Abstract
The stunning ability of bacteria to evolve and adapt has contributed to the success of these single cells, which have inhabited the Earth for billions of years and play vital roles in the environment and in human health. The goal of this chapter is to present and discuss the population-level organizational scheme of bacterial pangenomes, wherein genes are distributed among the strains of a species, such that each individual strain encodes only a subset of the genes available at the population level. Genes from the accessory/distributed genome (those present only in a subset of strains within a species) impart diverse functions or variations on a conserved function to strains. Moreover, horizontal gene transfer generates novel gene combinations. The maintenance and spread of any given gene arrangement are influenced by fitness. Further, the extent of genomic plasticity is regulated by restriction modification systems, phage-defense systems, and Clustered Regularly Interspaced Short Palindromic Repeats (CRISPR)—associated proteins (CRISPR-Cas). The combination of a pangenome structure and genomic plasticity reveals a successful strategy for bacterial adaptation to ever-changing environments. From a clinical perspective, pangenome analyses inform the selection of therapeutic targets, designed to focus either on an entire species or on virulence features within a species. Further, they provide a framework for modeling the efficacy of drugs and vaccines. In summary, following the explosion in sequencing technology, pangenome studies have revealed remarkable genomic organizations at the levels of species, with important implications to our understanding of evolution, and our ability to design therapeutics and predict their long-term outcomes.
You have full access to this open access chapter, Download chapter PDF
Similar content being viewed by others
Keywords
1 Introduction
Bacteria dominate our planet and can be traced back to billions of years in the geological record. They play critical roles in shaping our habitat, from adding oxygen to the atmosphere to fixing nitrogen in the soil. They also play a vital role in human health, with commensal/mutualistic bacteria influencing nutrition and immunity, and pathogenic bacteria causing diseases from epidemics like the Black Death of medieval times to modern-day chronic biofilm infections resulting in the spread of antibiotic resistance. A defining characteristic of bacteria in both the environment and health is their ability to rapidly evolve and adapt. Here we discuss the elegant population-level organizational scheme that bacterial species use wherein their genomes are distributed among large numbers of strains, with no single strain having more than a small minority of genes available at the population level. This distributed pan(supra)-genome provides for adaptation to countless novel challenges and environmental niches.
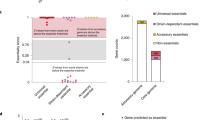
Individual bacterial genomes have a discrete number of genes. However, enormous differences in gene content exist even among the genomes of strains of a single species. Therefore, the gene content of a single strain is less than the full complement of different genes from all strains. The comprehensive set of genes within a species, i.e., all genes from all strains, is defined as the pangenome (or supragenome). The pangenome is organized into the core genome, which corresponds to the set of genes conserved across all strains in the species, and the accessory genome (or distributed genome), which are all noncore genes. We compiled pangenome papers from PubMed, identifying 295 species-specific pangenome projects performed on approximately 70 genera (Fig. 1). In all of these projects, the pangenome was found to be substantially larger than the core genome (Fig. 2).
Overview of the number of pangenomic studies per genus from a literature search between 2005 and 2018 (See “References for Fig. 1”)
The diversity within a species’ pangenome provides a reservoir of genetic material available to bacterial cells to respond to selective pressures. Horizontal gene transfer (HGT) is the process by which individual bacterial cells can uptake genetic material from their environment or neighboring bacteria and generate novel, strain-specific gene combinations. It seems logical that when HGT occurs among strains of the same species these events are more likely to be adaptive or work in concert within the biological network, when compared to random mutations or genes acquired from distantly related species. This has been demonstrated to be the case in multiple species, where the majority of accessory genes appear to be evolving in tandem with the core genome (Gladitz et al. 2005). In this manner, the pangenome allows a species to incorporate more solutions to environmental stresses and niches than can be encoded by a single strain (Ehrlich et al. 2005, 2010).
2 Steps in the Assembly of a Pangenome
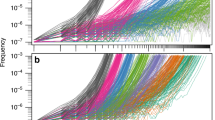
Pangenome analyses are performed on a set of strains from the same species, or very closely related species (often different species grouped together by genus, though we will not be examining those projects here). The set of all coding sequences (CDS) are clustered by sequence similarity with the objective of generating groups of orthologous genes. This is a multistep process that begins with whole-genome sequencing (WGS) of multiple independent bacterial (nonclonal, nonderivative) strains selected to represent the broadest geographic and phenotypic ranges of the species of interest. Following sequencing, the remaining steps are computational and include (1) assembly of genomes into contigs, (2) annotation of protein-coding sequences (CDS), and (3) clustering of CDSs based on the sequence similarity of nucleic acids or amino acids of their cognate encoded proteins. Once clusters are defined, they are classified based on strain prevalence into core or accessory (distributed) clusters. The accessory/distributed set of gene clusters is often further organized into those that are widely distributed (near core/soft core) in a population and those that are rare (shell) or unique (Fig. 3).
Histogram of the number of gene clusters present in a given number of genomes. Taken from a project examining 12 genomes of Moraxella catarrhalis (Davie et al. 2011), with a total of 2383 gene clusters
The tools and the parameters used to characterize gene clusters vary widely among projects (Fig. 4). Generally, the first project(s) within a species tend to focus on the basic characterization of the pangenome. Subsequent projects often emphasize specific areas of interest, such as the distribution of virulence factors, levels of horizontal gene transfer, or epigenetic factors. Our survey of 295 pangenome projects did not reveal a strong preference for any individual assembly program. This is likely because assembly programs and versions perform differently depending on the examined species and the employed DNA sequencing technology. Further, many pangenome projects utilize pre-assembled genomes from publicly available databases (GenBank, EMBL, DDJB, JGI, PubMLST, etc.). This survey found that the CD-HIT program was the most frequently used gene clustering software, though a diverse set of other programs were also utilized for this purpose. Finally, commonly used software for other analyses include gene annotation (RAST, Prokka, PHAST, and Prodigal) (Aziz et al. 2008; Seemann 2014; Zhou et al. 2011; Hyatt et al. 2010), genome/gene alignments (Muscle, Mauve, Mega, and ClustalW) (Edgar 2004; Darling et al. 2004; Kumar et al. 1994; Higgins and Sharp 1988), and phylogenetic tree building (Mega, RAxML, and PhyML) (Kumar et al. 1994; Stamatakis 2006; Guindon et al. 2010). Overall, there is high variability in the methods/software used for pangenome analyses, reflecting diversity in the scope and goals of these projects.
3 Size of the Pangenome
The size of a species’ pangenome, relative to the size of the core genome, is highly variable across the eubacteria. In Fig. 2, we display the variability we encountered in 295 species-specific pangenome projects (Figs. 1 and 2). Papers included in this summary span from 2005 [when the first pangenomes were described in S. agalactiae (Tettelin et al. 2005) and H. influenzae (Shen et al. 2005; Hogg et al. 2007)] through 2018. In all cases, the pangenome was significantly larger than the set of genes in a given strain. The size of the core genomes ranged from <20 to >60% of the pangenome (Fig. 2).
In some cases, calculations on the size of the pangenome may reflect inaccuracies in the current taxonomy, instead of the underlying biology. An instance of high genomic diversity is observed with Gardnerella vaginalis, where only 27% (746/2792) of its gene clusters are core (Ahmed et al. 2012). It is likely that G. vaginalis appears so genomically diverse because traditional biochemical tests used to identify strains within this taxa were unable to distinguish among the multiple genomically diverse species that are actually present. Thus, in this case, the apparent large size of the pangenome (and the corresponding small size of the core genome) arose from the unintentional merging of multiple species into a single species. In contrast, instances of low genomic diversity are observed in the genus Bacillus. Both Bacillus anthracis and Bacillus thuringiensis closely resemble B. cereus (Vilas-Bôas et al. 2007). B. thuringiensis appears to correspond to multiple phylogenetic clades (lineages) within B. cereus. B. anthracis (a species with one of the smallest pangenomes) likely represents a single phylogenetic lineage within the broader, more diverse definition of B. cereus that acquired a clinically important set of toxin genes (Okinaka and Keim 2016; Hall et al. 2010).
It is tempting to speculate that there are general principles that directly associate the size of the pangenome with the biology of the species. Factors that may play a substantial role are the extent of gene transfer, the degree of interactions with competing and cooperating species, the number of niches inhabited, or the lifestyle of the bacterium. The hypothesis that highly specialized environments lead to smaller genome sizes has been explored in the context of obligate intracellular species and pathogens (Merhej et al. 2009; Georgiades et al. 2011). A study of overall differences between the genomes of 12 highly pathogenic species compared to their most closely related nonpathogenic cousins found that, for the sets of bacteria studied, the most virulent species generally had smaller genomes, which suggests gene loss as well as loss-of-function mutations (Georgiades and Raoult 2011). The reduced genome size is hypothesized to be a consequence of extreme specialization of the pathogens to their hosts, while the less-specialized nonpathogens show greater levels of genomic variation due to selective pressure to remain competitive in more diverse environments (Georgiades and Raoult 2011). While this is an interesting idea, not all studies point to a relationship between pathogenicity and genome size (Bonar et al. 2018).
In a related vein, longitudinal comparative genomic studies of pathogenic clonal lineages of Pseudomonas aeruginosa, Burkholderia sp., and Haemophilus influenzae have captured microevolution and host adaptation in the human lung (Rau et al. 2012; Lee et al. 2017; Pettigrew et al. 2018; Moleres et al. 2018; Bianconi et al. 2018; Burns et al. 2001; Li et al. 2005; Jorth et al. 2015; Silva et al. 2016). In many cases, these changes reveal gene deletions when compared to their antecedents. For instance, serial isolates of H. influenzae clonal lineages in COPD patients display a significant association with loss-of-function mutations in the ompP1 (fadL) accessory gene. fadL is beneficial to this bacterium in early infection, as it promotes adhesion and intracellular invasion via interactions with the epithelial cell ligand hCEACAM1 (human carcinoembryonic antigen-related cell adhesion molecule 1). In contrast, it may hinder long-term survival in the lung, as its expression increases sensitivity to arachidonic acid, an exogenous mammalian long-chain fatty acid with bactericidal effects (Moleres et al. 2018). This is indicative of selective pressure in favor of ompP1 function in the nasopharynx and against its function in the lungs. These observations support the general concept that gene loss may accompany the ability to survive within highly circumscribed niches (Rau et al. 2012; Lee et al. 2017; Pettigrew et al. 2018; Moleres et al. 2018). Nonetheless, one must keep in mind that evolution in niches that do not support transmission may not be relevant to the evolution of the pangenome. Large-scale comparative pangenome and evolutionary studies promise to reveal the rules that shape the overall pangenome size, as well as identify disease and tissue-specific genes (and gene losses).
4 The Accessory Genome and Functional Diversity
In general, core genomes are enriched for housekeeping functions. These include energy production, amino acid metabolism, nucleotide metabolism, lipid transport, and translational machinery. Accessory genomes often encode genes involved in protein trafficking and defense, as well as many niche-specific functions. Further, plasmids, phage, and transposons are also often associated with accessory genomes. This section focuses on functional diversity as it pertains to the accessory genome.
Phenotypic traits can result from a blend of core genes with highly variable accessory genes. This is exemplified by the production of the capsule (Swartley et al. 1997; Bentley et al. 2006), synthesis of the extracellular polymeric substance (EPS) (Harris et al. 2017), and modification of the cell wall (Gerlach et al. 2018). Here, conserved modules encoded in the core and softcore genomes are modified by components encoded by the accessory genome, providing a procedure to generate phenotypic variability. In Neisseria meningitidis, capsule biosynthesis genes are encoded within a single syntenic cps chromosomal region, which encodes both core and accessory genes. Variations in the accessory genes yield diversity in capsular types (Harrison et al. 2013). In Lactobacillus salivarius, the EPS cluster 2 contributes to the biofilm matrix. The genes at the extremities of this multigene cluster genes are core, while there is extensive variation in the genes encoded in the center of the cluster. These differences in glycotransferases and EPS biosynthesis-related proteins contribute to variations in the EPS structure (Harris et al. 2017). Yet another example is observed in methicillin-resistant Staphylococcus aureus (MRSA), where strains evade host immunity by modification of wall teichoic acid (WTA) using an alternative WTA glycosyltransferase encoded on a prophage (Gerlach et al. 2018). These studies exemplify how diversity within the accessory genome can provide bacteria with a blueprint to generate variability. This genomic flexibility is likely to increase the adaptive potential of bacterial species in the face of environmental stresses.
Genes encoded by the accessory genome can influence pathogenic potential. A well-studied example is Escherichia coli; this species encodes a highly diverse pangenome, where variability within the accessory genome leads to strains that differ in their ability to colonize human cell types and to trigger pathogenicity (Rasko et al. 2008). E. coli strains are grouped into pathovars based on the presence of virulence markers, often encoded on mobile elements (Kaper et al. 2004). Whole-genome comparative analyses of pathovars demonstrate that strains of the same pathovar are not always phylogenetically clustered (Rasko et al. 2008; Salipante et al. 2015; Hazen et al. 2013). This pattern of clustering is consistent with the transfer of accessory genes among E. coli strains, as well as the independent acquisition of virulence traits by strains in the same pathovar. One prominent example of HGT among E. coli strains of different pathovars is observed in the highly pathogenic strain that caused the 2011 German food poisoning outbreak (Mahan et al. 2013). Multiple genomic studies ultimately concluded that the outbreak was caused by a Shiga toxin-producing E. coli (STEC) of serotype O104:H4, which harbored multiple genes commonly associated with enteroaggregative E. coli (EAEC) including: a plasmid-encoded type I aggregative adherence fimbriae that mediate colonization and biofilm formation, assortment of serine proteases (SPATEs), and chromosomally encoded Shigella enterotoxin 1 (Askar et al. 2011; Mellmann et al. 2011; Rasko et al. 2011). Moreover, the prevalence of genetic transfer among E. coli strains is highlighted by the lack of an exclusive genomic signature among commensal E. coli strains. The strains that asymptomatically colonize the human gastrointestinal tract are genetically diverse (Rasko et al. 2008). These commensal strains may serve as genetic repositories for virulence determinants and, in addition, gene transfer events may modify their pathogenic potential and drug sensitivity. In conclusion, the accessory genome of E. coli is a critical determinant of tissue tropism, pathogenic potential, and clinical presentation.
Non-orthologous accessory genes with related functions are often syntenic across strains. We propose that this genomic configuration allows one variant to be switched by another in the process of recombination, where the neighboring genes provide an anchor for homologous recombination. One example is the genomic region that encodes the DpnI, DpnII, or the DpnIII type II restriction enzymes in S. pneumoniae. These loci differ in the sequence of the enzymes, the number of genes in the locus, and their ability to restrict phages or transforming DNA (Johnston et al. 2013a; Eutsey et al. 2015). Another example is the genomic region that encodes bacteriocins downstream of the blp histidine kinase signal transduction system in S. pneumoniae. While the genes in this region are predicted to be bacteriocins, the number of genes, their sequence, and the cells they target differ across strains (Lux et al. 2007; Dawid et al. 2007; Valente et al. 2016; Rezaei Javan et al. 2018). Other examples of this proposed mechanism, wherein conserved flanking genes anchor multiple variants of pathogenicity genes, include the parologous vHiSLR genes of H. influenzae (Kress-Bennett et al. 2016) and the bro gene variants of Moraxella catarrhalis (Earl et al. 2016). Syntenic regions that encode non-homologous genes within a single functional class may provide a pangenomic “switch,” allowing cells to flip between variants of a single function to optimize fitness in diverse niches.
