Abstract
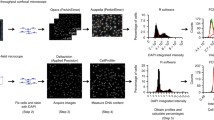
The development of technologies that allow measurement of the cell cycle at the single-cell level has revealed novel insights into the mechanisms that regulate cell cycle commitment and progression through DNA replication and cell division. These studies have also provided evidence of heterogeneity in cell cycle regulation among individual cells, even within a genetically identical population. Cell cycle mapping combines highly multiplexed imaging with manifold learning to visualize the diversity of “paths” that cells can take through the proliferative cell cycle or into various states of cell cycle arrest. In this chapter, we describe a general protocol of the experimental and computational components of cell cycle mapping. We also provide a comprehensive guide for the design and analysis of experiments, discussing key considerations in detail (e.g., antibody library preparation, analysis strategies, etc.) that may vary depending on the research question being addressed.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Nurse P (2000) A long twentieth century of the cell cycle and beyond. Cell 100:71–78. https://doi.org/10.1016/s0092-8674(00)81684-0
Sakaue-Sawano A, Kurokawa H, Morimura T et al (2008) Visualizing spatiotemporal dynamics of multicellular cell-cycle progression. Cell 132:487–498. https://doi.org/10.1016/j.cell.2007.12.033
Gut G, Tadmor MD, Pe’er D et al (2015) Trajectories of cell-cycle progression from fixed cell populations. Nat Methods 12:951–954. https://doi.org/10.1038/nmeth.3545
Gookin S, Min M, Phadke H et al (2017) A map of protein dynamics during cell-cycle progression and cell-cycle exit. PLoS Biol 15:e2003268. https://doi.org/10.1371/journal.pbio.2003268
Zerjatke T, Gak IA, Kirova D et al (2017) Quantitative cell cycle analysis based on an endogenous all-in-one reporter for cell tracking and classification. Cell Rep 19:1953–1966. https://doi.org/10.1016/j.celrep.2017.05.022
Rappez L, Rakhlin A, Rigopoulos A et al (2020) DeepCycle reconstructs a cyclic cell cycle trajectory from unsegmented cell images using convolutional neural networks. Mol Syst Biol 16:e9474. https://doi.org/10.15252/msb.20209474
Stallaert W, Kedziora KM, Taylor CD et al (2022) The structure of the human cell cycle. Cell Syst 13:230–240.e3. https://doi.org/10.1016/j.cels.2021.10.007
Spencer SL, Cappell SD, Tsai FC et al (2013) The proliferation-quiescence decision is controlled by a bifurcation in CDK2 activity at mitotic exit. Cell 155:369–383. https://doi.org/10.1016/j.cell.2013.08.062
Overton KW, Spencer SL, Noderer WL et al (2014) Basal p21 controls population heterogeneity in cycling and quiescent cell cycle states. Proc Natl Acad Sci U S A 111:E4386–E4393. https://doi.org/10.1073/pnas.1409797111
Stallaert W, Taylor SR, Kedziora KM et al (2022) The molecular architecture of cell cycle arrest. Mol Syst Biol 18:e11087. https://doi.org/10.15252/msb.202211087
Liu C, Konagaya Y, Chung M et al (2020) Altered G1 signaling order and commitment point in cells proliferating without CDK4/6 activity. Nat Commun 11:5305. https://doi.org/10.1038/s41467-020-18966-9
Knudsen ES, Kumarasamy V, Nambiar R et al (2022) CDK/cyclin dependencies define extreme cancer cell-cycle heterogeneity and collateral vulnerabilities. Cell Rep 38:110448. https://doi.org/10.1016/j.celrep.2022.110448
Stallaert W, Kedziora KM, Chao HX, Purvis JE (2019) Bistable switches as integrators and actuators during cell cycle progression. FEBS Lett 593:2805–2816. https://doi.org/10.1002/1873-3468.13628
Zou T, Lin Z (2021) The involvement of ubiquitination machinery in cell cycle regulation and cancer progression. Int J Mol Sci 22. https://doi.org/10.3390/ijms22115754
Maiani E, Milletti G, Nazio F et al (2021) AMBRA1 regulates cyclin D to guard S-phase entry and genomic integrity. Nature 592:799. https://doi.org/10.1038/s41586-021-03422-5
Simoneschi D, Rona G, Zhou N et al (2021) CRL4AMBRA1 is a master regulator of D-type cyclins. Nature 592:789. https://doi.org/10.1038/s41586-021-03445-y
Gut G, Herrmann MD, Pelkmans L (2018) Multiplexed protein maps link subcellular organization to cellular states. Science 361. https://doi.org/10.1126/science.aar7042
