. Bioinformatics 24(13), i223–i231 (2008)
Donoho, D., Jin, J.: Higher criticism for detecting sparse heterogeneous mixtures. Ann. Stat. 32(3), 962–994 (2004)
Efron, B.: Large-scale simultaneous hypothesis testing: the choice of a null hypothesis. J. Am. Stat. Assoc. 99(465), 96–104 (2004)
Efron, B.: Correlation and large-scale simultaneous significance testing. J. Am. Stat. Assoc. 102(477), 93–103 (2007)
Efron, B.: Size, power and false discovery rates. Ann. Stat. 35(4), 1351–1377 (2007)
Ghiassian, S.D., Menche, J., Barabási, A.L.: A DIseAse MOdule Detection (DIAMOnD) algorithm derived from a systematic analysis of connectivity patterns of disease proteins in the human interactome. PLoS Comput. Biol. 11(4), e1004120 (2015)
Glaz, J., Naus, J., Wallenstein, S.: Scan Statistics. Springer-Verlag, New York (2001). https://doi.org/10.1007/978-1-4757-3460-7
Gligorijević, V., Pržulj, N.: Methods for biological data integration: perspectives and challenges. J. Roy. Soc. Interface 12(112), 20150571 (2015)
Guo, Z., et al.: Edge-based scoring and searching method for identifying condition-responsive protein-protein interaction sub-network. Bioinformatics 23(16), 2121–2128 (2007)
Gurobi Optimization, LLC: Gurobi Optimizer Reference Manual (2021)
Halldórsson, B.V., Sharan, R.: Network-based interpretation of genomic variation data. J. Mol. Biol. 425(21), 3964–3969 (2013)
Hofree, M., Shen, J.P., Carter, H., Gross, A., Ideker, T.: Network-based stratification of tumor mutations. Nat. Methods 10(11), 1108–1115 (2013)
Hormozdiari, F., Penn, O., Borenstein, E., Eichler, E.E.: The discovery of integrated gene networks for autism and related disorders. Genome Res. 25(1), 142–154 (2015)
Horn, H., Lawrence, M.S., Chouinard, C.R., Shrestha, Y., Hu, J.X., et al.: NetSig: network-based discovery from cancer genomes. Nat. Methods 15(1), 61–66 (2018)
Huang, J.K., et al.: Systematic evaluation of molecular networks for discovery of disease genes. Cell Syst. 6(4), 484–495 (2018)
Ideker, T., Ozier, O., Schwikowski, B., Siegel, A.F.: Discovering regulatory and signalling circuits in molecular interaction networks. Bioinformatics 18(suppl 1), S233–S240 (2002)
Ideker, T., et al.: Integrated genomic and proteomic analyses of a systematically perturbed metabolic network. Science 292(5518), 929–934 (2001)
Jia, P., Zhao, Z.: Network assisted analysis to prioritize GWAS results: principles, methods and perspectives. Hum. Genet. 133(2), 125–138 (2014). https://doi.org/10.1007/s00439-013-1377-1
Kloumann, I.M., Ugander, J., Kleinberg, J.: Block models and personalized PageRank. Proc. Natl. Acad. Sci. 114(1), 33–38 (2017)
Kulldorff, M.: A spatial scan statistic. Commun. Stat. Theory Methods 26(6), 1481–1496 (1997)
Köhler, S., Bauer, S., Horn, D., Robinson, P.N.: Walking the interactome for prioritization of candidate disease genes. Am. J. Hum. Genet. 82(4), 949–958 (2008)
Lawrence, M.S., et al.: Discovery and saturation analysis of cancer genes across 21 tumour types. Nature 505(7484), 495–501 (2014)
Lazareva, O., Baumbach, J., List, M., Blumenthal, D.B.: On the limits of active module identification. Briefings Bioinf. 22(5), bbab066 (2021)
Lee, I., Blom, U.M., Wang, P.I., Shim, J.E., Marcotte, E.M.: Prioritizing candidate disease genes by network-based boosting of genome-wide association data. Genome Res. 21(7), 1109–1121 (2011)
Leiserson, M.D.M., Vandin, F., Wu, H.T., Dobson, J.R., et al.: Pan-cancer network analysis identifies combinations of rare somatic mutations across pathways and protein complexes. Nat. Genetics 47(2), 106–114 (2015)
Leiserson, M.D., Eldridge, J.V., Ramachandran, S., Raphael, B.J.: Network analysis of GWAS data. Curr. Opin. Genet. Dev. 23(6), 602–610 (2013)