In summary, many of the genes in the accessory genome provide new functions or variations on a conserved function in a manner that expands the ability of strains to survive or adapt in their environments. In this manner, the strain diversity resulting from variations in the accessory genome may serve as a population-level tool to ensure the survival of a bacterial species.
5 Pangenome Plasticity
Speaking teleologically, via intra- and inter-species gene transfer, individual bacterial strains can draw from an expanded set of genes for their own adaptation and evolutionary success. This phenomenon was observed as early as 1928 in the Griffith’s experiment, where a nonencapsulated strain of S. pneumoniae integrated DNA from an encapsulated isolate, leading to its conversion from avirulent to virulent (Griffith 1928). Almost a century later, the bacterial research community has described multitudinous instances of gene transfer among bacterial strains.
5.1 Gene Transfer Events Within and Across Species
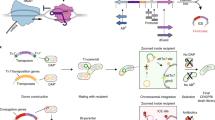
Gene transfer events can occur anywhere, and our literature review identified 19 manuscripts that describe bacterial in vivo gene transfer within human patients (Table 1). A common theme is the acquisition of antibiotic resistance; particularly in regard to carbapenems, β-lactamases, and quinolones. Resistance was commonly the result of genes acquired via bacteriophages, plasmids, or pathogenicity islands (Conlan et al. 2014; Bielaszewska et al. 2007; Datta et al. 2017; Feld et al. 2008; Langhanki et al. 2018; Mena et al. 2006; Neuwirth et al. 2001; Soto et al. 2011). In our set, five cases show HGT between different bacterial species: Serratia marcescens and Escherichia coli (Mata et al. 2010), two instances of Klebsiella pneumoniae and E. coli (Gona et al. 2014; Göttig et al. 2015), Staphylococcus aureus and Staphylococcus epidermidis (Hurdle et al. 2005), and Enterobacter cloacae and E. coli (Sidjabat et al. 2014). These studies highlight how bacteria occupying the same niche can evolve during the infectious disease process, posing new challenges for treatment.
Cross-species transfer events introduce new genes into the species, thus expanding the pangenome. A prominent example is acquisition of the type 3 secretion system (T3SS) by multiple Gram-negative bacteria. The T3SS allows for the transport of effector proteins from the bacterial cytosol directly into the host cells (Hacker et al. 1997; Hueck 1998). In most cases, the genes encoding this injection system, and their effectors, have been acquired by HGT (Brown and Finlay 2011). These T3SS systems are critical components of virulence. For instance, in Salmonella, acquisition of the SPI1 T3SS enables the bacterium to invade host cells, while acquisition of the SPI2 T3SS enables it to escape host defenses and survive within host cells inside a protective vacuole (Jennings et al. 2017; Ochman et al. 1996). Another example of cross-species transfer has been observed in S. pneumoniae, where a multigene locus was acquired from Streptococcus suis (Antic et al. 2017). This locus was acquired exclusively by a phylogenetically distinct subset of strains within the S. pneumoniae species—a subset much more likely to infect the conjunctiva. The genes acquired from S. suis appear to contribute to the tissue tropism by promoting adherence to the ocular epithelium. Thus, expansion of the pangenome by gene acquisition from outside the species can contribute to bacterial virulence and tropism.
Gene transfer among strains of the same species provides a mechanism to redistribute accessory/distributed genes within single strains. Studies on vaccine-escape strains of S. pneumoniae identified multiple genes acquired from a single donor (Golubchik et al. 2012). These recombination events ranged from 0.04 to 44 kb in size, and were located in various regions of the genome, including the capsular locus. Separate analyses of whole genomes of S. pneumoniae have captured multiple instances of serotype switches including from 23F to 3 and from 19F to 19A (Chewapreecha et al. 2014; Croucher et al. 2014a; Hiller et al. 2011). A current vaccine targets the 19F capsule, but not the 19A. Serotype 19F strains were widely prevalent pre-vaccine, while serotype 19A strains have spread in the USA during the post-vaccine era (Geno et al. 2015). This serotype switch has been observed in vaccinated and non-vaccinated populations. These observations are consistent with a model where HGT generates diverse genotypes, selective pressure from vaccines drives the spread of a subset of strains, and competition across strains shape the population and distribution of accessory genes.
Studies that describe recombination among strains driven by natural competence and transformation suggest that multiple transfers may occur both simultaneously and sequentially between individual donors and recipient strains. A study on S. pneumoniae captured the progressive accumulation of recombinations in a set of six clinical strains isolated from a pediatric patient over a 7-month period. One strain incurred multiple recombination events from the same donor, over two instances of recombination. These events introduced recombinations at 23 sites, and led to the exchange of over 7% of the genome (Hiller et al. 2010). Similarly, a laboratory study in H. influenzae also captured multiple gene transfer events after a bout of recombination (Mell et al. 2011). For this study, DNA from a clinical strain was used to transform a laboratory strain. Transformants were observed to have multiple recombination events over the length of the chromosome, collectively corresponding to ~1–3% of the genome. These analyses not only demonstrate HGT events across strains, but also suggest that strains may display multiple transfers during a single competence event.
HGT occurring through natural competence and transformation is unique among HGT mechanisms, in that it is driven by the recipient as opposed to by the donor (as is the case with mating and transduction). This means that it is an expressed phenotype that is triggered by the recipient cell. Thus, as a mechanism of mutation and evolution, it is expressed when a cell is stressed and provides a genetic means to adapt to a stressful environment resulting in mutation-on-demand (Ehrlich et al. 2005).
5.2 Constraints on Gene Transfer
While there is clear evidence of HGT among strains of the same species, distributed genes are not randomly distributed within a species. Instead, they tend to be associated with specific lineages, suggesting that pangenome evolution operates with forces that promote as well as limit gene transfer (Croucher et al. 2014b), as discussed in the next paragraphs.
There is increasing evidence that co-selection of genes limits gene transfer. A genome-wide study in S. pneumoniae demonstrated that a set of 876 loci, annotated to function in metabolism or transport, displayed a nonrandom distribution (Watkins et al. 2015). The authors show that groups of coevolved genes (alleles) are adapted to particular metabolic niches. They predict that disruption of these groups of alleles, a process mediated by HGT, would lead to a drop in strain fitness. A computational approach applied to S. pneumoniae and N. meningitidis also uncovered co-selection of genes associated with drug resistance and virulence (Pensar et al. 2019). Genome architecture may also limit gene transfer. Many bacterial genomes encode short sequences that are enriched in close proximity to the replication terminus. The location of these sequences is under selection, such that HGT events that disrupt these elements impose a fitness cost (Hendrickson et al. 2018). Thus, allele co-selection and genomic architecture illustrate genome-wide features that, when disturbed, can result in loss of fitness and consequently restrict gene flow.
In addition to factors that limit gene transfer via their influence on fitness, bacteria encode genes that serve as barriers to incoming DNA, such as restriction modification systems (RM), phage-defense systems, and Clustered Regularly Interspaced Short Palindromic Repeats (CRISPR)—associated proteins (CRISPR-Cas). Most RM and CRISPR-Cas systems exert their influence on double-stranded DNA. While DNA entering the cell by transformation is single stranded, these systems still appear to serve as barriers to transformation; a compelling model proposed that they do so via their activity on the transformed chromosome (Johnston et al. 2013b). Studies in N. meningitidis and S. pneumoniae illustrate the role of restriction modification (RM) systems in limiting HGT. Strains of N. meningitidis organize into distinct phylogenetic groups that are associated with the distribution of >20 RM systems (Budroni et al. 2011). This distribution is consistent with the hypothesis that the RM systems limit HGT among clades. Similarly, the PMEN1 pandemic lineage of S. pneumoniae displays asymmetric gene transfer. The heterologous gene transfer from PMEN1 to other strains is abundant, yet into PMEN1 is modest (Wyres et al. 2012). The DpnIII RM system contributes to this structure, as it appears to limits HGT into PMEN1 strains, and is almost exclusively found in the PMEN1 lineage (Eutsey et al. 2015). Type I RM systems can also limit gene transfer, however, their architecture may allow rapid evolution of HGT barriers. The type I RM systems have a multifunctional component, where modification in one sequence can lead to both changes in methylation and endonuclease activity. This is in contrast to type II RM systems, where the protein that directs methylation is distinct from the protein that directs endonuclease activity, such that changes in specificity require mutations in more than one protein (Wilson and Murray 2003). In this manner, type I RM systems can rapidly evolve new specificities and generate diversity. A recent study in S. pneumoniae demonstrated that phase variation in the SpnIV phase-variable Type I RM limits acquisition of genomic islands by transformation (Kwun et al. 2018). The work captures an instance of phase variation on a type I RM system that generated an HGT barrier between nearly identical strains. Together, these studies suggest that RM systems may foster genomic stability within subsets of strains.
Many bacteria encode an abortive infection (Abi) system, which appears to be altruistic mechanism to protect the population at-large. When bacteria possessing an Abi system are infected by phage, the system is activated and triggers the death of the bacterial host. In this manner, death of the infected isolate avoids spread of the phage across the bacterial community (Chopin et al. 2005). In an exciting twist, phage defense systems may also be encoded by prophage, illustrating cooperation between bacteria and phage to restrict unrelated phages (Dedrick et al. 2017; Bondy-Denomy et al. 2016).
CRISPR-Cas confers adaptive immunity in prokaryotes and has the ability to inhibit conjugation, transduction and transformation. The CRISPR-Cas are composed of arrays of palindromic nucleotide repeats that are interspersed by short unique DNA segments called spacers, and cas genes. The spacers are acquired from foreign DNA, usually bacteriophages. Following acquisition, spacers are transcribed and processed into small CRISPR RNA (crRNA) molecules. A complex formed by Cas proteins and crRNA leads to the degradation of invading foreign nucleic acid, protecting cells from future invasion (Jiang and Doudna 2017; Adli 2018). Many bacterial species and lineages are devoid of CRISPR-Cas systems. In vitro studies in multiple bacteria reveal an inverse correlation between HGT and the presence of a functional CRISPR-Cas system (Jiang et al. 2013; Watson et al. 2018). In Enterococcus faecalis, multidrug-resistant plasmids were observed in strains that lacked CRISPR-Cas systems, while the drug-sensitive strains encoded this system (Palmer and Gilmore 2010). Further, under selective pressure for the acquisition of antibiotic-resistant plasmids, Staphylococcus epidermidis strains acquired inactivating mutations in the CRISPR-Cas system (Jiang et al. 2013). These studies suggest that bacteria encounter a tradeoff: the fitness advantages associated with phage resistance afforded by CRISPR-Cas must be balanced against a decrease in genomic plasticity and the benefits conferred by acquisition of novel genes. Nonetheless, the role of phage protection systems in restricting gene flow is far from fully resolved. Some studies find contrasting results, and do not support the conclusion that CRISPR-Cas limits HGT. A large-scale computational study revealed that the activity of the CRISPR-Cas system was not associated with HGT events over long evolutionary timescales (Gophna et al. 2015). Further, a study in Pectobacterium atrosepticum suggests that CRISPR-Cas systems may actually contribute to HGT via their role in protecting bacteria against phage attack (Watson et al. 2018). Thus, more research is required to determine the ultimate influence of CRISPR-Cas systems on the genomic plasticity of bacterial populations.
In conclusion, the set of genes in a species’ pangenome can expand via the introduction of genes from other species, rearrange across strains via an intra-species exchange, or vary with mutations. The shuffling of accessory genes and alleles generates new combinations that are subsequently subjected to the forces of selection on gene products and genome-wide features. Moreover, RMs, CRISPR-Cas, and phage-defense systems may also influence gene flow across strains and species. All factors combined, genomic plasticity emerges as a successful strategy for bacterial survival.
6 A Balance in the Accessory Genome
A remarkable observation comes from recent mathematical models and population studies. Negative frequency-dependent selection may stabilize the proportion of individual accessory genes in a population of S. pneumoniae (Azarian et al. 2018; Corander et al. 2017). As expected, the authors observed that vaccination led to a dramatic drop in the representation of vaccine-sensitive strains. In doing so, the distribution of accessory genes within the population differed from that of the pre-vaccine population. Interestingly, over time, the frequency of the accessory genes trended toward that seen in the pre-vaccine population. These results suggest that the distribution of genes in the pneumococcal pangenome may have an equilibrium point. It remains to be determined whether similar patterns are observed in other species. The suggestion that the composition of pangenomes tends toward an equilibrium has important implications regarding our ability to predict the nature of replacement strains after the introduction of therapies that target subsets of strains within a bacterial population using a microbiome-sparing approach.
7 Clinical Applications
Pangenomic analyses can be utilized to identify potential therapeutic targets. Target specificity can be customized depending on the desired effect. The core genome can be used to target an entire species, as it contains genes possessed by every member of the species. Alternatively, targeting select members of the accessory genome, or the “microbiome-sparing” approach, will ensure that only strains containing the gene of interest are affected. Both strategies can be utilized to combat a wide variety of pathogens.
Current efforts to combat pathogenic bacteria include targeting the bacterial capsule, a large polysaccharide layer that is a major virulence determinant with a key role in immune evasion. Strains vary in the composition of their capsules: those with identical capsules are placed in the same serotype, and those with highly similar capsules within a serogroup. For example, there are over 97 different serotypes known for S. pneumoniae that fall into 46 serogroups (Bentley et al. 2006; Geno et al. 2015; Tzeng et al. 2016), and over 12 serotypes for N. meningitidis (Harrison et al. 2013; Geno et al. 2015; Tzeng et al. 2016; Claus et al. 1997). New serotypes can arise by HGT, like in the movement of SiaD genes between N. meningitidis strains, or through mispairing during gene replication, which is responsible for serotypes 15 B/C in S. pneumoniae (Claus et al. 1997; van Selm et al. 2003). Capsular polysaccharide vaccines are available for S. pneumoniae, S. typhi, and N. meningitidis (Geno et al. 2015; Tzeng et al. 2016; Hessel et al. 1999). These specifically target the bacterial capsule, but young children (under the age of two) fail to create antibodies against these vaccines. To combat this, polysaccharide–protein conjugate vaccines were designed, which combine the polysaccharide antigen with protein carriers and render them more immunogenic in young children (Finn 2004; Nair 2012; Szu et al. 1989; Lin et al. 2001). Development of conjugate vaccines faces major challenges, such as cost, host immune response, and bacterial structures (Nair 2012). Therefore, it would be ideal to create capsular polysaccharide vaccines with better immunogenicity. However, the structures of some capsule sugars are too similar to those found in mammalian tissues to be useful as polysaccharide vaccines. In these cases, vaccines could be designed to target virulence via accessory genes or to target these species as a whole via the core genome (Pichichero 2017; Daniels et al. 2016; Chan et al. 2018).