Radtke AJ, Chu CJ, Yaniv Z et al (2022) IBEX: an iterative immunolabeling and chemical bleaching method for high-content imaging of diverse tissues. Nat Protoc 1–26:378. https://doi.org/10.1038/s41596-021-00644-9
Black S, Phillips D, Hickey JW et al (2021) CODEX multiplexed tissue imaging with DNA-conjugated antibodies. Nat Protoc 16:3802. https://doi.org/10.1038/s41596-021-00556-8
Lin J-R, Izar B, Wang S et al (2018) Highly multiplexed immunofluorescence imaging of human tissues and tumors using t-CyCIF and conventional optical microscopes. elife 7. https://doi.org/10.7554/eLife.31657
Lee J-Y, Kitaoka M (2018) A beginner’s guide to rigor and reproducibility in fluorescence imaging experiments. Mol Biol Cell 29:1519–1525. https://doi.org/10.1091/mbc.E17-05-0276
Ogama T (2020) A beginner’s guide to improving image acquisition in fluorescence microscopy. Biochemist 42:22–27. https://doi.org/10.1042/BIO20200075
Montero Llopis P, Senft RA, Ross-Elliott TJ et al (2021) Best practices and tools for reporting reproducible fluorescence microscopy methods. Nat Methods 18:1463–1476. https://doi.org/10.1038/s41592-021-01156-w
van der Walt S, Schönberger JL, Nunez-Iglesias J et al (2014) Scikit-image: image processing in Python. PeerJ 2:e453. https://doi.org/10.7717/peerj.453
Chiu C-L, Clack N, The Napari Community (2022) Napari: a python multi-dimensional image viewer platform for the research community. Microsc Microanal 28:1576–1577. https://doi.org/10.1017/S1431927622006328
Schindelin J, Arganda-Carreras I, Frise E et al (2012) Fiji: an open-source platform for biological-image analysis. Nat Methods 9:676–682. https://doi.org/10.1038/nmeth.2019
Schapiro D, Sokolov A, Yapp C et al (2021) MCMICRO: a scalable, modular image-processing pipeline for multiplexed tissue imaging. Nat Methods 19:311–315. https://doi.org/10.1038/s41592-021-01308-y
McQuin C, Goodman A, Chernyshev V et al (2018) CellProfiler 3.0: next-generation image processing for biology. PLoS Biol 16:e2005970. https://doi.org/10.1371/journal.pbio.2005970
Stringer C, Wang T, Michaelos M, Pachitariu M (2021) Cellpose: a generalist algorithm for cellular segmentation. Nat Methods 18:100–106. https://doi.org/10.1038/s41592-020-01018-x
Bankhead P, Loughrey MB, Fernández JA et al (2017) QuPath: open source software for digital pathology image analysis. Sci Rep 7:16878. https://doi.org/10.1038/s41598-017-17204-5
Palla G, Spitzer H, Klein M et al (2022) Squidpy: a scalable framework for spatial omics analysis. Nat Methods 19:171–178. https://doi.org/10.1038/s41592-021-01358-2
Bannon D, Moen E, Schwartz M et al (2021) DeepCell Kiosk: scaling deep learning–enabled cellular image analysis with Kubernetes. Nat Methods 18:43–45. https://doi.org/10.1038/s41592-020-01023-0
Peng T, Thorn K, Schroeder T et al (2017) A BaSiC tool for background and shading correction of optical microscopy images. Nat Commun 8:14836. https://doi.org/10.1038/ncomms14836
BaSiC: a BaSiC tool for background and shading correction of optical microscopy images. Github. Available from: https://github.com/marrlab/BaSiC
BaSiCPy. Github. Available from: https://github.com/peng-lab/BaSiCPy
Chalfoun J, Majurski M, Blattner T et al (2017) MIST: accurate and scalable microscopy image stitching tool with stage modeling and error minimization. Sci Rep 7:1–10. https://doi.org/10.1038/s41598-017-04567-y
Muhlich JL, Chen Y-A, Yapp C et al (2022) Stitching and registering highly multiplexed whole-slide images of tissues and tumors using ASHLAR. Bioinformatics 38:4613–4621. https://doi.org/10.1093/bioinformatics/btac544
MIST: microscopy image stitching tool. Github. Available from: https://github.com/usnistgov/MIST
Fukai YT m2stitch: MIST-inspired microscope image stitching package. Github. Available from: https://github.com/yfukai/m2stitch
ashlar: ASHLAR: alignment by simultaneous harmonization of layer/adjacency registration. Github. Available from: https://github.com/labsyspharm/ashlar
Thévenaz P, Ruttimann UE, Unser M (1998) A pyramid approach to subpixel registration based on intensity. IEEE Trans Image Process 7:27–41. https://doi.org/10.1109/83.650848
StackReg. http://bigwww.epfl.ch/thevenaz/stackreg/. Accessed 2 May 2023
Lichtner G pystackreg: a python extension for the automatic alignment of a source image or a stack (movie) to a target image/reference frame. Github. Available from: https://github.com/glichtner/pystackreg
