Abstract
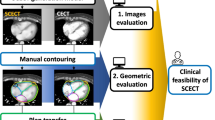
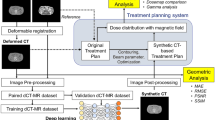
Radiation therapy is one of the most important strategies for treating patients with tumor. The rationale is to deliver high radiation doses to the tumor in order to damage its DNA while sparing, at the same time, healthy tissues. In order to optimize such a process, biomedical images play a fundamental role; in particular, Magnetic Resonance (MR) produces well-contrasted images for precisely contouring tumors and organs at risk. However, due to the physical information stored in it, Computed Tomography (CT) is mandatory for accurate radiation dose calculation. To overcome this limitation, several algorithms, usually based on deep learning techniques, were proposed for converting MR–to–synthetic CT (sCT). In this paper, we report about the evaluation of the impact of three different train losses, commonly used for non–medical applications. Tests were ran on a cohort of 15 brain MR/CT image pairs. An algorithm for MR–to–sCT conversion, previously developed for MR-only radiotherapy, was used as benchmark platform. Predicted sCT images were compared on the basis of intensities and edges reconstruction. Results show that potential improvements can be achieved if non–medical image loss will be adapted to this application.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
Notes
- 1.
Two types of MR acquisitions that, by taking into account different matter properties, lead to different contrast among tissues.
References
Yang, X., et al.: MRI-based proton radiotherapy for prostate cancer using deep convolutional neural networks. Int. J. Radiat. Oncol. Biol. Phys. 105, S200 (2019)
Sjölund, J., Forsberg, D., Andersson, M., Knutsson, H.: Generating patient specific pseudo-CT of the head from MR using atlas-based regression. Phys. Med. Biol. 60, 825–839 (2015)
Dowling, J.A., et al.: Automatic substitute computed tomography generation and contouring for magnetic resonance imaging (MRI)-alone external beam radiation therapy from standard MRI sequences. Int. J. Radiat. Oncol. Biol. Phys. 93, 1144–1153 (2015)
Arabi, H., Koutsouvelis, N., Rouzaud, M., Miralbell, R., Zaidi, H.: Atlas-guided generation of pseudo-CT images for MRI-only and hybrid PET-MRI-guided radiotherapy treatment planning. Phys. Med. Biol. 61, 6531–6552 (2016)
Burgos, N., et al.: Iterative framework for the joint segmentation and CT synthesis of MR images: application to MRI-only radiotherapy treatment planning. Phys. Med. Biol. 62, 4237 (2017)
Han, X.: MR-based synthetic CT generation using a deep convolutional neural network method. Med. Phys. 44, 02 (2017)
Fritscher, K., Raudaschl, P., Zaffino, P., Spadea, M.F., Sharp, G.C., Schubert, R.: Deep neural networks for fast segmentation of 3D medical images. In: Ourselin, S., Joskowicz, L., Sabuncu, M.R., Unal, G., Wells, W. (eds.) MICCAI 2016. LNCS, vol. 9901, pp. 158–165. Springer, Cham (2016). https://doi.org/10.1007/978-3-319-46723-8_19
Zaffino, P., et al.: Multi atlas based segmentation: should we prefer the best atlas group over the group of best atlases? Phys. Med. Biol. 63(12), 12NT01 (2018)
Zaffino, P., et al.: Radiotherapy of Hodgkin and non-Hodgkin lymphoma: a nonrigid image-based registration method for automatic localization of prechemotherapy gross tumor volume. Technol. Cancer Res. Treat. 15(2), 355–364 (2016)
Spadea, M.F., et al.: Contrast-enhanced proton radiography for patient set-up by using x-ray CT prior knowledge. Int. J. Radiat. Oncol. Biol. Phys. 90(3), 628–636 (2014)
Zaffino, P., et al.: Fully automatic catheter segmentation in MRI with 3D convolutional neural networks: application to MRI-guided gynecologic brachytherapy. Phys. Med. Biol. 64(16), 165008 (2019)
Bruno, P., et al.: Using CNNs for designing and implementing an automatic vascular segmentation method of biomedical images. In: Ghidini, C., Magnini, B., Passerini, A., Traverso, P. (eds.) AI*IA 2018. LNCS (LNAI), vol. 11298, pp. 60–70. Springer, Cham (2018). https://doi.org/10.1007/978-3-030-03840-3_5
Tappeiner, E., et al.: Multi-organ segmentation of the head and neck area: an efficient hierarchical neural networks approach. Int. J. Comput. Assist. Radiol. Surg. 14(5), 745–754 (2019). https://doi.org/10.1007/s11548-019-01922-4
Fritscher, K.D., Peroni, M., Zaffino, P., Spadea, M.F., Schubert, R., Sharp, G.: Automatic segmentation of head and neck CT images for radiotherapy treatment planning using multiple atlases, statistical appearance models, and geodesic active contours. Med. Phys. 41(5), 051910 (2014)