Using the accessory genome to create strain-specific drugs and vaccines has wide implications. For example, it is easy to imagine the creation of therapies against bacterial pathogens that are able to spare the larger microbiome. Commensal bacteria in the microbiome and pathogenic bacteria of the same species may share the same core genome, but can have vast differences in the content of their accessory genomes. If a therapy targets protein products from genes found only in the accessory genomes of pathogenic bacteria, it will not disturb the patient’s microflora as the commensal bacteria would lack the proteins the therapy is created against. This strategy has the potential to greatly improve patient health and recovery following a bacterial infection.
Pangenomic studies can aid in the development of diagnostic tools. As with vaccines and drug development, accessory genes can be used to identify a particular strain/phenotype and core genes to identify a specific species. A study of 17 clinical isolates of G. vaginalis was used to propose the reclassification of G. vaginalis as a genus, based on the extent of pangenomic variation (Ahmed et al. 2012). Previously, metronidazole was used as a blanket antibiotic for the treatment of bacterial vaginosis. However, the understanding that metronidazole-resistant clades of G. vaginalis are actually different species creates room for the development of diagnostic tools to inform antibiotic treatment for patients with bacterial vaginosis (Balashov et al. 2014). Similarly, pangenomic studies among phenotypically divergent M. catarrhalis strains led to the characterization of a deep phylogenetic clade structure that separated the pathogenic sero-resistant strains from commensal sero-sensitive strains (Earl et al. 2016). In yet another example, Staphylococcus epidermidis was divided into two phylogenetic groups. One group included both commensals and pathogens, the other composed exclusively of commensal strains. Strains in the second group-encoded formate dehydrogenase, revealing a potential diagnostic marker (Conlan et al. 2012). A study in Helicobacter pylori identified lineage-specific genes; some have already been associated with acid resistance and virulence, and thus are potential targets to guide treatments (van Vliet 2017). Moreover, when studies associating pangenome and phenotype identify unannotated genes as diagnostic markers, they provide genetic fodder for linking new functions, distribution, and disease outcome (Ehrlich et al. 2010). One caution to consider in the development of diagnostics is that chronic infections can be caused by multiple strains of the same species, and analysis of a single strain could misdirect treatment.
A crucial benefit of pangenomic analyses is their ability to determine the presence or absence of antibiotic-resistant markers. Prescription of an ineffective antibiotic is both detrimental to patient’s health and adds to the problem of global antibiotic resistance. Some examples of pangenomic analyses to study the distribution and transmission of resistance genes have been performed on E. coli strains collected from wastewater treatment plants (Mahfouz et al. 2018), community-associated Clostridium difficile strains isolated from farm animals and humans (Knetsch et al. 2018), and strains of Stenotrophomonas maltophilia collected from cystic fibrosis (CF) patients (Esposito et al. 2017). Given that related strains often differ in their drug resistance profile, probing the accessory genome for genes that encode drug resistance will be a critical component of personalized medicine.
Genome-scale models (GEMs) of metabolism can provide great insight into the link between metabolism and pathogenesis. These network reconstructions provide context for the relationship between gene, gene product, and phenotype. Pangenomic analyses in three species observed that the majority of core genes are associated with metabolism (Cornejo et al. 2013; Bosi et al. 2016; Vieira et al. 2011). Pangenomic analysis of inflammatory bowel disease (IBD)-associated E. coli strains reported metabolic differences between IBD-associated strains and nonassociated strains, where the former set appeared to utilize energy more efficiently (Fang et al. 2018). The differences in metabolic capabilities in disease and healthy states provide a promising place to explore diagnostic applications of the pangenome. Furthermore, the link between metabolism and virulence can be explored, and be used diagnostically to differentiate strains that cause mild or severe symptom presentation (Bosi et al. 2016).
Beyond the use of pangenomic analyses to select targets for vaccines, therapeutics, and diagnosis, it has also served as an epidemiological tool. The origin of the 2010 cholera outbreak in Haiti was traced using pangenomic analysis of Vibrio cholerae. Initially, it was unclear whether the epidemic originated with a local strain or Asian strain. A pangenomic analysis revealed that the epidemic was caused by strains originated in Southeast Asia (Reimer et al. 2011; Hendriksen et al. 2011; Chin et al. 2011; Mutreja et al. 2011; Orata et al. 2014; Hasan et al. 2012). Such epidemiological studies allow better strategic planning to avoid future epidemics.
8 Conclusions
The Distributed Genome Hypothesis provides both a historical and theoretical framework for understanding bacterial genomic plasticity, and puts it in the context of other classes of chronic pathogens (viruses and eukaryotic parasites) that have developed different mechanistic strategies for the generation of genetic diversity in situ. Viruses such as HIV-1 utilize an error-prone DNA polymerase (reverse transcriptase) to generate enormous diversity resulting in the development of a quasispecies within days of infection (Korber et al. 2001). Trypanosomes utilize a cassetting mechanism for antigen switching wherein they have an entire chromosome of outer surface protein cassettes that they can exchange within the larger functional protein whenever the host adaptive immune response recognizes the previous cassette (Horn 2014). Thus, within this context, we can view HGT of distributed genes among bacterial strains of a species as yet another means of “programmed” variation (Ehrlich et al. 2010).
9 Perspectives
The plasticity provided by the eubacterial pangenome may be driving the evolution of other domains of life. The rapid recombination of bacterial strains provided the evolutionary pressure for the development of the vertebrate adaptive immune system—which is mechanistically similar to what the bacteria are doing—it is essentially a random gene rearrangement phenomenon, very similar to HGT (Hu et al. 2007). Lastly, as the variability in species becomes apparent, it triggers the question of how best to define a species. While pangenomic analyses do not offer the ultimate solution, they may provide a useful definition. Once the core genome of a species is defined, strains can be assigned, or not assigned, to a species based on the extent to which they share the same core genome (Nistico et al. 2014).
References
Adli M (2018) The CRISPR tool kit for genome editing and beyond. Nat Commun 9:1911
Agodi A, Zarrilli R, Barchitta M, Anzaldi A, Di Popolo A, Mattaliano A, Ghiraldi E, Travali S (2006) Alert surveillance of intensive care unit-acquired Acinetobacter infections in a Sicilian hospital. Clin Microbiol Infect 12(3):241–247
Ahmed A, Earl J, Retchless A, Hillier SL, Rabe LK, Cherpes TL et al (2012) Comparative genomic analyses of 17 clinical isolates of Gardnerella vaginalis provide evidence of multiple genetically isolated clades consistent with subspeciation into genovars. J Bacteriol 194:3922–3937
Antic I, Brothers KM, Stolzer M, Lai H, Powell E, Eutsey R et al (2017) Gene acquisition by a distinct phyletic group within Streptococcus pneumoniae promotes adhesion to the ocular epithelium. mSphere 2:e00213. https://doi.org/10.1128/mSphere.00213-17
Askar M, Faber MS, Frank C, Bernard H, Gilsdorf A, Fruth A, et al (2011) Update on the ongoing outbreak of haemolytic uraemic syndrome due to Shiga toxin-producing Escherichia coli (STEC) serotype O104, Germany, May 2011. Euro Surveill 16. Available https://www.ncbi.nlm.nih.gov/pubmed/21663710
Azarian T, Grant LR, Arnold BJ, Hammitt LL, Reid R, Santosham M et al (2018) The impact of serotype-specific vaccination on phylodynamic parameters of Streptococcus pneumoniae and the pneumococcal pan-genome. PLoS Pathog 14:e1006966
Aziz RK, Bartels D, Best AA, DeJongh M, Disz T, Edwards RA et al (2008) The RAST server: rapid annotations using subsystems technology. BMC Genomics 9:75
Balashov SV, Mordechai E, Adelson ME, Gygax SE (2014) Identification, quantification and subtyping of Gardnerella vaginalis in noncultured clinical vaginal samples by quantitative PCR. J Med Microbiol 63:162–175
Bentley SD, Aanensen DM, Mavroidi A, Saunders D, Rabbinowitsch E, Collins M et al (2006) Genetic analysis of the capsular biosynthetic locus from all 90 pneumococcal serotypes. PLoS Genet 2:e31
Bianconi I, D’Arcangelo S, Esposito A, Benedet M, Piffer E, Dinnella G et al (2018) Persistence and microevolution of Pseudomonas aeruginosa in the cystic fibrosis lung: a single-patient longitudinal genomic study. Front Microbiol 9:3242
Bielaszewska M, Prager R, Köck R, Mellmann A, Zhang W, Tschäpe H et al (2007) Shiga toxin gene loss and transfer in vitro and in vivo during enterohemorrhagic Escherichia coli O26 infection in humans. Appl Environ Microbiol 73:3144–3150
Bonar EA, Bukowski M, Hydzik M, Jankowska U, Kedracka-Krok S, Groborz M et al (2018) Joint genomic and proteomic analysis identifies meta-trait characteristics of virulent and non-virulent strains. Front Cell Infect Microbiol 8:313
Bondy-Denomy J, Qian J, Westra ER, Buckling A, Guttman DS, Davidson AR et al (2016) Prophages mediate defense against phage infection through diverse mechanisms. ISME J 10:2854–2866
Bosi E, Monk JM, Aziz RK, Fondi M, Nizet V, Palsson BØ (2016) Comparative genome-scale modelling of Staphylococcus aureus strains identifies strain-specific metabolic capabilities linked to pathogenicity. Proc Natl Acad Sci U S A 113:E3801–E3809
Brown NF, Finlay BB (2011) Potential origins and horizontal transfer of type III secretion systems and effectors. Mob Genet Elem 1:118–121
Brynildsrud OB, Eldholm V, Bohlin J, Uadiale K, Obaro S, Caugant DA (2018) Acquisition of Virulence genes by a carrier strain gave rise to the ongoing epidemics of meningococcal disease in West Africa. Proc Natl Acad Sci U S A 115(21):5510–5515
Budroni S, Siena E, Dunning Hotopp JC, Seib KL, Serruto D, Nofroni C et al (2011) Neisseria meningitidis is structured in clades associated with restriction modification systems that modulate homologous recombination. Proc Natl Acad Sci U S A 108:4494–4499
Burns JL, Gibson RL, McNamara S, Yim D, Emerson J, Rosenfeld M et al (2001) Longitudinal assessment of Pseudomonas aeruginosa in young children with cystic fibrosis. J Infect Dis 183:444–452
Chan W-Y, Entwisle C, Ercoli G, Ramos-Sevillano E, McIlgorm A, Cecchini P et al (2018) A novel, multiple-antigen pneumococcal vaccine protects against lethal Streptococcus pneumoniae challenge. Infect Immun 87(3):e00846. https://doi.org/10.1128/IAI.00846-18
Chewapreecha C, Harris SR, Croucher NJ, Turner C, Marttinen P, Cheng L et al (2014) Dense genomic sampling identifies highways of pneumococcal recombination. Nat Genet 46:305–309
Chin C-S, Sorenson J, Harris JB, Robins WP, Charles RC, Jean-Charles RR et al (2011) The origin of the Haitian cholera outbreak strain. N Engl J Med 364:33–42
Chopin M-C, Chopin A, Bidnenko E (2005) Phage abortive infection in Lactococci: variations on a theme. Curr Opin Microbiol 8:473–479
Claus H, Vogel U, Mühlenhoff M, Gerardy-Schahn R, Frosch M (1997) Molecular divergence of the sia locus in different serogroups of Neisseria meningitidis expressing polysialic acid capsules. Mol Gen Genet 257:28–34
Conlan S, Mijares LA, NISC Comparative Sequencing Program, Becker J, Blakesley RW, Bouffard GG et al (2012) Staphylococcus epidermidis pan-genome sequence analysis reveals diversity of skin commensal and hospital infection-associated isolates. Genome Biol 13:R64
Conlan S, Thomas PJ, Deming C, Park M, Lau AF, Dekker JP et al (2014) Single-molecule sequencing to track plasmid diversity of hospital-associated carbapenemase-producing Enterobacteriaceae. Sci Transl Med 6:254ra126
Corander J, Fraser C, Gutmann MU, Arnold B, Hanage WP, Bentley SD et al (2017) Frequency-dependent selection in vaccine-associated pneumococcal population dynamics. Nat Ecol Evol 1:1950–1960
Cornejo OE, Lefébure T, Bitar PDP, Lang P, Richards VP, Eilertson K et al (2013) Evolutionary and population genomics of the cavity causing bacteria Streptococcus mutans. Mol Biol Evol 30:881–893
Croucher NJ, Chewapreecha C, Hanage WP, Harris SR, McGee L, van der Linden M et al (2014a) Evidence for soft selective sweeps in the evolution of pneumococcal multidrug resistance and vaccine escape. Genome Biol Evol 6:1589–1602
Croucher NJ, Coupland PG, Stevenson AE, Callendrello A, Bentley SD, Hanage WP (2014b) Diversification of bacterial genome content through distinct mechanisms over different timescales. Nat Commun 5:5471
Daniels CC, Rogers PD, Shelton CM (2016) A review of pneumococcal vaccines: current polysaccharide vaccine recommendations and future protein antigens. J Pediatr Pharmacol Ther 21:27–35