Johnson HJ, McCormick MM, Ibanez L (2015) The ITK software guide Book 2: design and functionality, 4th edn. Kitware, Incorporated
Yaniv Z, Lowekamp BC, Johnson HJ, Beare R (2018) SimpleITK image-analysis notebooks: a collaborative environment for education and reproducible research. J Digit Imaging 31:290–303. https://doi.org/10.1007/s10278-017-0037-8
Moen E, Bannon D, Kudo T et al (2019) Deep learning for cellular image analysis. Nat Methods 16:1233–1246. https://doi.org/10.1038/s41592-019-0403-1
Bilodeau A, Bouchard C, Lavoie-Cardinal F (2022) Automated microscopy image segmentation and analysis with machine learning. Methods Mol Biol 2440:349–365. https://doi.org/10.1007/978-1-0716-2051-9_20
Ravindran S (2022) Five ways deep learning has transformed image analysis. Nature 609:864–866. https://doi.org/10.1038/d41586-022-02964-6
Schmidt U, Weigert M, Broaddus C, Myers G (2018) Cell detection with star-convex polygons. In: Medical image computing and computer assisted intervention—MICCAI 2018. Springer International Publishing, pp 265–273
stardist: StarDist—object detection with star-convex shapes. Github. Available from: https://github.com/stardist/stardist
Schmidt U, Weigert M StarDist. In: ImageJ Wiki. https://imagej.net/plugins/stardist. Accessed 2 May 2023. Available from: https://imagej.net/plugins/stardist
qupath-extension-stardist: QuPath extension to run StarDist nucleus detection. Github. Available from: https://github.com/qupath/qupath-extension-stardist
cellpose: a generalist algorithm for cellular segmentation with human-in-the-loop capabilities. Github. Available from: https://github.com/MouseLand/cellpose
qupath-extension-cellpose: an extension that wraps a Cellpose environment such that WSI can be analyzed using Cellpose through QuPath. Github. Available from: https://github.com/BIOP/qupath-extension-cellpose
ijl-utilities-wrappers: wrappers for external software calls (Cellpose, Elastix, Ilastix...) and java utilities (object conversions and display). Github. Available from: https://github.com/BIOP/ijl-utilities-wrappers
deepcell-tf: deep learning library for single cell analysis. Github. Available from: https://github.com/vanvalenlab/deepcell-tf
kiosk-console: DeepCell kiosk distribution for kubernetes on GKE and AWS. Github. Available from: https://github.com/vanvalenlab/kiosk-console
Yang P, Huang H, Liu C (2021) Feature selection revisited in the single-cell era. Genome Biol 22:321. https://doi.org/10.1186/s13059-021-02544-3
Moon KR, van Dijk D, Wang Z et al (2019) Visualizing structure and transitions in high-dimensional biological data. Nat Biotechnol 37:1482–1492. https://doi.org/10.1038/s41587-019-0336-3
PHATE: PHATE (Potential of Heat-diffusion for Affinity-based Transition Embedding) is a tool for visualizing high dimensional data. Github. Available from: https://github.com/KrishnaswamyLab/PHATE
Pardee AB (1974) A restriction point for control of normal animal cell proliferation. Proc Natl Acad Sci U S A 71:1286–1290. https://doi.org/10.1073/pnas.71.4.1286
Zetterberg A, Larsson O (1985) Kinetic analysis of regulatory events in G1 leading to proliferation or quiescence of Swiss 3T3 cells. Proc Natl Acad Sci U S A 82:5365–5369. https://doi.org/10.1073/pnas.82.16.5365
BenchSci BenchSci: reimagining research. https://www.benchsci.com/. Accessed 8 May 2023
Kramer BA, Sarabia Del Castillo J, Pelkmans L (2022) Multimodal perception links cellular state to decision-making in single cells. Science 377:642–648. https://doi.org/10.1126/science.abf4062
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2024 The Author(s), under exclusive license to Springer Science+Business Media, LLC, part of Springer Nature
About this protocol
Cite this protocol
Kedziora, K.M., Stallaert, W. (2024). Cell Cycle Mapping Using Multiplexed Immunofluorescence. In: Castro, A., Lacroix, B. (eds) Cell Cycle Control. Methods in Molecular Biology, vol 2740. Humana, New York, NY. https://doi.org/10.1007/978-1-0716-3557-5_15
Download citation
DOI: https://doi.org/10.1007/978-1-0716-3557-5_15
Published:
Publisher Name: Humana, New York, NY
Print ISBN: 978-1-0716-3556-8
Online ISBN: 978-1-0716-3557-5
eBook Packages: Springer Protocols