Abstract
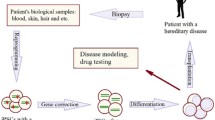
Insights into genome engineering in cells have allowed researchers to cultivate and modify cells as organoids that display structural and phenotypic features of human diseases or normal health status. The generation of targeted mutants is a crucial step toward studying the biomedical effect of genes of interest. Modified organoids derived from patients’ tissue cells are used as models to study diseases and test novel drugs. CRISPR-Cas9 technology has contributed to an explosion of advances that have the ability to edit genomes for the study of monogenic diseases and cancers. The generation of such mutants in human induced pluripotent stem cells (iPSCs) is of utmost importance as these cells carry the potential to be differentiated into any cell lineage. We describe recent developments that are broadening our understanding and extend DNA specificity, product selectivity, and fundamental capabilities. Furthermore, fundamental capabilities and remarkable advancements in basic research, biotechnology, and therapeutics development in cell engineering are detailed within this chapter. Using the CRISPR/Cas9 nuclease system for induction of targeted double-strand breaks, gene editing of target loci in iPSCs can be achieved with high efficiency. This chapter includes detailed protocols for the preparation of reagents to target loci of interest and transfection to genotype single cell-derived iPSC clones. Furthermore, we provide a protocol for the convenient generation of ribonucleoprotein (RNP) delivered directly to cells.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Zakrzewski W et al (2019) Stem cells: past, present, and future. Stem Cell Res Ther 10(1):68. https://doi.org/10.1186/s13287-019-1165-5
Volarevic V et al (2018) Ethical and safety issues of stem cell-based therapy. Int J Med Sci 15(1):36–45. https://doi.org/10.7150/ijms.21666
Wen W et al (2016) Enhanced generation of integration-free iPSCs from human adult peripheral blood mononuclear cells with an optimal combination of episomal vectors. Stem Cell Reports 6(6):873–884. https://doi.org/10.1016/j.stemcr.2016.04.005
Thomas KR, Capecchi MR (1987) Site-directed mutagenesis by gene targeting in mouse embryo-derived stem cells. Cell 51(3):503–512. https://doi.org/10.1016/0092-8674(87)90646-5
San Filippo J, Sung P, Klein H (2008) Mechanism of eukaryotic homologous recombination. Annu Rev Biochem 77:229–257. https://doi.org/10.1146/annurev.biochem.77.061306.125255
Capecchi MR (1989) Altering the genome by homologous recombination. Science 244(4910):1288–1292. https://doi.org/10.1126/science.2660260
Urnov FD et al (2010) Genome editing with engineered zinc finger nucleases. Nat Rev Genet 11(9):636–646. https://doi.org/10.1038/nrg2842
Joung JK, Sander JD (2013) TALENs: a widely applicable technology for targeted genome editing. Nat Rev Mol Cell Biol 14(1):49–55. https://doi.org/10.1038/nrm3486
Jinek M et al (2012) A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Science 337(6096):816–821. https://doi.org/10.1126/science.1225829
Cong L et al (2013) Multiplex genome engineering using CRISPR/Cas systems. Science 339(6121):819–823. https://doi.org/10.1126/science.1231143
Makarova KS et al (2015) An updated evolutionary classification of CRISPR-Cas systems. Nat Rev Microbiol 13(11):722–736. https://doi.org/10.1038/nrmicro3569
Lieber MR (2010) The mechanism of double-strand DNA break repair by the nonhomologous DNA end-joining pathway. Annu Rev Biochem 79:181–211. https://doi.org/10.1146/annurev.biochem.052308.093131
Wang H et al (2013) One-step generation of mice carrying mutations in multiple genes by CRISPR/Cas-mediated genome engineering. Cell 153(4):910–918. https://doi.org/10.1016/j.cell.2013.04.025
Hsu PD, Lander ES, Zhang F (2014) Development and applications of CRISPR-Cas9 for genome engineering. Cell 157(6):1262–1278. https://doi.org/10.1016/j.cell.2014.05.010
Jang HK et al (2020) Current trends in gene recovery mediated by the CRISPR-Cas system. Exp Mol Med 52(7):1016–1027. https://doi.org/10.1038/s12276-020-0466-1