Darling ACE, Mau B, Blattner FR, Perna NT (2004) Mauve: multiple alignment of conserved genomic sequence with rearrangements. Genome Res 14:1394–1403
Datta S, Mitra S, Chattopadhyay P, Som T, Mukherjee S, Basu S (2017) Spread and exchange of bla NDM-1 in hospitalized neonates: role of mobilizable genetic elements. Eur J Clin Microbiol Infect Dis 36:255–265
Davie JJ, Earl J, de Vries SPW, Ahmed A, Hu FZ, Bootsma HJ et al (2011) Comparative analysis and supragenome modeling of twelve Moraxella catarrhalis clinical isolates. BMC Genomics 12:70
Dawid S, Roche AM, Weiser JN (2007) The blp bacteriocins of Streptococcus pneumoniae mediate intraspecies competition both in vitro and in vivo. Infect Immun 75:443–451
Dedrick RM, Jacobs-Sera D, Bustamante CAG, Garlena RA, Mavrich TN, Pope WH et al (2017) Prophage-mediated defence against viral attack and viral counter-defence. Nat Microbiol 2:16251
Earl JP, de Vries SPW, Ahmed A, Powell E, Schultz MP, Hermans PWM et al (2016) Comparative genomic analyses of the Moraxella catarrhalis serosensitive and seroresistant lineages demonstrate their independent evolution. Genome Biol Evol 8:955–974
Edgar RC (2004) MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res 32:1792–1797
Ehrlich GD, Hu FZ, Shen K, Stoodley P, Post JC (2005) Bacterial plurality as a general mechanism driving persistence in chronic infections. Clin Orthop Relat Res 437:20–24
Ehrlich GD, Ahmed A, Earl J, Hiller NL, Costerton JW, Stoodley P et al (2010) The distributed genome hypothesis as a rubric for understanding evolution in situ during chronic bacterial biofilm infectious processes. FEMS Immunol Med Microbiol 59:269–279
Esposito A, Pompilio A, Bettua C, Crocetta V, Giacobazzi E, Fiscarelli E et al (2017) Evolution of Stenotrophomonas maltophilia in cystic fibrosis lung over chronic infection: a genomic and phenotypic population study. Front Microbiol 8:1590
Eutsey RA, Powell E, Dordel J, Salter SJ, Clark TA, Korlach J et al (2015) Genetic stabilization of the drug-resistant PMEN1 Pneumococcus lineage by its distinctive DpnIII restriction-modification system. MBio 6:e00173
Fang X, Monk JM, Mih N, Du B, Sastry AV, Kavvas E et al (2018) Escherichia coli B2 strains prevalent in inflammatory bowel disease patients have distinct metabolic capabilities that enable colonization of intestinal mucosa. BMC Syst Biol 12:66
Feld L, Schjørring S, Hammer K, Licht TR, Danielsen M, Krogfelt K et al (2008) Selective pressure affects transfer and establishment of a Lactobacillus plantarum resistance plasmid in the gastrointestinal environment. J Antimicrob Chemother 61:845–852
Finn A (2004) Bacterial polysaccharide-protein conjugate vaccines. Br Med Bull 70:1–14
Geno KA, Gilbert GL, Song JY, Skovsted IC, Klugman KP, Jones C et al (2015) Pneumococcal capsules and their types: past, present, and future. Clin Microbiol Rev 28:871–899
Georgiades K, Raoult D (2011) Genomes of the most dangerous epidemic bacteria have a virulence repertoire characterized by fewer genes but more toxin-antitoxin modules. PLoS One 6:e17962
Georgiades K, Merhej V, El Karkouri K, Raoult D, Pontarotti P (2011) Gene gain and loss events in Rickettsia and Orientia species. Biol Direct 6:6
Gerlach D, Guo Y, De Castro C, Kim S-H, Schlatterer K, Xu F-F et al (2018) Methicillin-resistant Staphylococcus aureus alters cell wall glycosylation to evade immunity. Nature 563:705–709
Gladitz J, Shen K, Antalis P, Hu FZ, Post JC, Ehrlich GD (2005) Codon usage comparison of novel genes in clinical isolates of Haemophilus influenzae. Nucleic Acids Res 33:3644–3658
Golubchik T, Brueggemann AB, Street T, Gertz RE, Spencer CCA, Ho T et al (2012) Pneumococcal genome sequencing tracks a vaccine escape variant formed through a multi-fragment recombination event. Nat Genet 44:352–355
Gona F, Barbera F, Pasquariello AC, Grossi P, Gridelli B, Mezzatesta ML et al (2014) In vivo multiclonal transfer of bla(KPC-3) from Klebsiella pneumoniae to Escherichia coli in surgery patients. Clin Microbiol Infect 20:O633–O635
Gophna U, Kristensen DM, Wolf YI, Popa O, Drevet C, Koonin EV (2015) No evidence of inhibition of horizontal gene transfer by CRISPR-Cas on evolutionary timescales. ISME J 9:2021–2027
Göttig S, Gruber TM, Stecher B, Wichelhaus TA, Kempf VAJ (2015) In vivo horizontal gene transfer of the carbapenemase OXA-48 during a nosocomial outbreak. Clin Infect Dis 60:1808–1815
Griffith F (1928) The significance of Pneumococcal types. J Hyg 27:113–159
Guindon S, Dufayard J-F, Lefort V, Anisimova M, Hordijk W, Gascuel O (2010) New algorithms and methods to estimate maximum-likelihood phylogenies: assessing the performance of PhyML 3.0. Syst Biol 59:307–321
Gumpert H, Kubicek-Sutherland JZ, Porse A, Karami N, Munck C, Linkevicius M, Adlerberth I, Wold AE, Andersson DI, Sommer MOA (2017) Transfer and persistence of a multi-drug resistance plasmid in situ of the infant gut microbiota in the absence of antibiotic treatment. Front Microbiol 8(September):1852
Hacker J, Blum-Oehler G, Mühldorfer I, Tschäpe H (1997) Pathogenicity islands of virulent bacteria: structure, function and impact on microbial evolution. Mol Microbiol 23:1089–1097
Hall BG, Ehrlich GD, Hu FZ (2010) Pan-genome analysis provides much higher strain typing resolution than multi-locus sequence typing. Microbiology 156:1060–1068
Hammerum AM, Hansen F, Nielsen HL, Jakobsen L, Stegger M, Andersen PS, Jensen P et al (2016) Use of WGS data for investigation of a long-term NDM-1-producing Citrobacter freundii outbreak and secondary in vivo spread of blaNDM-1 to Escherichia coli, Klebsiella pneumoniae and Klebsiella oxytoca. J Antimicrob Chemother 71(11):3117–3124
Harris HMB, Bourin MJB, Claesson MJ, O’Toole PW (2017) Phylogenomics and comparative genomics of, a mammalian gut commensal. Microb Genom 3:e000115
Harrison OB, Claus H, Jiang Y, Bennett JS, Bratcher HB, Jolley KA et al (2013) Description and nomenclature of Neisseria meningitidis capsule locus. Emerg Infect Dis 19:566–573
Hasan NA, Choi SY, Eppinger M, Clark PW, Chen A, Alam M et al (2012) Genomic diversity of 2010 Haitian cholera outbreak strains. Proc Natl Acad Sci U S A 109:E2010
Hazen TH, Sahl JW, Fraser CM, Donnenberg MS, Scheutz F, Rasko DA (2013) Refining the pathovar paradigm via phylogenomics of the attaching and effacing Escherichia coli. Proc Natl Acad Sci U S A 110:12810–12815
Hendrickson HL, Barbeau D, Ceschin R, Lawrence JG (2018) Chromosome architecture constrains horizontal gene transfer in bacteria. PLoS Genet 14:e1007421
Hendriksen RS, Price LB, Schupp JM, Gillece JD, Kaas RS, Engelthaler DM et al (2011) Population genetics of Vibrio cholerae from Nepal in 2010: evidence on the origin of the Haitian outbreak. MBio 2:e00157
Hessel L, Debois H, Fletcher M, Dumas R (1999) Experience with Salmonella typhi Vi capsular polysaccharide vaccine. Eur J Clin Microbiol Infect Dis 18:609–620
Higgins DG, Sharp PM (1988) CLUSTAL: a package for performing multiple sequence alignment on a microcomputer. Gene 73:237–244
Hiller NL, Ahmed A, Powell E, Martin DP, Eutsey R, Earl J et al (2010) Generation of genic diversity among Streptococcus pneumoniae strains via horizontal gene transfer during a chronic polyclonal pediatric infection. PLoS Pathog 6:e1001108
Hiller NL, Eutsey RA, Powell E, Earl JP, Janto B, Martin DP et al (2011) Differences in genotype and virulence among four multidrug-resistant Streptococcus pneumoniae isolates belonging to the PMEN1 clone. PLoS One 6:e28850
Hogg JS, Hu FZ, Janto B, Boissy R, Hayes J, Keefe R et al (2007) Characterization and modeling of the Haemophilus influenzae core and supragenomes based on the complete genomic sequences of Rd and 12 clinical nontypeable strains. Genome Biol 8:R103
Horn D (2014) Antigenic variation in African trypanosomes. Mol Biochem Parasitol 195:123–129
Hu FZ, Hogg J, Hiller NL, Janto B, Boissy R, Post JC, Ehrlich GD (2007) Biofilms as bacterial breeding grounds: a counterpoint to the adaptive host response. Abstract MS15 9th international symposium on recent advances in otitis media, June 3–7, 2007, in St. Pete Beach, FL
Hueck CJ (1998) Type III protein secretion systems in bacterial pathogens of animals and plants. Microbiol Mol Biol Rev 62:379–433
Hurdle JG, O’Neill AJ, Mody L, Chopra I, Bradley SF (2005) In vivo transfer of high-level mupirocin resistance from Staphylococcus epidermidis to methicillin-resistant Staphylococcus aureus associated with failure of mupirocin prophylaxis. J Antimicrob Chemother 56:1166–1168
Hyatt D, Chen G-L, Locascio PF, Land ML, Larimer FW, Hauser LJ (2010) Prodigal: prokaryotic gene recognition and translation initiation site identification. BMC Bioinf 11:119
Jennings E, Thurston TLM, Holden DW (2017) Salmonella SPI-2 type III secretion system effectors: molecular mechanisms and physiological consequences. Cell Host Microbe 22:217–231
Jiang F, Doudna JA (2017) CRISPR-Cas9 structures and mechanisms. Annu Rev Biophys 46:505–529
Jiang W, Maniv I, Arain F, Wang Y, Levin BR, Marraffini LA (2013) Dealing with the evolutionary downside of CRISPR immunity: bacteria and beneficial plasmids. PLoS Genet 9:e1003844
Johnston C, Polard P, Claverys J-P (2013a) The DpnI/DpnII pneumococcal system, defense against foreign attack without compromising genetic exchange. Mob Genet Elem 3:e25582
Johnston C, Martin B, Polard P, Claverys J-P (2013b) Postreplication targeting of transformants by bacterial immune systems? Trends Microbiol 21:516–521
Jorth P, Staudinger BJ, Wu X, Hisert KB, Hayden H, Garudathri J et al (2015) Regional isolation drives bacterial diversification within cystic fibrosis lungs. Cell Host Microbe 18:307–319
Kaper JB, Nataro JP, Mobley HL (2004) Pathogenic Escherichia coli. Nat Rev Microbiol 2:123–140
Knetsch CW, Kumar N, Forster SC, Connor TR, Browne HP, Harmanus C et al (2018) Zoonotic transfer of Clostridium difficile harboring antimicrobial resistance between farm animals and humans. J Clin Microbiol 56(3):e01384. https://doi.org/10.1128/JCM.01384-17
Korber B, Gaschen B, Yusim K, Thakallapally R, Kesmir C, Detours V (2001) Evolutionary and immunological implications of contemporary HIV-1 variation. Br Med Bull 58:19–42
Kress-Bennett JM, Hiller NL, Eutsey RA, Powell E, Longwell MJ, Hillman T et al (2016) Identification and characterization of msf, a novel virulence factor in Haemophilus influenzae. PLoS One 11:e0149891
Kumar S, Tamura K, Nei M (1994) MEGA: molecular evolutionary genetics analysis software for microcomputers. Comput Appl Biosci 10:189–191
Kwun MJ, Oggioni MR, De Ste Croix M, Bentley SD, Croucher NJ (2018) Excision-reintegration at a pneumococcal phase-variable restriction-modification locus drives within- and between-strain epigenetic differentiation and inhibits gene acquisition. Nucleic Acids Res 46:11438–11453
Langhanki L, Berger P, Treffon J, Catania F, Kahl BC, Mellmann A (2018) In vivo competition and horizontal gene transfer among distinct Staphylococcus aureus lineages as major drivers for adaptational changes during long-term persistence in humans. BMC Microbiol 18:152
Lee AH-Y, Flibotte S, Sinha S, Paiero A, Ehrlich RL, Balashov S et al (2017) Phenotypic diversity and genotypic flexibility of Burkholderia cenocepacia during long-term chronic infection of cystic fibrosis lungs. Genome Res 27:650–662
Li Z, Kosorok MR, Farrell PM, Laxova A, West SEH, Green CG et al (2005) Longitudinal development of mucoid Pseudomonas aeruginosa infection and lung disease progression in children with cystic fibrosis. JAMA 293:581–588
Lin FY, Ho VA, Khiem HB, Trach DD, Bay PV, Thanh TC et al (2001) The efficacy of a Salmonella typhi Vi conjugate vaccine in two-to-five-year-old children. N Engl J Med 344:1263–1269
Lux T, Nuhn M, Hakenbeck R, Reichmann P (2007) Diversity of bacteriocins and activity spectrum in Streptococcus pneumoniae. J Bacteriol 189:7741–7751
Mahan MJ, Kubicek-Sutherland JZ, Heithoff DM (2013) Rise of the microbes. Virulence 4:213–222
Mahfouz N, Caucci S, Achatz E, Semmler T, Guenther S, Berendonk TU et al (2018) High genomic diversity of multi-drug resistant wastewater Escherichia coli. Sci Rep 8:8928
Mata C, Miró E, Mirelis B, Garcillán-Barcia MP, de la Cruz F, Coll P et al (2010) In vivo transmission of a plasmid coharbouring bla and qnrB genes between Escherichia coli and Serratia marcescens. FEMS Microbiol Lett 308:24–28
McAdam PR, Vander Broek CW, Lindsay DSJ, Ward MJ, Hanson MF, Gillies M, Watson M, Stevens JM, Edwards GF, Fitzgerald JR (2014) Gene flow in environmental Legionella pneumophila leads to genetic and pathogenic heterogeneity within a Legionnaires’ disease outbreak. Genome Biol 15(11):504
Mell JC, Shumilina S, Hall IM, Redfield RJ (2011) Transformation of natural genetic variation into Haemophilus influenzae genomes. PLoS Pathog 7:e1002151
Mellmann A, Harmsen D, Cummings CA, Zentz EB, Leopold SR, Rico A et al (2011) Prospective genomic characterization of the German enterohemorrhagic Escherichia coli O104:H4 outbreak by rapid next generation sequencing technology. PLoS One 6:e22751
Mena A, Plasencia V, García L, Hidalgo O, Ayestarán JI, Alberti S et al (2006) Characterization of a large outbreak by CTX-M-1-producing Klebsiella pneumoniae and mechanisms leading to in vivo carbapenem resistance development. J Clin Microbiol 44:2831–2837
Merhej V, Royer-Carenzi M, Pontarotti P, Raoult D (2009) Massive comparative genomic analysis reveals convergent evolution of specialized bacteria. Biol Direct 4:13
