Abstract
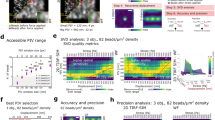
Immune cells rely on the generation of mechanical force to carry out their function. Consequently, there is a pressing need for quantitative methodologies that permit the probing of the spatio-temporal distribution of mechanical forces generated by immune cells. In this chapter, we provide a guide to quantify immune cell force generation using traction force microscopy (TFM), with a specific focus on its application to the study of the T-cell immunological synapse.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Huse M (2017) Mechanical forces in the immune system. Nat Rev Immunol 17:679–690. https://doi.org/10.1038/nri.2017.74
Pageon SV, Govendir MA, Kempe D, Biro M (2018) Mechanoimmunology: molecular-scale forces govern immune cell functions. MBoC 29:1919–1926. https://doi.org/10.1091/mbc.E18-02-0120
Basu R, Whitlock BM, Husson J, Le Floc’h A, Jin W, Oyler-Yaniv A, Dotiwala F, Giannone G, Hivroz C, Biais N, Lieberman J, Kam LC, Huse M (2016) Cytotoxic T cells use mechanical force to potentiate target cell killing. Cell 165:100–110. https://doi.org/10.1016/j.cell.2016.01.021
Li R, Ma C, Cai H, Chen W (2020) The CAR T-cell mechanoimmunology at a glance. Adv Sci 7:2002628. https://doi.org/10.1002/advs.202002628
de la Zerda A, Kratochvil MJ, Suhar NA, Heilshorn SC (2018) Review: bioengineering strategies to probe T cell mechanobiology. APL Bioeng 2:021501. https://doi.org/10.1063/1.5006599
Zancla A, Mozetic P, Orsini M, Forte G, Rainer A (2022) A primer to traction force microscopy. J Biol Chem 298:101867. https://doi.org/10.1016/j.jbc.2022.101867
Mustapha F, Sengupta K, Puech P-H (2022) May the force be with your (immune) cells: an introduction to traction force microscopy in immunology. Front Immunol 13:898558. https://doi.org/10.3389/fimmu.2022.898558
Mustapha F, Sengupta K, Puech P-H (2022) Protocol for measuring weak cellular traction forces using well-controlled ultra-soft polyacrylamide gels. STAR Protoc 3:101133. https://doi.org/10.1016/j.xpro.2022.101133
Vorselen D, Wang Y, de Jesus MM, Shah PK, Footer MJ, Huse M, Cai W, Theriot JA (2020) Microparticle traction force microscopy reveals subcellular force exertion patterns in immune cell–target interactions. Nat Commun 11:20. https://doi.org/10.1038/s41467-019-13804-z
Colin-York H, Shrestha D, Felce JH, Waithe D, Moeendarbary E, Davis SJ, Eggeling C, Fritzsche M (2016) Super-resolved traction force microscopy (STFM). Nano Lett 16:2633–2638. https://doi.org/10.1021/acs.nanolett.6b00273
Colin-York H, Javanmardi Y, Barbieri L, Li D, Korobchevskaya K, Guo Y, Hall C, Taylor A, Khuon S, Sheridan GK, Chew T-L, Li D, Moeendarbary E, Fritzsche M (2019) Spatiotemporally super-resolved volumetric traction force microscopy. Nano Lett 19:4427–4434. https://doi.org/10.1021/acs.nanolett.9b01196
Barbieri L, Colin-York H, Korobchevskaya K, Li D, Wolfson DL, Karedla N, Schneider F, Ahluwalia BS, Seternes T, Dalmo RA, Dustin ML, Li D, Fritzsche M (2021) Two-dimensional TIRF-SIM–traction force microscopy (2D TIRF-SIM-TFM). Nat Commun 12:2169. https://doi.org/10.1038/s41467-021-22377-9
Tse JR, Engler AJ (2010) Preparation of hydrogel substrates with tunable mechanical properties. Curr Protoc Cell Biol 47:10.16.1–10.16.16. https://doi.org/10.1002/0471143030.cb1016s47
Butler JP, Tolić-Nørrelykke IM, Fabry B, Fredberg JJ (2002) Traction fields, moments, and strain energy that cells exert on their surroundings. Am J Phys Cell Phys 282:C595–C605. https://doi.org/10.1152/ajpcell.00270.2001
Bauer A, Prechová M, Fischer L, Thievessen I, Gregor M, Fabry B (2021) pyTFM: a tool for traction force and monolayer stress microscopy. PLOS Comput Biol 17:e1008364. https://doi.org/10.1371/journal.pcbi.1008364
Tambe DT, Croutelle U, Trepat X, Park CY, Kim JH, Millet E, Butler JP, Fredberg JJ (2013) Monolayer stress microscopy: limitations, artifacts, and accuracy of recovered intercellular stresses. PLoS One 8:e55172. https://doi.org/10.1371/journal.pone.0055172
Han SJ, Oak Y, Groisman A, Danuser G (2015) Traction microscopy to identify force modulation in subresolution adhesions. Nat Methods 12:653–656. https://doi.org/10.1038/nmeth.3430
Saha K, Kim J, Irwin E, Yoon J, Momin F, Trujillo V, Schaffer DV, Healy KE, Hayward RC (2010) Surface creasing instability of soft polyacrylamide cell culture substrates. Biophys J 99:L94–L96. https://doi.org/10.1016/j.bpj.2010.09.045
Trujillo V, Kim J, C. Hayward R (2008) Creasing instability of surface-attached hydrogels. Soft Matter 4:564–569. https://doi.org/10.1039/B713263H
Acknowledgments
M.F., H.C.Y., and M.I. acknowledge generous funding from the Rosalind Franklin Institute, the Kennedy Trust for Rheumatology Research, the Wellcome Trust (212343/Z/18/Z), the EPSRC (EP/S004459/1), and the Oxford-ZEISS Centre of Excellence.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2023 The Author(s), under exclusive license to Springer Science+Business Media, LLC, part of Springer Nature
About this protocol
Cite this protocol
Issler, M., Colin-York, H., Fritzsche, M. (2023). Quantifying Immune Cell Force Generation Using Traction Force Microscopy. In: Baldari, C.T., Dustin, M.L. (eds) The Immune Synapse. Methods in Molecular Biology, vol 2654. Humana, New York, NY. https://doi.org/10.1007/978-1-0716-3135-5_23
Download citation
DOI: https://doi.org/10.1007/978-1-0716-3135-5_23
Published:
Publisher Name: Humana, New York, NY
Print ISBN: 978-1-0716-3134-8
Online ISBN: 978-1-0716-3135-5
eBook Packages: Springer Protocols