Yamamoto Y, Gerbi SA (2018) Making ends meet: targeted integration of DNA fragments by genome editing. Chromosoma 127(4):405–420. https://doi.org/10.1007/s00412-018-0677-6
Paquet D et al (2016) Efficient introduction of specific homozygous and heterozygous mutations using CRISPR/Cas9. Nature 533(7601):125–129. https://doi.org/10.1038/nature17664
Zhang JP et al (2017) Efficient precise knockin with a double cut HDR donor after CRISPR/Cas9-mediated double-stranded DNA cleavage. Genome Biol 18(1):35. https://doi.org/10.1186/s13059-017-1164-8
Li XL et al (2018) Highly efficient genome editing via CRISPR-Cas9 in human pluripotent stem cells is achieved by transient BCL-XL overexpression. Nucleic Acids Res 46(19):10195–10215. https://doi.org/10.1093/nar/gky804
Yao X et al (2017) Homology-mediated end joining-based targeted integration using CRISPR/Cas9. Cell Res 27(6):801–814. https://doi.org/10.1038/cr.2017.76
Chen X et al (2017) In trans paired nicking triggers seamless genome editing without double-stranded DNA cutting. Nat Commun 8(1):657. https://doi.org/10.1038/s41467-017-00687-1
Hsu PD et al (2013) DNA targeting specificity of RNA-guided Cas9 nucleases. Nat Biotechnol 31(9):827–832. https://doi.org/10.1038/nbt.2647
Zou J et al (2009) Gene targeting of a disease-related gene in human induced pluripotent stem and embryonic stem cells. Cell Stem Cell 5(1):97–110. https://doi.org/10.1016/j.stem.2009.05.023
He X et al (2016) Knock-in of large reporter genes in human cells via CRISPR/Cas9-induced homology-dependent and independent DNA repair. Nucleic Acids Res 44(9):e85. https://doi.org/10.1093/nar/gkw064
Byrne SM et al (2015) Multi-kilobase homozygous targeted gene replacement in human induced pluripotent stem cells. Nucleic Acids Res 43(3):e21. https://doi.org/10.1093/nar/gku1246
Hayashi Y, Ohnuma K, Furue MK (2019) Pluripotent stem cell heterogeneity. Adv Exp Med Biol 1123:71–94. https://doi.org/10.1007/978-3-030-11096-3_6
De Los Angeles A et al (2015) Hallmarks of pluripotency. Nature 525(7570):469–478. https://doi.org/10.1038/nature15515
Montagnani S et al (2016) Adult stem cells in tissue maintenance and regeneration. Stem Cells Int 2016:7362879. https://doi.org/10.1155/2016/7362879
Kim EJ, Kang KH, Ju JH (2017) CRISPR-Cas9: a promising tool for gene editing on induced pluripotent stem cells. Korean J Intern Med 32(1):42–61. https://doi.org/10.3904/kjim.2016.198
Reisman M, Adams KT (2014) Stem cell therapy: a look at current research, regulations, and remaining hurdles. P T 39(12):846–857
Barman A, Deb B, Chakraborty S (2020) A glance at genome editing with CRISPR-Cas9 technology. Curr Genet 66(3):447–462. https://doi.org/10.1007/s00294-019-01040-3
Memi F, Ntokou A, Papangeli I (2018) CRISPR/Cas9 gene-editing: research technologies, clinical applications and ethical considerations. Semin Perinatol 42(8):487–500. https://doi.org/10.1053/j.semperi.2018.09.003
Shinwari ZK, Tanveer F, Khalil AT (2018) Ethical issues regarding CRISPR mediated genome editing. Curr Issues Mol Biol 26:103–110. https://doi.org/10.21775/cimb.026.103
Conflict of Interest
Stephen H. Tsang receives grant support from Abeona Therapeutics, Inc and Emendo. He is also the founder of Rejuvitas and is on the scientific and clinical advisory board for Nanoscope Therapeutics and Medical Excellence Capital.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2023 The Author(s), under exclusive license to Springer Science+Business Media, LLC, part of Springer Nature
About this protocol
Cite this protocol
Chang, YJ., Cui, X., Levi, S.R., Jenny, L.A., Tsang, S.H. (2023). CRISPR Manipulations in Stem Cell Lines. In: Tsang, S.H., Quinn, P.M. (eds) Retinitis Pigmentosa. Methods in Molecular Biology, vol 2560. Humana, New York, NY. https://doi.org/10.1007/978-1-0716-2651-1_23
Download citation
DOI: https://doi.org/10.1007/978-1-0716-2651-1_23
Published:
Publisher Name: Humana, New York, NY
Print ISBN: 978-1-0716-2650-4
Online ISBN: 978-1-0716-2651-1
eBook Packages: Springer Protocols