Moleres J, Fernández-Calvet A, Ehrlich RL, Martí S, Pérez-Regidor L, Euba B et al (2018) Antagonistic pleiotropy in the bifunctional surface protein FadL (OmpP1) during adaptation of Haemophilus influenzae to chronic lung infection associated with chronic obstructive pulmonary disease. MBio 9:e01176. https://doi.org/10.1128/mBio.01176-18
Moore PC, Lindsay JA (2001) Genetic variation among hospital isolates of methicillin-sensitive Staphylococcus aureus: evidence for horizontal transfer of Virulence genes. J Clin Microbiol 39(8):2760–2767
Mutreja A, Kim DW, Thomson NR, Connor TR, Lee JH, Kariuki S et al (2011) Evidence for several waves of global transmission in the seventh cholera pandemic. Nature 477:462–465
Nair M (2012) Protein conjugate polysaccharide vaccines: challenges in development and global implementation. Indian J Community Med 37:79–82
Neuwirth C, Siebor E, Pechinot A, Duez JM, Pruneaux M, Garel F et al (2001) Evidence of in vivo transfer of a plasmid encoding the extended-spectrum beta-lactamase TEM-24 and other resistance factors among different members of the family Enterobacteriaceae. J Clin Microbiol 39:1985–1988
Nistico L, Earl J, Hiller L, Ahmed A, Retchless A, Janto B, Costerton JC, Hu FZ, Ehrlich GD (2014) Using the core and supra genomes to determine diversity and natural proclivities among bacterial strains. In: Skovhus TL, Caffrey S, Hubert C (eds) Application of molecular microbiological methods. Caister Academic Press, Norfolk, UK, p 1
Ochman H, Soncini FC, Solomon F, Groisman EA (1996) Identification of a pathogenicity island required for Salmonella survival in host cells. Proc Natl Acad Sci 93:7800–7804
Okinaka RT, Keim P (2016) The phylogeny of Bacillus cereus sensu lato. Microbiol Spectr 4(1). https://doi.org/10.1128/microbiolspec.TBS-0012-2012
Orata FD, Keim PS, Boucher Y (2014) The 2010 cholera outbreak in Haiti: how science solved a controversy. PLoS Pathog 10:e1003967
Palmer KL, Gilmore MS (2010) Multidrug-resistant enterococci lack CRISPR-cas. MBio 1:e00227. https://doi.org/10.1128/mBio.00227-10
Pensar J, Puranen S, MacAlasdair N, Kuronen J, Tonkin-Hill G, Pesonen M et al (2019) Genome-wide epistasis and co-selection study using mutual information. Genomics. bioRxiv
Pettigrew MM, Ahearn CP, Gent JF, Kong Y, Gallo MC, Munro JB et al (2018) Haemophilus influenzae genome evolution during persistence in the human airways in chronic obstructive pulmonary disease. Proc Natl Acad Sci U S A 115:E3256–E3265
Pichichero ME (2017) Pneumococcal whole-cell and protein-based vaccines: changing the paradigm. Expert Rev Vaccines 16:1181–1190
Rasko DA, Rosovitz MJ, Myers GSA, Mongodin EF, Fricke WF, Gajer P et al (2008) The pangenome structure of Escherichia coli: comparative genomic analysis of E. coli commensal and pathogenic isolates. J Bacteriol 190:6881–6893
Rasko DA, Webster DR, Sahl JW, Bashir A, Boisen N, Scheutz F et al (2011) Origins of the E. coli strain causing an outbreak of hemolytic-uremic syndrome in Germany. N Engl J Med 365:709–717
Rau MH, Marvig RL, Ehrlich GD, Molin S, Jelsbak L (2012) Deletion and acquisition of genomic content during early stage adaptation of Pseudomonas aeruginosa to a human host environment. Environ Microbiol 14:2200–2211
Reimer AR, Van Domselaar G, Stroika S, Walker M, Kent H, Tarr C et al (2011) Comparative genomics of Vibrio cholerae from Haiti, Asia, and Africa. Emerg Infect Dis 17:2113–2121
Rezaei Javan R, van Tonder AJ, King JP, Harrold CL, Brueggemann AB (2018) Genome sequencing reveals a large and diverse repertoire of antimicrobial peptides. Front Microbiol 9:2012
Salipante SJ, Roach DJ, Kitzman JO, Snyder MW, Stackhouse B, Butler-Wu SM et al (2015) Large-scale genomic sequencing of extraintestinal pathogenic Escherichia coli strains. Genome Res 25:119–128
Schjørring S, Struve C, Krogfelt KA (2008) Transfer of antimicrobial resistance plasmids from Klebsiella pneumoniae to Escherichia coli in the mouse intestine. J Antimicrob Chemother 62(5):1086–1093
Seemann T (2014) Prokka: rapid prokaryotic genome annotation. Bioinformatics 30:2068–2069
Shen K, Antalis P, Gladitz J, Sayeed S, Ahmed A, Yu S et al (2005) Identification, distribution, and expression of novel genes in 10 clinical isolates of nontypeable Haemophilus influenzae. Infect Immun 73:3479–3491
Sidjabat HE, Heney C, George NM, Nimmo GR, Paterson DL (2014) Interspecies transfer of blaIMP-4 in a patient with prolonged colonization by IMP-4-producing Enterobacteriaceae. J Clin Microbiol 52:3816–3818
Silva IN, Santos PM, Santos MR, Zlosnik JEA, Speert DP, Buskirk SW et al (2016) Long-term evolution of Burkholderia multivorans during a chronic cystic fibrosis infection reveals shifting forces of selection. mSystems 1:e00029. https://doi.org/10.1128/mSystems.00029-16
Soto SM, Zúñiga S, Ulleryd P, Vila J (2011) Acquisition of a pathogenicity island in an Escherichia coli clinical isolate causing febrile urinary tract infection. Eur J Clin Microbiol Infect Dis 30:1543–1550
Stamatakis A (2006) RAxML-VI-HPC: maximum likelihood-based phylogenetic analyses with thousands of taxa and mixed models. Bioinformatics 22:2688–2690
Stanczak-Mrozek KI, Manne A, Knight GM, Gould K, Witney AA, Lindsay JA (2015) Within-host diversity of MRSA antimicrobial resistances. J Antimicrob Chemother 70(8):2191–2198
Swartley JS, Marfin AA, Edupuganti S, Liu LJ, Cieslak P, Perkins B et al (1997) Capsule switching of Neisseria meningitidis. Proc Natl Acad Sci U S A 94:271–276
Szu SC, Li XR, Schneerson R, Vickers JH, Bryla D, Robbins JB (1989) Comparative immunogenicities of Vi polysaccharide-protein conjugates composed of cholera toxin or its B subunit as a carrier bound to high- or lower-molecular-weight Vi. Infect Immun 57:3823–3827
Tettelin H, Masignani V, Cieslewicz MJ, Donati C, Medini D, Ward NL et al (2005) Genome analysis of multiple pathogenic isolates of Streptococcus agalactiae: implications for the microbial “pan-genome”. Proc Natl Acad Sci U S A 102:13950–13955
Tzeng Y-L, Thomas J, Stephens DS (2016) Regulation of capsule in Neisseria meningitidis. Crit Rev Microbiol 42:759–772
Valente C, Dawid S, Pinto FR, Hinds J, Simões AS, Gould KA et al (2016) The blp locus of Streptococcus pneumoniae plays a limited role in the selection of strains that can Cocolonize the human nasopharynx. Appl Environ Microbiol 82:5206–5215
van Selm S, van Cann LM, Kolkman MAB, van der Zeijst BAM, van Putten JPM (2003) Genetic basis for the structural difference between Streptococcus pneumoniae serotype 15B and 15C capsular polysaccharides. Infect Immun 71:6192–6198
van Vliet AHM (2017) Use of pan-genome analysis for the identification of lineage-specific genes of Helicobacter pylori. FEMS Microbiol Lett 364(2):fnw296. https://doi.org/10.1093/femsle/fnw296
Vieira G, Sabarly V, Bourguignon P-Y, Durot M, Le Fèvre F, Mornico D et al (2011) Core and panmetabolism in Escherichia coli. J Bacteriol 193:1461–1472
Vilas-Bôas GT, Peruca APS, Arantes OMN (2007) Biology and taxonomy of Bacillus cereus, Bacillus anthracis, and Bacillus thuringiensis. Can J Microbiol 53:673–687
Watkins ER, Penman BS, Lourenço J, Buckee CO, Maiden MCJ, Gupta S (2015) Vaccination drives changes in metabolic and virulence profiles of Streptococcus pneumoniae. PLoS Pathog 11:e1005034
Watson BNJ, Staals RHJ, Fineran PC (2018) CRISPR-Cas-mediated phage resistance enhances horizontal gene transfer by transduction. MBio 9:e02406. https://doi.org/10.1128/mBio.02406-17
Wilson GG, Murray NE (2003) Restriction and modification systems. Annual Reviews. Palo Alto, CA. https://doi.org/10.1146/annurev.ge.25.120191.003101
Wyres KL, Lambertsen LM, Croucher NJ, McGee L, von Gottberg A, Liñares J et al (2012) The multidrug-resistant PMEN1 pneumococcus is a paradigm for genetic success. Genome Biol 13:R103
Zhou Y, Liang Y, Lynch KH, Dennis JJ, Wishart DS (2011) PHAST: a fast phage search tool. Nucleic Acids Res 39:W347–W352
References for Fig. 1
Abreu VAC, Popin RV, Alvarenga DO, Schaker PDC, Hoff-Risseti C, Varani AM, Fiore MF (2018) Genomic and genotypic characterization of Cylindrospermopsis raciborskii: toward an intraspecific phylogenetic evaluation by comparative genomics. Front Microbiol 9(February):306
Alhashash F, Wang X, Paszkiewicz K, Diggle M, Zong Z, McNally A (2016) Increase in bacteraemia cases in the east midlands region of the UK due to MDR Escherichia coli ST73: high levels of genomic and plasmid diversity in causative isolates. J Antimicrob Chemother 71(2):339–343
Ali A, Soares SC, Santos AR, Guimarães LC, Barbosa E, Almeida SS, Abreu VAC et al (2012) Campylobacter fetus subspecies: comparative genomics and prediction of potential virulence targets. Gene 508(2):145–156
Ali A, Naz A, Soares SC, Bakhtiar M, Tiwari S, Hassan SS, Hanan F et al (2015) Pan-genome analysis of human gastric pathogen H. pylori: comparative genomics and pathogenomics approaches to identify regions associated with pathogenicity and prediction of potential core therapeutic targets. Biomed Res Int:139580
Anastasi E, MacArthur I, Scortti M, Alvarez S, Giguère S, Vázquez-Boland JA (2016) Pangenome and phylogenomic analysis of the pathogenic actinobacterium Rhodococcus equi. Genome Biol Evol 8(10):3140–3148
Andreani NA, Carraro L, Martino ME, Fondi M, Fasolato L, Miotto G, Magro M, Vianello F, Cardazzo B (2015) A genomic and transcriptomic approach to investigate the blue pigment phenotype in Pseudomonas fluorescens. Int J Food Microbiol 213(November):88–98
Arboleya S, Bottacini F, O’Connell-Motherway M, Ryan CA, Ross RP, van Sinderen D, Stanton C (2018) Gene-trait matching across the Bifidobacterium longum pan-genome reveals considerable diversity in carbohydrate catabolism among human infant strains. BMC Genomics 19(1):33
Argemi X, Matelska D, Ginalski K, Riegel P, Hansmann Y, Bloom J, Pestel-Caron M, Dahyot S, Lebeurre J, Prévost G (2018a) Comparative genomic analysis of Staphylococcus lugdunensis shows a closed pan-genome and multiple barriers to horizontal gene transfer. BMC Genomics 19(1):621
Argemi X, Nanoukon C, Affolabi D, Keller D, Hansmann Y, Riegel P, Baba-Moussa L, Prévost G (2018b) Comparative genomics and identification of an enterotoxin-bearing pathogenicity island, SEPI-1/SECI-1, in Staphylococcus epidermidis pathogenic strains. Toxins 10(3):93. https://doi.org/10.3390/toxins10030093
Asenjo F, Olmos A, Henríquez-Piskulich P, Polanco V, Aldea P, Ugalde JA, Trombert AN (2016) Genome sequencing and analysis of the first complete genome of Lactobacillus kunkeei strain MP2, an Apis mellifera gut isolate. PeerJ 4(April):e1950
Baddam R, Kumar N, Shaik S, Lankapalli AK, Ahmed N (2014) Genome dynamics and evolution of Salmonella typhi strains from the typhoid-endemic zones. Sci Rep 4(December):7457
Bakshi U, Sarkar M, Paul S, Dutta C (2016) Assessment of virulence potential of uncharacterized Enterococcus faecalis strains using pan genomic approach - identification of pathogen-specific and habitat-specific genes. Sci Rep 6(December):38648
Baltrus DA, Nishimura MT, Romanchuk A, Chang JH, Mukhtar MS, Cherkis K, Roach J, Grant SR, Jones CD, Dangl JL (2011) Dynamic evolution of pathogenicity revealed by sequencing and comparative genomics of 19 Pseudomonas syringae isolates. PLoS Pathog 7(7):e1002132
Bansal K, Midha S, Kumar S, Patil PB (2017) Ecological and evolutionary insights into Xanthomonas citri pathovar diversity. Appl Environ Microbiol 83(9):e02993. https://doi.org/10.1128/AEM.02993-16
Baraúna RA, Ramos RTJ, Veras AAO, Pinheiro KC, Benevides LJ, Viana MVC, Guimarães LC et al (2017) Assessing the genotypic differences between strains of Corynebacterium Pseudotuberculosis Biovar Equi through comparative genomics. PLoS One 12(1):e0170676
Bazinet AL (2017) Pan-genome and phylogeny of Bacillus cereus Sensu Lato. BMC Evol Biol 17(1):176
Benevides L, Burman S, Martin R, Robert V, Thomas M, Miquel S, Chain F et al (2017) New insights into the diversity of the genus Faecalibacterium. Front Microbiol 8(September):1790
Bergsveinson J, Ziola B (2017) Comparative genomic and plasmid analysis of beer-spoiling and non- beer-spoiling Lactobacillus brevis isolates. Can J Microbiol 63(12):970–983
Bhardwaj T, Somvanshi P (2017) Pan-genome analysis of Clostridium botulinum reveals unique targets for drug development. Gene 623(August):48–62
Bhattacharyya C, Bakshi U, Mallick I, Mukherji S, Bera B, Ghosh A (2017) Genome-guided insights into the plant growth promotion capabilities of the physiologically versatile Bacillus aryabhattai strain AB211. Front Microbiol 8(March):411
Boissy R, Ahmed A, Janto B, Earl J, Hall BG, Hogg JS, Pusch GD et al (2011) Comparative supragenomic analyses among the pathogens Staphylococcus aureus, Streptococcus pneumoniae, and Haemophilus influenzae using a modification of the finite supragenome model. BMC Genomics 12(April):187
Bolotin E, Hershberg R (2015) Gene loss dominates as a source of genetic variation within clonal pathogenic bacterial species. Genome Biol Evol 7(8):2173–2187
Borneman AR, McCarthy JM, Chambers PJ, Bartowsky EJ (2012) Comparative analysis of the Oenococcus oeni pan genome reveals genetic diversity in industrially-relevant pathways. BMC Genomics 13(August):373
Bottacini F, Motherway MO’C, Kuczynski J, O’Connell KJ, Serafini F, Duranti S, Milani C et al (2014) Comparative genomics of the Bifidobacterium breve taxon. BMC Genomics 15(March):170
Breurec S, Criscuolo A, Diancourt L, Rendueles O, Vandenbogaert M, Passet V, Caro V, Rocha EPC, Touchon M, Brisse S (2016) Genomic epidemiology and global diversity of the emerging bacterial pathogen Elizabethkingia anophelis. Sci Rep 6(July):30379
Brito PH, Chevreux B, Serra CR, Schyns G, Henriques AO, Pereira-Leal JB (2018) Genetic competence drives genome diversity in Bacillus subtilis. Genome Biol Evol 10(1):108–124
