Abstract
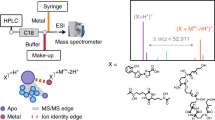
The roles of small molecules in every aspect of life have been gaining increased recognition. Many are known to exert their effect by binding proteins—but a comprehensive overview of protein–metabolite interactions (PMIs) is missing. Recently we devised a non-targeted method for detecting PMIs using size-exclusion chromatography followed by proteomic and metabolomic analysis: PROMIS. Under test this method was able to identify known PMIs such as enzyme–cofactor complexes as well as novel ones.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Veyel D, Kierszniowska S, Kosmacz M et al (2017) System-wide detection of protein-small molecule complexes suggests extensive metabolite regulation in plants. Sci Rep 7. https://doi.org/10.1038/srep42387
Veyel D, Sokolowska EM, Moreno JC et al (2018) PROMIS, global analysis of PROtein-metabolite interactions using size separation in Arabidopsis thaliana. J Biol Chem 293:12440–12453
Luzarowski M, Vicente R, Kiselev A et al (2021) Global mapping of protein–metabolite interactions in Saccharomyces cerevisiae reveals that Ser-Leu dipeptide regulates phosphoglycerate kinase activity. Commun Biol 4. https://doi.org/10.1038/s42003-021-01684-3
Luzarowski M, Skirycz A (2019) Emerging strategies for the identification of protein-metabolite interactions. J Exp Bot. https://doi.org/10.1093/jxb/erz228
Luzarowski M, Wojciechowska I, Skirycz A (2018) 2 in 1: one-step affinity purification for the parallel analysis of protein-protein and protein-metabolite complexes. J Vis Exp 2018. https://doi.org/10.3791/57720
Luzarowski M, Kosmacz M, Sokolowska E et al (2017) Affinity purification with metabolomic and proteomic analysis unravels diverse roles of nucleoside diphosphate kinases. J Exp Bot 68. https://doi.org/10.1093/jxb/erx183
Maeda K, Anand K, Chiapparino A et al (2013) Interactome map uncovers phosphatidylserine transport by oxysterol-binding proteins. Nature 501:257–261
Piazza I, Kochanowski K, Cappelletti V et al (2018) A map of protein-metabolite interactions reveals principles of chemical communication. Cell 172:358–372
Savitski MM, Reinhard FBM, Franken H et al (2014) Tracking cancer drugs in living cells by thermal profiling of the proteome. Science 346:1255784
Diether M, Nikolaev Y, Allain FHT, Sauer U (2019) Systematic mapping of protein-metabolite interactions in central metabolism of Escherichia coli. Mol Syst Biol 15:e9008
Gallego O, Betts MJ, Gvozdenovic-Jeremic J et al (2010) A systematic screen for protein–lipid interactions in Saccharomyces cerevisiae. Mol Syst Biol 6:430
Li X, Gianoulis TA, Yip KY et al (2010) Extensive in vivo metabolite-protein interactions revealed by large-scale systematic analyses. Cell 143:639–650
Giavalisco P, Li Y, Matthes A et al (2011) Elemental formula annotation of polar and lipophilic metabolites using 13C, 15N and 34S isotope labelling, in combination with high-resolution mass spectrometry. Plant J 68:364–376
Cox J, Mann M (2008) MaxQuant enables high peptide identification rates, individualized ppb-range mass accuracies and proteome-wide protein quantification. Nat Biotechnol 26:1367–1372
Tyanova S, Temu T, Cox J (2016) The MaxQuant computational platform for mass spectrometry-based shotgun proteomics. Nat Protoc 11:2301
Cox J, Neuhauser N, Michalski A et al (2011) Andromeda: a peptide search engine integrated into the MaxQuant environment. J Proteome Res 10:1794–1805
Acknowledgments
We thank Hezi Tenenboim for his excellent help with English editing.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2023 The Author(s), under exclusive license to Springer Science+Business Media, LLC, part of Springer Nature
About this protocol
Cite this protocol
Luzarowski, M., Sokolowska, E.M., Schlossarek, D., Skirycz, A. (2023). PROMIS: Co-fractionation Mass Spectrometry for Analysis of Protein–Metabolite Interactions. In: Skirycz, A., Luzarowski, M., Ewald, J.C. (eds) Cell-Wide Identification of Metabolite-Protein Interactions. Methods in Molecular Biology, vol 2554. Humana, New York, NY. https://doi.org/10.1007/978-1-0716-2624-5_10
Download citation
DOI: https://doi.org/10.1007/978-1-0716-2624-5_10
Published:
Publisher Name: Humana, New York, NY
Print ISBN: 978-1-0716-2623-8
Online ISBN: 978-1-0716-2624-5
eBook Packages: Springer Protocols