Levi, H., Elkon, R., Shamir, R.: DOMINO: a network-based active module identification algorithm with reduced rate of false calls. Mol. Syst. Biol. 17(1), e9593 (2021)
Liu, Y., et al.: SigMod: an exact and efficient method to identify a strongly interconnected disease-associated module in a gene network. Bioinformatics 33(10), 1536–1544 (2017)
Luo, Y., et al.: A network integration approach for drug-target interaction prediction and computational drug repositioning from heterogeneous information. Nat. Commun. 8(1), 573 (2017)
McLachlan, G., Bean, R.W., Jones, L.B.T.: A simple implementation of a normal mixture approach to differential gene expression in multiclass microarrays. Bioinformatics 22(13), 1608–1615 (2006)
Menche, J., et al.: Uncovering disease-disease relationships through the incomplete human interactome. Science 347(6224), 1257601 (2015)
Mitra, K., Carvunis, A.R., Ramesh, S.K., Ideker, T.: Integrative approaches for finding modular structure in biological networks. Nat. Rev. Genet. 14(10), 719–732 (2013)
Nabieva, E., Jim, K., Agarwal, A., Chazelle, B., Singh, M.: Whole-proteome prediction of protein function via graph-theoretic analysis of interaction maps. Bioinformatics 21, i302–i310 (2005)
Nibbe, R.K., Koyutürk, M., Chance, M.R.: An integrative-omics approach to identify functional sub-networks in human colorectal cancer. PLoS Comput. Biol. 6(1), e1000639 (2010)
Nikolayeva, I., Pla, O.G., Schwikowski, B.: Network module identification-a widespread theoretical bias and best practices. Methods 132, 19–25 (2018)
Page, L., Brin, S., Motwani, R., Winograd, T.: The PageRank citation ranking: Bringing order to the web. Technical report 1999-66, Stanford InfoLab, November 1999
Pan, W., Lin, J., Le, C.T.: A mixture model approach to detecting differentially expressed genes with microarray data. Funct. Integr. Genomics 3(3), 117–124 (2003). https://doi.org/10.1007/s10142-003-0085-7
Paull, E.O., Carlin, D.E., Niepel, M., Sorger, P.K., Haussler, D., Stuart, J.M.: Discovering causal pathways linking genomic events to transcriptional states using Tied Diffusion Through Interacting Events (TieDIE). Bioinformatics 29(21), 2757–2764 (2013)
Picart-Armada, S., Barrett, S.J., Willé, D.R., Perera-Lluna, A., Gutteridge, A., Dessailly, B.H.: Benchmarking network propagation methods for disease gene identification. PLoS Comput. Biol. 15(9), 1–24 (2019)
Pounds, S., Morris, S.W.: Estimating the occurrence of false positives and false negatives in microarray studies by approximating and partitioning the empirical distribution of p-values. Bioinformatics 19(10), 1236–1242 (2003)
Radivojac, P., et al.: A large-scale evaluation of computational protein function prediction. Nat. Methods 10(3), 221–227 (2013)
Reyna, M.A., Chitra, U., Elyanow, R., Raphael, B.J.: NetMix: a network-structured mixture model for reduced-bias estimation of altered subnetworks. J. Computat. Biol. 28(5), 469–484 (2021)
Reyna, M.A., Leiserson, M.D., Raphael, B.J.: Hierarchical HotNet: identifying hierarchies of altered subnetworks. Bioinformatics 34(17), i972–i980 (2018)
Robinson, S., Nevalainen, J., Pinna, G., Campalans, A., Radicella, J.P., Guyon, L.: Incorporating interaction networks into the determination of functionally related hit genes in genomic experiments with Markov random fields. Bioinformatics 33(14), i170–i179 (2017)
Sharan, R., Ulitsky, I., Shamir, R.: Network-based prediction of protein function. Mol. Syst. Biol. 3, 88 (2007)
Sharpnack, J., Krishnamurthy, A., Singh, A.: Near-optimal anomaly detection in graphs using Lovász extended scan statistic. In: Proceedings of the 26th International Conference on Neural Information Processing Systems, NIPS 2013, vol. 2. pp. 1959–1967 (2013)