Broadbent JR, Neeno-Eckwall EC, Stahl B, Tandee K, Cai H, Morovic W, Horvath P et al (2012) Analysis of the Lactobacillus casei supragenome and its influence in species evolution and lifestyle adaptation. BMC Genomics 13(October):533
Bulagonda EP, Manivannan B, Mahalingam N, Lama M, Chanakya PP, Khamari B, Jadhao S, Vasudevan M, Nagaraja V (2018) Comparative genomic analysis of a naturally competent Elizabethkingia anophelis isolated from an eye infection. Sci Rep 8(1):8447
Cao D-M, Qun-Feng L, Li S-B, Wang J-P, Chen Y-L, Huang Y-Q, Bi H-K (2016) Comparative genomics of H. pylori and non-Pylori helicobacter species to identify new regions associated with its pathogenicity and adaptability. BioMed Res Int:6106029
Cao P, Guo D, Liu J, Jiang Q, Xu Z, Liandong Q (2017) Genome-wide analyses reveal genes subject to positive selection in Pasteurella multocida. Front Microbiol 8(May):961
Castillo D, Christiansen RH, Dalsgaard I, Madsen L, Espejo R, Middelboe M (2016) Comparative genome analysis provides insights into the pathogenicity of Flavobacterium psychrophilum. PLoS One 11(4):e0152515
Castillo D, Alvise PD, Xu R, Zhang F, Middelboe M, Gram L (2017) Comparative genome analyses of Vibrio anguillarum strains reveal a link with pathogenicity traits. mSystems 2(1):e00001. https://doi.org/10.1128/mSystems.00001-17
Castillo D, Pérez-Reytor D, Plaza N, Ramírez-Araya S, Blondel CJ, Corsini G, Bastías R et al (2018) Exploring the genomic traits of non-toxigenic Vibrio parahaemolyticus strains isolated in Southern Chile. Front Microbiol 9(February):161
Ceapa C, Davids M, Ritari J, Lambert J, Wels M, Douillard FP, Smokvina T, de Vos WM, Knol J, Kleerebezem M (2016) The variable regions of Lactobacillus rhamnosus genomes reveal the dynamic evolution of metabolic and host-adaptation repertoires. Genome Biol Evol 8(6):1889–1905
Chan AP, Sutton G, DePew J, Krishnakumar R, Choi Y, Huang X-Z, Beck E et al (2015) A novel method of consensus pan-chromosome assembly and large-scale comparative analysis reveal the highly flexible pan-genome of Acinetobacter baumannii. Genome Biol 16(July):143
Chaplin AV, Efimov BA, Smeianov VV, Kafarskaia LI, Pikina AP, Shkoporov AN (2015) Intraspecies genomic diversity and long-term persistence of Bifidobacterium longum. PLoS One 10(8):e0135658
Chen C, Wu L, Cao Q, Shao H, Li X, Zhang Y, Wang H, Tan X (2018) Genome comparison of different Zymomonas mobilis strains provides insights on conservation of the evolution. PLoS One 13(4):e0195994
Choo SW, Wee WY, Ngeow YF, Mitchell W, Tan JL, Wong GJ, Zhao Y, Xiao J (2014) Genomic reconnaissance of clinical isolates of emerging human pathogen Mycobacterium abscessus reveals high evolutionary potential. Sci Rep 4(February):4061
Chun J, Grim CJ, Hasan NA, Lee JH, Choi SY, Haley BJ, Taviani E et al (2009) Comparative genomics reveals mechanism for short-term and long-term clonal transitions in pandemic Vibrio cholerae. Proc Natl Acad Sci U S A 106(36):15442–15447
Chun BH, Kim KH, Jeon HH, Lee SH, Jeon CO (2017) Pan-genomic and transcriptomic analyses of Leuconostoc mesenteroides provide insights into its genomic and metabolic features and roles in Kimchi fermentation. Sci Rep 7(1):11504
Cortés MP, Mendoza SN, Travisany D, Gaete A, Siegel A, Cambiazo V, Maass A (2017) Analysis of Piscirickettsia salmonis metabolism using genome-scale reconstruction, modeling, and testing. Front Microbiol 8(December):2462
Coscollá M, Comas I, González-Candelas F (2011) Quantifying nonvertical inheritance in the evolution of Legionella pneumophila. Mol Biol Evol 28(2):985–1001
D’Amato F, Eldin C, Georgiades K, Edouard S, Delerce J, Labas N, Raoult D (2015) Loss of TSS1 in hypervirulent Coxiella burnetii 175, the causative agent of Q fever in French Guiana. Comp Immunol Microbiol Infect Dis 41(August):35–41
D’Auria G, Jiménez-Hernández N, Peris-Bondia F, Moya A, Latorre A (2010) Legionella pneumophila pangenome reveals strain-specific virulence factors. BMC Genomics 11(March):181
Delmont TO, Murat Eren A (2018) Linking pangenomes and metagenomes: the Prochlorococcus metapangenome. PeerJ 6(January):e4320
De M, Pieter WYC, Rubagotti E, Venter SN, Toth IK, Birch PRJ, Coutinho TA (2014) Analysis of the Pantoea ananatis pan-genome reveals factors underlying its ability to colonize and interact with plant, insect and vertebrate hosts. BMC Genomics 15(May):404
Deng X, Phillippy AM, Li Z, Salzberg SL, Zhang W (2010) Probing the pan-genome of Listeria monocytogenes: new insights into intraspecific niche expansion and genomic diversification. BMC Genomics 11(September):500
Dias GM, Bidault A, Le Chevalier P, Choquet G, Der Sarkissian C, Orlando L, Medigue C et al (2018) Vibrio tapetis displays an original type IV secretion system in strains pathogenic for Bivalve molluscs. Front Microbiol 9(February):227
Ding W, Baumdicker F, Neher RA (2018) panX: pan-genome analysis and exploration. Nucleic Acids Res 46(1):e5
Donati C, Luisa Hiller N, Tettelin H, Muzzi A, Croucher NJ, Angiuoli SV, Oggioni M et al (2010) Structure and dynamics of the pan-genome of Streptococcus pneumoniae and closely related species. Genome Biol 11(10):R107
Duchaud E, Rochat T, Habib C, Barbier P, Loux V, Guérin C, Dalsgaard I et al (2018) Genomic diversity and evolution of the fish pathogen Flavobacterium psychrophilum. Front Microbiol 9(February):138
Duranti S, Milani C, Lugli GA, Turroni F, Mancabelli L, Sanchez B, Ferrario C et al (2015) Insights from genomes of representatives of the human gut commensal Bifidobacterium bifidum. Environ Microbiol 17(7):2515–2531
Duranti S, Milani C, Lugli GA, Mancabelli L, Turroni F, Ferrario C, Mangifesta M et al (2016) Evaluation of genetic diversity among strains of the human gut commensal Bifidobacterium adolescentis. Sci Rep 6(April):23971
Eppinger M, Worsham PL, Nikolich MP, Riley DR, Sebastian Y, Mou S, Achtman M, Lindler LE, Ravel J (2010) Genome sequence of the deep-rooted Yersinia pestis strain Angola reveals new insights into the evolution and pangenome of the Plague bacterium. J Bacteriol 192(6):1685–1699
Fang Y, Li Z, Liu J, Shu C, Wang X, Zhang X, Yu X et al (2011) A pangenomic study of Bacillus thuringiensis. J Genet Genomics 38(12):567–576
Fernández-Romero N, Romero-Gómez MP, Mora-Rillo M, Rodríguez-Baño J, López- Cerero L, Pascual Á, Mingorance J (2015) Uncoupling between core genome and virulome in extraintestinal pathogenic Escherichia coli. Can J Microbiol 61(9):647–652
Ferrario C, Ricci G, Milani C, Lugli GA, Ventura M, Eraclio G, Borgo F, Fortina MG (2013) Lactococcus garvieae: where is it from? A first approach to explore the evolutionary history of this emerging pathogen. PLoS One 8(12):e84796
Fischer W, Windhager L, Rohrer S, Zeiller M, Karnholz A, Hoffmann R, Zimmer R, Haas R (2010) Strain-specific genes of Helicobacter pylori: genome evolution driven by a novel type IV secretion system and genomic island transfer. Nucleic Acids Res 38(18):6089–6101
Forgetta V, Oughton MT, Marquis P, Brukner I, Blanchette R, Haub K, Magrini V et al (2011) Fourteen-genome comparison identifies DNA markers for severe-disease-associated strains of Clostridium difficile. J Clin Microbiol 49(6):2230–2238
Frese SA, Benson AK, Tannock GW, Loach DM, Kim J, Zhang M, Phaik Lyn O et al (2011) The evolution of host specialization in the vertebrate gut symbiont Lactobacillus reuteri. PLoS Genet 7(2):e1001314
Galardini M, Mengoni A, Brilli M, Pini F, Fioravanti A, Lucas S, Lapidus A et al (2011) Exploring the symbiotic pangenome of the nitrogen-fixing bacterium Sinorhizobium meliloti. BMC Genomics 12(May):235
Galardini M, Pini F, Bazzicalupo M, Biondi EG, Mengoni A (2013) Replicon-dependent bacterial genome evolution: the case of Sinorhizobium meliloti. Genome Biol Evol 5(3):542–558
Garrido-Sanz D, Meier-Kolthoff JP, Göker M, Martín M, Rivilla R, Redondo-Nieto M (2016) Genomic and genetic diversity within the Pseudomonas fluorescens complex. PloS One 11(2):e0150183
Ghatak S, Blom J, Das S, Sanjukta R, Puro K, Mawlong M, Shakuntala I et al (2016) Pan-genome analysis of Aeromonas hydrophila, Aeromonas veronii and Aeromonas caviae indicates phylogenomic diversity and greater pathogenic potential for Aeromonas hydrophila. Antonie van Leeuwenhoek 109(7):945–956
Giampetruzzi A, Saponari M, Loconsole G, Boscia D, Savino VN, Almeida RPP, Zicca S, Landa BB, Chacón-Diaz C, Saldarelli P (2017) Genome-wide analysis provides evidence on the genetic relatedness of the emergent Xylella fastidiosa genotype in Italy to isolates from Central America. Phytopathology 107(7):816–827
Gómez-Lunar Z, Hernández-González I, Rodríguez-Torres M-D, Souza V, Olmedo-Álvarez G (2016) Microevolution analysis of Bacillus coahuilensis unveils differences in phosphorus acquisition strategies and their regulation. Front Microbiol 7(February):58
Gomila M, Busquets A, Mulet M, García-Valdés E, Lalucat J (2017) Clarification of taxonomic status within the Pseudomonas syringae species group based on a phylogenomic analysis. Front Microbiol 8(December):2422
Gordon DM, Geyik S, Clermont O, O’Brien CL, Huang S, Abayasekara C, Rajesh A et al (2017) Fine-scale structure analysis shows epidemic patterns of clonal complex 95, a cosmopolitan Escherichia coli lineage responsible for extraintestinal infection. mSphere 2(3):e00168. https://doi.org/10.1128/mSphere.00168-17
Grosso-Becerra M-V, Santos-Medellín C, González-Valdez A, Méndez J-L, Delgado G, Morales-Espinosa R, Servín-González L, Alcaraz L-D, Soberón-Chávez G (2014) Pseudomonas aeruginosa clinical and environmental isolates constitute a single population with high phenotypic diversity. BMC Genomics 15(April):318
Guo X, Li S, Zhang J, Feifan W, Li X, Dan W, Zhang M et al (2017) Genome sequencing of 39 Akkermansia muciniphila isolates reveals its population structure, genomic and functional diversity, and global distribution in mammalian gut microbiotas. BMC Genomics 18(1):800
Hall AB, Yassour M, Sauk J, Garner A, Jiang X, Arthur T, Lagoudas GK et al (2017) A novel Ruminococcus gnavus clade enriched in inflammatory bowel disease patients. Genome Med 9(1):103
Hassan A, Naz A, Obaid A, Paracha RZ, Naz K, Awan FM, Muhmmad SA, Janjua HA, Ahmad J, Ali A (2016) Pangenome and immuno-proteomics analysis of Acinetobacter baumannii strains revealed the core peptide vaccine targets. BMC Genomics 17(1):732
He E-M, Chen C-W, Guo Y, Hsu M-H, Liang Z, Chen H-L, Zhao G-P, Chiu C-H, Zhou Y (2017) The genome of serotype VI Streptococcus agalactiae serotype VI and comparative analysis. Gene 597(January):59–65
Hilker R, Munder A, Klockgether J, Losada PM, Chouvarine P, Cramer N, Davenport CF et al (2015) Interclonal gradient of virulence in the Pseudomonas aeruginosa pangenome from disease and environment. Environ Microbiol 17(1):29–46
Hiller NL, Janto B, Hogg JS, Boissy R, Susan Y, Powell E, Keefe R et al (2007) Comparative genomic analyses of seventeen Streptococcus pneumoniae strains: insights into the pneumococcal supragenome. J Bacteriol 189(22):8186–8195
Hollensteiner J, Poehlein A, Spröer C, Bunk B, Sheppard AE, Rosentstiel P, Schulenburg H, Liesegang H (2017) Complete genome sequence of the nematicidal Bacillus thuringiensis MYBT18246. Stand Genomic Sci 12(August):48
Holm KO, Bækkedal C, Söderberg JJ, Haugen P (2018) Complete genome sequences of seven Vibrio anguillarum strains as derived from PacBio sequencing. Genome Biol Evol 10(4):1127–1131
Howell KJ, Weinert LA, Chaudhuri RR, Luan S-L, Peters SE, Corander J, Harris D et al (2014) The use of genome wide association methods to investigate pathogenicity, population structure and serovar in Haemophilus parasuis. BMC Genomics 15(December):1179
Howell KJ, Weinert LA, Peters SE, Wang J, Hernandez-Garcia J, Chaudhuri RR, Luan S-L et al (2017) ‘Pathotyping’ multiplex PCR assay for Haemophilus parasuis: a tool for prediction of virulence. J Clin Microbiol 55(9):2617–2628
Huang Y, Kittichotirat W, Mayer MPA, Hall R, Bumgarner R, Chen C (2013) Comparative genomic hybridization and transcriptome analysis with a pan-genome microarray reveal distinctions between JP2 and non-JP2 genotypes of Aggregatibacter actinomycetemcomitans. Mol Oral Microbiol 28(1):1–17
Humbert J-F, Barbe V, Latifi A, Gugger M, Calteau A, Coursin T, Lajus A et al (2013) A tribute to disorder in the genome of the bloom-forming freshwater cyanobacterium Microcystis aeruginosa. PLoS One 8(8):e70747
Hurtado R, Carhuaricra D, Soares S, Viana MVC, Azevedo V, Maturrano L, Aburjaile F (2018) Pan-genomic approach shows insight of genetic divergence and pathogenic-adaptation of Pasteurella multocida. Gene 670(September):193–206
Imperi F, Antunes LCS, Blom J, Villa L, Iacono M, Visca P, Carattoli A (2011) The genomics of Acinetobacter baumannii: insights into genome plasticity, antimicrobial resistance and pathogenicity. IUBMB Life 63(12):1068–1074
Jacobsen A, Hendriksen RS, Aaresturp FM, Ussery DW, Friis C (2011) The Salmonella enterica pan-genome. Microb Ecol 62(3):487–504
Janvilisri T, Scaria J, Thompson AD, Nicholson A, Limbago BM, Arroyo LG, Glenn Songer J, Gröhn YT, Chang Y-F (2009) Microarray identification of Clostridium difficile core components and divergent regions associated with host origin. J Bacteriol 191(12):3881–3891
Jeong D-W, Heo S, Ryu S, Blom J, Lee J-H (2017) Genomic insights into the virulence and salt tolerance of Staphylococcus equorum. Sci Rep 7(1):5383
Joseph SJ, Didelot X, Gandhi K, Dean D, Read TD (2011) Interplay of recombination and selection in the genomes of Chlamydia trachomatis. Biol Direct 6(May):28