Raudaschl, P.F., et al.: Evaluation of segmentation methods on head and neck CT: auto-segmentation challenge 2015. Med. Phys. 44(5), 2020–2036 (2017)
Ciardo, D., et al.: Atlas-based segmentation in breast cancer radiotherapy: evaluation of specific and generic-purpose atlases. Breast 32, 44–52 (2017)
Milletari, F., Navab, N., Ahmadi, S.-A.: V-net: fully convolutional neural networks for volumetric medical image segmentation. In: 2016 Fourth International Conference on 3D Vision (3DV), pp. 565–571. IEEE (2016)
Litjens, G., et al.: A survey on deep learning in medical image analysis. Med. Image Anal. 42, 60–88 (2017)
Lee, H., Chen, Y.-P.P.: Image based computer aided diagnosis system for cancer detection. Expert Syst. Appl. 42(12), 5356–5365 (2015)
Moccia, S., Penza, V., Vanone, G.O., De Momi, E., Mattos, L.S.: Automatic workflow for narrow-band laryngeal video stitching. In: 2016 38th Annual International Conference of the IEEE Engineering in Medicine and Biology Society (EMBC), pp. 1188–1191. IEEE (2016)
Pileggi, G., et al.: Proton range shift analysis on brain pseudo-CT generated from T1 and T2 MR. Acta Oncologica 57(11), 1521–1531 (2018)
Spadea, M.F., et al.: Deep convolution neural network (DCNN) multiplane approach to synthetic CT generation from MR images-application in brain proton therapy. Int. J. Radiat. Oncol. Biol. Phys. 105(3), 495–503 (2019)
Johnstone, E., et al.: Systematic review of synthetic computed tomography generation methodologies for use in magnetic resonance imaging-only radiation therapy. Int. J. Radiat. Oncol. Biol. Phys. 100(1), 199–217 (2018)
Edmund, J.M., Nyholm, T.: A review of substitute CT generation for MRI-only radiation therapy. Radiat. Oncol. 12(1), 28 (2017). https://doi.org/10.1186/s13014-016-0747-y
Dowling, J.A., et al.: An atlas-based electron density mapping method for magnetic resonance imaging (MRI)-alone treatment planning and adaptive MRI-based prostate radiation therapy. Int. J. Radiat. Oncol. Biol. Phys. 83(1), e5–e11 (2012)
Xiang, L., Wang, Q., Nie, D., Qiao, Y., Shen, D.: Deep embedding convolutional neural network for synthesizing CT image from t1-weighted MR image. CoRR, vol. abs/1709.02073 (2017)
Wolterink, J.M., Dinkla, A.M., Savenije, M.H.F., Seevinck, P.R., van den Berg, C.A.T., Išgum, I.: Deep MR to CT synthesis using unpaired data. In: Tsaftaris, S.A., Gooya, A., Frangi, A.F., Prince, J.L. (eds.) SASHIMI 2017. LNCS, vol. 10557, pp. 14–23. Springer, Cham (2017). https://doi.org/10.1007/978-3-319-68127-6_2
Emami, H., Dong, M., Nejad-Davarani, S., Glide-Hurst, C.: Generating synthetic CTs from magnetic resonance images using generative adversarial networks. Med. Phys. 45, 06 (2018)
Zaffino, P., Raudaschl, P., Fritscher, K., Sharp, G.C., Spadea, M.F.: Plastimatch mabs, an open source tool for automatic image segmentation. Med. Phys. 43(9), 5155–5160 (2016)
Johnson, J., Alahi, A., Fei-Fei, L.: Perceptual losses for real-time style transfer and super-resolution. In: Leibe, B., Matas, J., Sebe, N., Welling, M. (eds.) ECCV 2016. LNCS, vol. 9906, pp. 694–711. Springer, Cham (2016). https://doi.org/10.1007/978-3-319-46475-6_43
Thummerer, A., et al.: Comparison of CBCT based synthetic CT methods suitable for proton dose calculations in adaptive proton therapy. Phys. Med. Biol. 65, 095002 (2020)
Ronneberger, O., Fischer, P., Brox, T.: U-net: convolutional networks for biomedical image segmentation. CoRR, vol. abs/1505.04597 (2015)
Simonyan, K., Zisserman, A.: Very deep convolutional networks for large-scale image recognition. arXiv preprint arXiv:1409.1556 (2014)
Canny, J.: A computational approach to edge detection. IEEE Trans. Pattern Anal. Mach. Intell. 6, 679–698 (1986)
Dice, L.R.: Measures of the amount of ecologic association between species. Ecology 26(3), 297–302 (1945)
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2020 Springer Nature Switzerland AG
About this paper
Cite this paper
Sherwani, M.K., Zaffino, P., Bruno, P., Spadea, M.F., Calimeri, F. (2020). Evaluating the Impact of Training Loss on MR to Synthetic CT Conversion. In: Nicosia, G., et al. Machine Learning, Optimization, and Data Science. LOD 2020. Lecture Notes in Computer Science(), vol 12565. Springer, Cham. https://doi.org/10.1007/978-3-030-64583-0_50
Download citation
DOI: https://doi.org/10.1007/978-3-030-64583-0_50
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-030-64582-3
Online ISBN: 978-3-030-64583-0
eBook Packages: Computer ScienceComputer Science (R0)