Sharpnack, J., Rinaldo, A., Singh, A.: Detecting anomalous activity on networks with the graph Fourier scan statistic. IEEE Trans. Signal Process. 64(2), 364–379 (2016)
Sharpnack, J., Singh, A., Rinaldo, A.: Changepoint detection over graphs with the spectral scan statistic. In: Artificial Intelligence and Statistics, pp. 545–553 (2013)
Shrestha, R., et al.: HIT’nDRIVE: patient-specific multidriver gene prioritization for precision oncology. Genome Res. 27(9), 1573–1588 (2017)
Szklarczyk, D., et al.: STRING v10: protein-protein interaction networks, integrated over the tree of life. Nucleic Acids Res. 43(D1), D447–D452 (2015)
Tate, J.G., et al.: COSMIC: the catalogue of somatic mutations in cancer. Nucleic Acids Res. 47(D1), D941–D947 (2019)
Ulitsky, I., Shamir, R.: Identification of functional modules using network topology and high-throughput data. BMC Syst. Biol. 1(1), 8 (2007). https://doi.org/10.1186/1752-0509-1-8
Vandin, F., Clay, P., Upfal, E., Raphael, B.J.: Discovery of mutated subnetworks associated with clinical data in cancer. In: Pacific Symposium on Biocomputing, vol. 17, pp. 55–66 (2012)
Vandin, F., Upfal, E., Raphael, B.J.: Algorithms for detecting significantly mutated pathways in cancer. J. Comput. Biol. 18(3), 507–522 (2011)
Vandin, F., Upfal, E., Raphael, B.J.: De novo discovery of mutated driver pathways in cancer. Genome Res. 22(2), 375–385 (2012)
Vanunu, O., Magger, O., Ruppin, E., Shlomi, T., Sharan, R.: Associating genes and protein complexes with disease via network propagation. PLoS Comput. Biol. 6(1), e1000641 (2010)
Velghe, A., et al.: PDGFRA alterations in cancer: characterization of a gain-of-function V536E transmembrane mutant as well as loss-of-function and passenger mutations. Oncogene 33(20), 2568–2576 (2014)
Vlaic, S., et al.: ModuleDiscoverer: identification of regulatory modules in protein-protein interaction networks. Sci. Rep. 8(1), 433 (2018)
Wang, X., Terfve, C., Rose, J.C., Markowetz, F.: HTSanalyzeR: an R/Bioconductor package for integrated network analysis of high-throughput screens. Bioinformatics 27(6), 879–880 (2011)
Weston, J., Elisseeff, A., Zhou, D., Leslie, C.S., Noble, W.S.: Protein ranking: from local to global structure in the protein similarity network. Proc. Nat. Acad. Sci. 101(17), 6559–6563 (2004)
Xia, J., Gill, E.E., Hancock, R.E.W.: NetworkAnalyst for statistical, visual and network-based meta-analysis of gene expression data. Nat. Protoc. 10(6), 823–844 (2015)
Zhou, D., Bousquet, O., Lal, T., Weston, J., Schölkopf, B.: Learning with local and global consistency. In: Advances in Neural Information Processing Systems, vol. 16. MIT Press (2004)
Acknowledgement
The authors would like to thank Jasper C. H. Lee and Christopher Musco for helpful discussions, as well as Matthew A. Myers and Palash Sashittal for reviewing early versions of the manuscript. U.C. is supported by NSF GRFP DGE 2039656. B.J.R. is supported by grant U24CA264027 from the National Cancer Institute (NCI).
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2022 The Author(s), under exclusive license to Springer Nature Switzerland AG
About this paper
Cite this paper
Chitra, U., Park, T.Y., Raphael, B.J. (2022). NetMix2: Unifying Network Propagation and Altered Subnetworks. In: Pe'er, I. (eds) Research in Computational Molecular Biology. RECOMB 2022. Lecture Notes in Computer Science(), vol 13278. Springer, Cham. https://doi.org/10.1007/978-3-031-04749-7_12
Download citation
DOI: https://doi.org/10.1007/978-3-031-04749-7_12
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-031-04748-0
Online ISBN: 978-3-031-04749-7
eBook Packages: Computer ScienceComputer Science (R0)