Joseph SJ, Cox D, Wolff B, Morrison SS, Kozak-Muiznieks NA, Frace M, Didelot X et al (2016) Dynamics of genome change among Legionella species. Sci Rep 6(September):33442
Kaas RS, Friis C, Ussery DW, Aarestrup FM (2012) Estimating variation within the genes and inferring the phylogeny of 186 sequenced diverse Escherichia coli genomes. BMC Genomics 13(October):577
Kadam A, Janto B, Eutsey R, Earl JP, Powell E, Dahlgren ME, Hu FZ, Ehrlich GD, Luisa Hiller N (2015) Streptococcus pneumoniae supragenome hybridization arrays for profiling of genetic content and gene expression. Curr Protoc Microbiol 36(February):9D-4
Kant R, Rintahaka J, Yu X, Sigvart-Mattila P, Paulin L, Mecklin J-P, Saarela M, Palva A, von Ossowski I (2014) A comparative pan-genome perspective of niche-adaptable cell-surface protein phenotypes in Lactobacillus rhamnosus. PLoS One 9(7):e102762
Karlsen C, Hjerde E, Klemetsen T, Willassen NP (2017) Pan genome and CRISPR analyses of the bacterial fish pathogen Moritella viscosa. BMC Genomics 18(1):313
Kawasaki M, Delamare-Deboutteville J, Bowater RO, Walker MJ, Beatson S, Ben Zakour NL, Barnes AC (2018) Microevolution of aquatic Streptococcus agalactiae ST-261 from Australia indicates dissemination via imported tilapia and ongoing adaptation to marine hosts or environment. Appl Environ Microbiol 84(16):e00859. https://doi.org/10.1128/AEM.00859-18
Kayansamruaj P, Pirarat N, Kondo H, Hirono I, Rodkhum C (2015) Genomic comparison between pathogenic Streptococcus agalactiae isolated from Nile Tilapia in Thailand and fish-derived ST7 strains. Infect Genet Evol 36(December):307–314
Kayansamruaj P, Dong HT, Hirono I, Kondo H, Senapin S, Rodkhum C (2017) Comparative genome analysis of fish pathogen Flavobacterium columnare reveals extensive sequence diversity within the species. Infect Genet Evol 54(October):7–17
Kelleher P, Bottacini F, Mahony J, Kilcawley KN, van Sinderen D (2017) Comparative and functional genomics of the Lactococcus lactis taxon; insights into evolution and niche adaptation. BMC Genomics 18(1):267
Kim EB, Marco ML (2014) Nonclinical and clinical Enterococcus faecium strains, but not Enterococcus faecalis strains, have distinct structural and functional genomic features. Appl Environ Microbiol 80(1):154–165
Kim Y, Koh I, Mi YL, Chung W-H, Rho M (2017) Pan-genome analysis of Bacillus for microbiome profiling. Sci Rep 7(1):10984
Kirk KF, Méric G, Nielsen HL, Pascoe B, Sheppard SK, Thorlacius-Ussing O, Nielsen H (2018) Molecular epidemiology and comparative genomics of campylobacter concisus strains from saliva, faeces and gut mucosal biopsies in inflammatory bowel disease. Sci Rep 8(1):1902
Kittichotirat W, Bumgarner RE, Asikainen S, Chen C (2011) Identification of the pangenome and its components in 14 distinct Aggregatibacter actinomycetemcomitans strains by comparative genomic analysis. PLoS One 6(7):e22420
Kiu R, Caim S, Alexander S, Pachori P, Hall LJ (2017) Probing genomic aspects of the multi-host pathogen Clostridium perfringens reveals significant pangenome diversity, and a diverse array of virulence factors. Front Microbiol 8(December):2485
Klockgether J, Cramer N, Wiehlmann L, Davenport CF, Tümmler B (2011) Pseudomonas aeruginosa genomic structure and diversity. Front Microbiol 2(July):150
Knight DR, Squire MM, Collins DA, Riley TV (2016) Genome analysis of Clostridium difficile PCR ribotype 014 lineage in Australian pigs and humans reveals a diverse genetic repertoire and signatures of long-range interspecies transmission. Front Microbiol 7:2138
Koton Y, Gordon M, Chalifa-Caspi V, Bisharat N (2014) Comparative genomic analysis of clinical and environmental Vibrio vulnificus isolates revealed biotype 3 evolutionary relationships. Front Microbiol 5:803
Kubasova T, Cejkova D, Matiasovicova J, Sekelova Z, Polansky O, Medvecky M, Rychlik I, Juricova H (2016) Antibiotic resistance, core-genome and protein expression in IncHI1 plasmids in Salmonella typhimurium. Genome Biol Evol 8(6):1661–1671
Kuenne C, Billion A, Mraheil MA, Strittmatter A, Daniel R, Goesmann A, Barbuddhe S, Hain T, Chakraborty T (2013) Reassessment of the Listeria monocytogenes pan-genome reveals dynamic integration hotspots and mobile genetic elements as major components of the accessory genome. BMC Genomics 14(January):47
Kunadu P-H, Angela MH, Miller EL, Grant AJ (2018) Microbiological quality and antimicrobial resistance characterization of Salmonella spp. in fresh milk value chains in Ghana. Int J Food Microbiol 277(July):41–49
Lacey JA, Allnutt TR, Vezina B, Van TTH, Stent T, Han X, Rood JI et al (2018) Whole genome analysis reveals the diversity and evolutionary relationships between necrotic enteritis-causing strains of Clostridium perfringens. BMC Genomics 19(1):379
Laing C, Villegas A, Taboada EN, Kropinski A, Thomas JE, Gannon VPJ (2011) Identification of Salmonella enterica species- and subgroup-specific genomic regions using Panseq 2.0. Infect Genet Evol 11(8):2151–2161
Laing CR, Whiteside MD, Gannon VPJ (2017) Pan-genome analyses of the species Salmonella enterica, and identification of genomic markers predictive for species, subspecies, and serovar. Front Microbiol 8(July):1345
Lebreton F, van Schaik W, McGuire AM, Godfrey P, Griggs A, Mazumdar V, Corander J et al (2013) Emergence of epidemic multidrug-resistant Enterococcus faecium from animal and commensal strains. mBio 4(4):e00534. https://doi.org/10.1128/mBio.00534-13
Lee J-Y, Han GG, Choi J, Jin G-D, Kang S-K, Chae BJ, Kim EB, Choi Y-J (2017a) Pan-genomic approaches in Lactobacillus reuteri as a porcine probiotic: investigation of host adaptation and antipathogenic activity. Microb Ecol 74(3):709–721
Lee J-Y, Han GG, Kim EB, Choi Y-J (2017b) Comparative genomics of Lactobacillus salivarius strains focusing on their host adaptation. Microbiol Res 205(December):48–58
Leekitcharoenphon P, Lukjancenko O, Friis C, Aarestrup FM, Ussery DW (2012) Genomic variation in Salmonella enterica core genes for epidemiological typing. BMC Genomics 13(March):88
Liang W, Zhao Y, Chen C, Cui X, Yu J, Xiao J, Kan B (2012) Pan-genomic analysis provides insights into the genomic variation and evolution of Salmonella paratyphi A. PLoS One 7(9):e45346
Li G, Shen M, Le S, Tan Y, Li M, Zhao X, Shen W et al (2016) Genomic analyses of multidrug resistant Pseudomonas aeruginosa PA1 resequenced by single-molecule real-time sequencing. Biosci Rep 36(6):e00418. https://doi.org/10.1042/BSR20160282
Lira F, Berg G, Martínez JL (2017) Double-face meets the bacterial world: the opportunistic pathogen Stenotrophomonas maltophilia. Front Microbiol 8(November):2190
Liu W, Fang L, Li M, Li S, Guo S, Luo R, Feng Z et al (2012) Comparative genomics of mycoplasma: analysis of conserved essential genes and diversity of the pan-genome. PLoS One 7(4):e35698
Liu L, Zhu W, Cao Z, Biao X, Wang G, Luo M (2015) High correlation between genotypes and phenotypes of environmental bacteria Comamonas testosteroni strains. BMC Genomics 16(February):110
Liu G, Zhang W, Chengping L (2013a) Comparative genomics analysis of Streptococcus agalactiae reveals that isolates from cultured Tilapia in China are closely related to the human strain A909. BMC Genomics 14(November):775
Liu W-Y, Wong C-F, Chung KM-K, Jiang J-W, Leung FC-C (2013b) Comparative genome analysis of Enterobacter cloacae. PLoS One 8(9):e74487
Liu Y-Y, Chen C-C, Chiou C-S (2016a) Construction of a pan-genome allele database of Salmonella enterica serovar enteritidis for molecular subtyping and disease cluster identification. Front Microbiol 7(December):2010
Liu Y-Y, Chiou C-S, Chen C-C (2016b) PGAdb-builder: a web service tool for creating pan-genome allele database for molecular fine typing. Sci Rep 6(November):36213
Loper JE, Hassan KA, Mavrodi DV, Davis EW 2nd, Lim CK, Shaffer BT, Elbourne LDH et al (2012) Comparative genomics of plant-associated Pseudomonas spp.: insights into diversity and inheritance of traits involved in multitrophic interactions. PLoS Genet 8(7):e1002784
López-Hermoso C, de la Haba RR, Sánchez-Porro C, Ventosa A (2018) Salinivibrio kushneri sp. nov., a moderately halophilic bacterium isolated from salterns. Syst Appl Microbiol 41(3):159–166
Lukjancenko O, Wassenaar TM, Ussery DW (2010) Comparison of 61 sequenced Escherichia Coli genomes. Microb Ecol 60(4):708–720
Lu W, Wise MJ, Tay CY, Windsor HM, Marshall BJ, Peacock C, Perkins T (2014) Comparative analysis of the full genome of Helicobacter pylori isolate Sahul64 identifies genes of high divergence. J Bacteriol 196(5):1073–1083
Malmstrom RR, Rodrigue S, Huang KH, Kelly L, Kern SE, Thompson A, Roggensack S, Berube PM, Henn MR, Chisholm SW (2013) Ecology of uncultured Prochlorococcus clades revealed through single-cell genomics and biogeographic analysis. ISME J 7(1):184–198
Mann RA, Smits THM, Bühlmann A, Blom J, Goesmann A, Frey JE, Plummer KM et al (2013) Comparative genomics of 12 strains of Erwinia Amylovora identifies a pan-genome with a large conserved core. PLoS One 8(2):e55644
Manzano-Marín A, Lamelas A, Moya A, Latorre A (2012) Comparative genomics of Serratia spp.: two paths towards endosymbiotic life. PLoS One 7(10):e47274
Meng P, Lu C, Zhang Q, Lin J, Chen F (2017) Exploring the genomic diversity and cariogenic differences of Streptococcus mutans strains through pan-genome and comparative genome analysis. Curr Microbiol 74(10):1200–1209
Méric G, Yahara K, Mageiros L, Pascoe B, Maiden MCJ, Jolley KA, Sheppard SK (2014) A reference pan-genome approach to comparative bacterial genomics: identification of novel epidemiological markers in pathogenic campylobacter. PLoS One 9(3):e92798
Méric G, Miragaia M, de Been M, Yahara K, Pascoe B, Mageiros L, Mikhail J et al (2015) Ecological overlap and horizontal gene transfer in Staphylococcus aureus and Staphylococcus epidermidis. Genome Biol Evol 7(5):1313–1328
Méric G, Mageiros L, Pascoe B, Woodcock DJ, Mourkas E, Lamble S, Bowden R, Jolley KA, Raymond B, Sheppard SK (2018) Lineage-specific plasmid acquisition and the evolution of specialized pathogens in Bacillus thuringiensis and the Bacillus cereus group. Mol Ecol 27(7):1524–1540
Milani C, Duranti S, Lugli GA, Bottacini F, Strati F, Arioli S, Foroni E, Turroni F, van Sinderen D, Ventura M (2013) Comparative genomics of Bifidobacterium animalis subsp. lactis reveals a strict monophyletic bifidobacterial taxon. Appl Environ Microbiol 79(14):4304–4315
Miyauchi E, Toh H, Nakano A, Tanabe S, Morita H (2012) Comparative genomic analysis of Lactococcus garvieae strains isolated from different sources reveals candidate virulence genes. Int J Microbiol 2012(May):728276
Mongodin EF, Casjens SR, Bruno JF, Yun X, Drabek EF, Riley DR, Cantarel BL et al (2013) Inter- and intra-specific pan-genomes of Borrelia burgdorferi Sensu Lato: genome stability and adaptive radiation. BMC Genomics 14(October):693
Monk JM, Charusanti P, Aziz RK, Lerman JA, Premyodhin N, Orth JD, Feist AM, Palsson BØ (2013) Genome-scale metabolic reconstructions of multiple Escherichia coli strains highlight strain-specific adaptations to nutritional environments. Proc Natl Acad Sci U S A 110(50):20338–20343
Mosquera-Rendón J, Rada-Bravo AM, Cárdenas-Brito S, Corredor M, Restrepo-Pineda E, Benítez-Páez A (2016) Pangenome-wide and molecular evolution analyses of the Pseudomonas aeruginosa species. BMC Genomics 17(January):45
Murillo T, Ramírez-Vargas G, Riedel T, Overmann J, Andersen JM, Guzmán-Verri C, Chaves-Olarte E, Rodríguez C (2018) Two groups of cocirculating, epidemic clostridiodes difficile strains microdiversify through different mechanisms. Genome Biol Evol 10(3):982–998
Nguyen TL, Kim D-H (2018) Genome-wide comparison reveals a probiotic strain Lactococcus lactis WFLU12 isolated from the gastrointestinal tract of olive flounder (Paralichthys olivaceus) harboring genes supporting probiotic action. Mar Drugs 16(5):140. https://doi.org/10.3390/md16050140
Nourdin-Galindo G, Sánchez P, Molina CF, Espinoza-Rojas DA, Oliver C, Ruiz P, Vargas-Chacoff L et al (2017) Comparative pan-genome analysis of Piscirickettsia salmonis reveals genomic divergences within genogroups. Front Cell Infect Microbiol 7(October):459
O’Callaghan A, Bottacini F, O’Connell Motherway M, van Sinderen D (2015) Pangenome analysis of Bifidobacterium longum and site-directed mutagenesis through by-pass of restriction-modification systems. BMC Genomics 16(October):832
Ogunremi D, Devenish J, Amoako K, Kelly H, Dupras AA, Belanger S, Wang LR (2014) High resolution assembly and characterization of genomes of Canadian isolates of Salmonella enteritidis. BMC Genomics 15(August):713
Ojala T, Kankainen M, Castro J, Cerca N, Edelman S, Westerlund-Wikström B, Paulin L, Holm L, Auvinen P (2014) Comparative genomics of Lactobacillus crispatus suggests novel mechanisms for the competitive exclusion of Gardnerella vaginalis. BMC Genomics 15(December):1070
Okura M, Nozawa T, Watanabe T, Murase K, Nakagawa I, Takamatsu D, Osaki M et al (2017) A locus encoding variable defence systems against invading DNA identified in Streptococcus suis. Genome Biol Evol 9(4):1000. https://doi.org/10.1093/gbe/evx062
Orata FD, Kirchberger PC, Méheust R, Barlow EJ, Tarr CL, Boucher Y (2015) The dynamics of genetic interactions between Vibrio metoecus and Vibrio cholerae, two close relatives co-occurring in the environment. Genome Biol Evol 7(10):2941–2954
Otchere ID, Harris SR, Busso SL, Asante-Poku A, Osei-Wusu S, Koram K, Parkhill J, Gagneux S, Yeboah-Manu D (2016) The first population structure and comparative genomics analysis of mycobacterium africanum strains from Ghana reveals higher diversity of lineage. Int J Mycobact 5(Suppl 1):S80–S81
Palmer SR, Miller JH, Abranches J, Zeng L, Lefebure T, Richards VP, Lemos JA, Stanhope MJ, Burne RA (2013) Phenotypic heterogeneity of genomically-diverse isolates of Streptococcus mutans. PLoS One 8(4):e61358
Periwal V, Patowary A, Vellarikkal SK, Gupta A, Singh M, Mittal A, Jeyapaul S et al (2015) Comparative whole-genome analysis of clinical isolates reveals characteristic architecture of Mycobacterium tuberculosis pangenome. PLoS One 10(4):e0122979
Qin X, Galloway-Peña JR, Sillanpaa J, Roh JH, Nallapareddy SR, Chowdhury S, Bourgogne A et al (2012) Complete genome sequence of Enterococcus faecium strain TX16 and comparative genomic analysis of Enterococcus faecium genomes. BMC Microbiol 12(July):135
Remenant B, Coupat-Goutaland B, Guidot A, Cellier G, Wicker E, Allen C, Fegan M et al (2010) Genomes of three tomato pathogens within the Ralstonia solanacearum species complex reveal significant evolutionary divergence. BMC Genomics 11(June):379
Reno ML, Held NL, Fields CJ, Burke PV, Whitaker RJ (2009) Biogeography of the Sulfolobus islandicus pan-genome. Proc Natl Acad Sci U S A 106(21):8605–8610
Roisin S, Gaudin C, De Mendonça R, Bellon J, Van Vaerenbergh K, De Bruyne K, Byl B, Pouseele H, Denis O, Supply P (2016) Pan-genome multilocus sequence typing and outbreak-specific reference-based single nucleotide polymorphism analysis to resolve two concurrent Staphylococcus aureus outbreaks in neonatal services. Clin Microbiol Infect 22(6):520–526
Rouleau FD, Vincent AT, Charette SJ (2018) Genomic and phenotypic characterization of an atypical Aeromonas salmonicida strain isolated from a lumpfish and producing unusual granular structures. J Fish Dis 41(4):673–681
Rouli L, MBengue M, Robert C, Ndiaye M, La Scola B, Raoult D (2014) Genomic analysis of three African strains of Bacillus anthracis demonstrates that they are part of the clonal expansion of an exclusively pathogenic bacterium. New Microb New Infect 2(6):161–169
Sangal V, Blom J, Sutcliffe IC, von Hunolstein C, Burkovski A, Hoskisson PA (2015) Adherence and invasive properties of Corynebacterium diphtheriae strains correlates with the predicted membrane-associated and secreted proteome. BMC Genomics 16(October):765
Sassi M, Drancourt M (2014) Genome analysis reveals three genomospecies in Mycobacterium abscessus. BMC Genomics 15(May):359
Scaria J, Ponnala L, Janvilisri T, Yan W, Mueller LA, Chang Y-F (2010) Analysis of ultra low genome conservation in Clostridium difficile. PLoS One 5(12):e15147
Sela U, Euler CW, Correa da Rosa J, Fischetti VA (2018) Strains of bacterial species induce a greatly varied acute adaptive immune response: the contribution of the accessory genome. PLoS Pathog 14(1):e1006726
Shariati J V, Malboobi MA, Tabrizi Z, Tavakol E, Owilia P, Safari M (2017) Comprehensive genomic analysis of a plant growth-promoting rhizobacterium pantoea agglomerans strain P5. Sci Rep 7(1):15610
Sharma PK, Jilagamazhi F, Zhang X, Fristensky B, Sparling R, Levin DB (2014) Genome features of Pseudomonas putida LS46, a novel polyhydroxyalkanoate producer and its comparison with other P. putida strains. AMB Express 4(May):37
Silby MW, Cerdeño-Tárraga AM, Vernikos GS, Giddens SR, Jackson RW, Preston GM, Zhang X-X et al (2009) Genomic and genetic analyses of diversity and plant interactions of Pseudomonas fluorescens. Genome Biol 10(5):R51
Smokvina T, Wels M, Polka J, Chervaux C, Brisse S, Boekhorst J, Vlieg JET v H, Siezen RJ (2013) Lactobacillus paracasei comparative genomics: towards species pan-genome definition and exploitation of diversity. PLoS One 8(7):e68731
Snipen L, Almøy T, Ussery DW (2009) Microbial comparative pan-genomics using binomial mixture models. BMC Genomics 10(August):385
Soares SC, Silva A, Trost E, Blom J, Ramos R, Carneiro A, Ali A et al (2013) The pan-genome of the animal pathogen Corynebacterium pseudotuberculosis reveals differences in genome plasticity between the Biovar ovis and equi strains. PLoS One 8(1):e53818
Song L, Wang W, Conrads G, Rheinberg A, Sztajer H, Reck M, Wagner-Döbler I, Ping Zeng A (2013) Genetic variability of Mutans streptococci revealed by wide whole-genome sequencing. BMC Genomics 14(June):430
Spring-Pearson SM, Stone JK, Doyle A, Allender CJ, Okinaka RT, Mayo M, Broomall SM et al (2015) Pangenome analysis of Burkholderia pseudomallei: genome evolution preserves gene order despite high recombination rates. PLoS One 10(10):e0140274
Stabler RA, Gerding DN, Songer JG, Drudy D, Brazier JS, Trinh HT, Witney AA, Hinds J, Wren BW (2006) Comparative phylogenomics of Clostridium difficile reveals clade specificity and microevolution of hypervirulent strains. J Bacteriol 188(20):7297–7305
Stanborough T, Fegan N, Powell SM, Singh T, Tamplin M, Scott Chandry P (2018) Genomic and metabolic characterization of spoilage-associated Pseudomonas species. Int J Food Microbiol 268(March):61–72
Sternes PR, Borneman AR (2016) Consensus pan-genome assembly of the specialised wine bacterium Oenococcus oeni. BMC Genomics 17(April):308
Stice SP, Stumpf SD, Gitaitis RD, Kvitko BH, Dutta B (2018) Pantoea ananatis genetic diversity analysis reveals limited genomic diversity as well as accessory genes correlated with onion pathogenicity. Front Microbiol 9(February):184
Sultanov RI, Arapidi GP, Vinogradova SV, Govorun VM, Luster DG, Ignatov AN (2016) Comprehensive analysis of draft genomes of two closely related Pseudomonas syringae phylogroup 2b strains infecting mono- and dicotyledon host plants. BMC Genomics 17(Suppl 14):1010
Sun S, Xiao J, Zhang H, Zhang Z (2016) Pangenome evidence for higher codon usage bias and stronger translational selection in core genes of Escherichia coli. Front Microbiol 7(August):1180
Sváb D, Bálint B, Maróti G, Tóth I (2016) Cytolethal distending toxin producing Escherichia coli O157:H43 strain T22 represents a novel evolutionary lineage within the O157 serogroup. Infect Genet Evol 46(December):110–117
Thépault A, Méric G, Rivoal K, Pascoe B, Mageiros L, Touzain F, Rose V, Béven V, Chemaly M, Sheppard SK (2017) Genome-wide identification of host- segregating epidemiological markers for source attribution in Campylobacter jejuni. Appl Environ Microbiol 83(7):e03085. https://doi.org/10.1128/AEM.03085-16
Timms VJ, Rockett R, Bachmann NL, Martinez E, Wang Q, Chen SC-A, Jeoffreys N et al (2018) Genome sequencing links persistent outbreak of legionellosis in Sydney (New South Wales, Australia) to an emerging clone of Legionella pneumophila sequence type 211. Appl Environ Microbiol 84(5):e02020. https://doi.org/10.1128/AEM.02020-17
Tomida S, Nguyen L, Chiu B-H, Liu J, Sodergren E, Weinstock GM, Li H (2013) Pan-genome and comparative genome analyses of Propionibacterium acnes reveal its genomic diversity in the healthy and diseased human skin microbiome. mBio 4(3):e00003
Touchon M, Hoede C, Tenaillon O, Barbe V, Baeriswyl S, Bidet P, Bingen E et al (2009) Organised genome dynamics in the Escherichia coli species results in highly diverse adaptive paths. PLoS Genet 5(1):e1000344
Trost E, Blom J, Soares S d C, Huang I-H, Al-Dilaimi A, Schröder J, Jaenicke S et al (2012) Pangenomic study of Corynebacterium diphtheriae that provides insights into the genomic diversity of pathogenic isolates from cases of classical diphtheria, endocarditis, and pneumonia. J Bacteriol 194(12):3199–3215
Uchiyama I, Albritton J, Fukuyo M, Kojima KK, Yahara K, Kobayashi I (2016) A novel approach to Helicobacter pylori pan-genome analysis for identification of genomic islands. PLoS One 11(8):e0159419
Udaondo Z, Molina L, Segura A, Duque E, Ramos JL (2016) Analysis of the core genome and pangenome of Pseudomonas putida. Environ Microbiol 18(10):3268–3283
van Schaik W, Top J, Riley DR, Boekhorst J, Vrijenhoek JEP, Schapendonk CME, Hendrickx APA et al (2010) Pyrosequencing-based comparative genome analysis of the nosocomial pathogen Enterococcus faecium and identification of a large transferable pathogenicity island. BMC Genomics 11(April):239
Vesth T, Wassenaar TM, Hallin PF, Snipen L, Lagesen K, Ussery DW (2010) On the origins of a Vibrio species. Microb Ecol 59(1):1–13
Viver T, Orellana L, González-Torres P, Díaz S, Urdiain M, Farías ME, Benes V et al (2018) Genomic comparison between members of the Salinibacteraceae family, and description of a new species of Salinibacter (Salinibacter altiplanensis sp. nov.) isolated from high altitude hypersaline environments of the Argentinian Altiplano. Syst Appl Microbiol 41(3):198–212
Wang J, Haapalainen M, Schott T, Thompson SM, Smith GR, Nissinen AI, Pirhonen M (2017) Genomic sequence of ‘candidatus liberibacter solanacearum’ haplotype C and its comparison with haplotype A and B genomes. PLoS One 12(2):e0171531
Wegmann U, MacKenzie DA, Zheng J, Goesmann A, Roos S, Swarbreck D, Walter J, Crossman LC, Juge N (2015) The pan-genome of Lactobacillus reuteri strains originating from the pig gastrointestinal tract. BMC Genomics 16(December):1023
Weller-Stuart T, De Maayer P, Coutinho T (2017) Pantoea ananatis: genomic insights into a versatile pathogen. Mol Plant Pathol 18(9):1191–1198
Wilkinson DA, O’Donnell AJ, Akhter RN, Fayaz A, Mack HJ, Rogers LE, Biggs PJ, French NP, Midwinter AC (2018) Updating the genomic taxonomy and epidemiology of Campylobacter hyointestinalis. Sci Rep 8(1):2393
Willenbrock H, Hallin PF, Wassenaar TM, Ussery DW (2007) Characterization of probiotic Escherichia coli isolates with a novel pan-genome microarray. Genome Biol 8(12):R267
Williams TM, Loman NJ, Ebruke C, Musher DM, Adegbola RA, Pallen MJ, Weinstock GM, Antonio M (2012) Genome analysis of a highly virulent serotype 1 strain of Streptococcus pneumoniae from West Africa. PLoS One 7(10):e26742
Xu Z, Chen X, Li L, Li T, Wang S, Chen H, Zhou R (2010) Comparative genomic characterization of Actinobacillus pleuropneumoniae. J Bacteriol 192(21):5625–5636
Yang J, Yang S (2017) Comparative analysis of corynebacterium glutamicum genomes: a new perspective for the industrial production of amino acids. BMC Genomics 18(Suppl 1):940
Yu G, Wang XC, Tian WH, Shi JC, Wang B, Ye Q, Dong SG, Zeng M, Wang JZ (2015) Genomic diversity and evolution of Bacillus subtilis. Biomed Environ Sci 28(8):620–625
Yu D, Yin Z, Li B, Jin Y, Ren H, Zhou J, Zhou W, Liang L, Yue J (2016) Gene flow, recombination, and positive selection in Stenotrophomonas maltophilia: mechanisms underlying the diversity of the widespread opportunistic pathogen. Genome 59(12):1063–1075
Yu J, Zhao J, Song Y, Zhang J, Yu Z, Zhang H, Sun Z (2018) Comparative genomics of the herbivore gut symbiont Lactobacillus reuteri reveals genetic diversity and lifestyle adaptation. Front Microbiol 9(June):1151
Yue M, Rankin SC, Blanchet RT, Nulton JD, Edwards RA, Schifferli DM (2012) Diversification of the Salmonella fimbriae: a model of macro- and microevolution. PLoS One 7(6):e38596
Zhang A, Yang M, Hu P, Wu J, Chen B, Hua Y, Yu J, Chen H, Xiao J, Jin M (2011a) Comparative genomic analysis of Streptococcus suis reveals significant genomic diversity among different serotypes. BMC Genomics 12(October):523
Zhang J, van Aartsen JJ, Jiang X, Shao Y, Tai C, He X, Tan Z et al (2011b) Expansion of the known Klebsiella pneumoniae species gene pool by characterization of novel alien DNA islands integrated into tmRNA gene sites. J Microbiol Methods 84(2):283–289
Zhang D-F, Zhi X-Y, Zhang J, Paoli GC, Cui Y, Shi C, Shi X (2017) Preliminary comparative genomics revealed pathogenic potential and international spread of Staphylococcus argenteus. BMC Genomics 18(1):808
Zhang X, Liu X, Yang F, Chen L (2018) Pan-genome analysis links the hereditary variation of Leptospirillum ferriphilum with its evolutionary adaptation. Front Microbiol 9(March):577
Zhao Y, Sun C, Zhao D, Zhang Y, You Y, Jia X, Yang J et al (2018) PGAP-X: extension on pan-genome analysis pipeline. BMC Genomics 19(Suppl 1):36
Zhou Z, McCann A, Litrup E, Murphy R, Cormican M, Fanning S, Brown D, Guttman DS, Brisse S, Achtman M (2013) Neutral genomic microevolution of a recently emerged pathogen, Salmonella enterica serovar agona. PLoS Genet 9(4):e1003471
Zhu Ge X, Jiang J, Pan Z, Hu L, Wang S, Wang H, Leung FC, Dai J, Fan H (2014) Comparative genomic analysis shows that avian pathogenic Escherichia coli isolate IMT5155 (O2:K1:H5; ST Complex 95, ST140) shares close relationship with ST95 APEC O1:K1 and human ExPEC O18:K1 strains. PLoS One 9(11):e112048
Author information
Authors and Affiliations
Corresponding authors
Editor information
Editors and Affiliations
Rights and permissions
Open Access This chapter is licensed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license and indicate if changes were made.
The images or other third party material in this chapter are included in the chapter's Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the chapter's Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder.
Copyright information
© 2020 The Author(s)
About this chapter
Cite this chapter
Innamorati, K.A., Earl, J.P., Aggarwal, S.D., Ehrlich, G.D., Hiller, N.L. (2020). The Bacterial Guide to Designing a Diversified Gene Portfolio. In: Tettelin, H., Medini, D. (eds) The Pangenome. Springer, Cham. https://doi.org/10.1007/978-3-030-38281-0_3
Download citation
DOI: https://doi.org/10.1007/978-3-030-38281-0_3
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-030-38280-3
Online ISBN: 978-3-030-38281-0
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)