Abstract
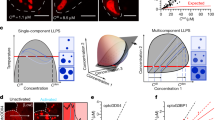
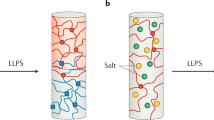
Phase separation is one of the fundamental processes to compartmentalize biomolecules in living cells. RNA–protein complexes (RNPs) often scaffold biomolecular condensates formed through phase separation. We here present a statistical thermodynamics approach to investigate intracellular phase separation. We first present the statistical thermodynamic theory of the liquid-liquid phase separation (LLPS) of two molecules (such as proteins and solvent molecules) and of a polymer solution (such as RNPs and solvent molecules). Condensates produced by LLPS show coarsening and/or coalescence to minimize their total surface area. In addition to the LLPS, there are other types of self-assembly, such as microphase separation, micellization, emulsification, and vesiculation, with which the growth of the assembly stops with optimal size and shape. We also describe a scaling theory of micelles of block copolymers, where their structures are analogous to the core-shell structure of paraspeckle nuclear bodies scaffolded by RNPs of NEAT1_2 long noncoding RNAs (lncRNAs) and RNA-binding proteins (RBPs). These theories treat the self-assembly of polymers in the thermodynamic equilibrium, where their concentrations and compositions do not change with time. In contrast, RNPs are produced according to the transcription of RNAs and are degraded with time. We therefore take into account the dynamical aspect of the production of RNPs in an extension of the theory of the self-assembly of soft matter. Finally, we discuss the structure of paraspeckles as an example to demonstrate that an approach combining experiment and theory is powerful to investigate the mechanism of intracellular phase separation.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Banani SF et al (2017) Biomolecular condensates: organizers of cellular biochemistry. Nat Rev Mol Cell Biol 18(5):285–298
Schmidt HB, Gorlich D (2016) Transport selectivity of nuclear pores, phase separation, and Membraneless organelles. Trends Biochem Sci 41(1):46–61
Fujioka Y et al (2020) Phase separation organizes the site of autophagosome formation. Nature 578(7794):301–305
Spannl S et al (2019) Biomolecular condensates in neurodegeneration and cancer. Traffic 20(12):890–911
Shin Y, Brangwynne CP (2017) Liquid phase condensation in cell physiology and disease. Science 357(6357):eaaf4382
Mathieu C, Pappu RV, Taylor JP (2020) Beyond aggregation: pathological phase transitions in neurodegenerative disease. Science 370(6512):56–60
Zbinden A et al (2020) Phase separation and neurodegenerative diseases: a disturbance in the force. Dev Cell 55(1):45–68
Chujo T, Yamazaki T, Hirose T (2016) Architectural RNAs (arcRNAs): a class of long noncoding RNAs that function as the scaffold of nuclear bodies. Biochim Biophys Acta 1859(1):139–146
Maharana S et al (2018) RNA buffers the phase separation behavior of prion-like RNA binding proteins. Science 360(6391):918–921
Van Treeck B, Parker R (2018) Emerging roles for intermolecular RNA-RNA interactions in RNP assemblies. Cell 174(4):791–802
Yamazaki T, Nakagawa S, Hirose T (2020) Architectural RNAs for Membraneless nuclear body formation. Cold Spring Harb Symp Quant Biol 84:227–237
Ninomiya K, Hirose T (2020) Short tandem repeat-enriched architectural RNAs in nuclear bodies: functions and associated diseases. Noncoding RNA 6(1):6
Chujo T et al (2017) Unusual semi-extractability as a hallmark of nuclear body-associated architectural noncoding RNAs. EMBO J 36(10):1447–1462
Yap K et al (2018) A short tandem repeat-enriched RNA assembles a nuclear compartment to control alternative splicing and promote cell survival. Mol Cell 72(3):525–540.e13
Safran SA (2003) Statistical thermodynamics of surfaces, interfaces, and membranes. Westview Press, CO., Routledge
Doi M (2013) Soft matter physics. Oxford Univ. Press, Oxford
Doi M (1996) Introduction to polymer physics. Oxford Univ. Press, Oxford
de Gennes PG (1979) Scaling concepts in polymer physics. Cornell Univ. Press, New York
Rubinstein M, Colby R (2003) Polymer physics. Oxford Univ. Press, New York
Yamazaki T, Yamamoto T, Yoshino H, Souquere S, Nakagawa S, Pierron G, Hirose T (2021) Paraspeckles are constructed as block copolymer micelles. EMBO J 40(12):e107270. https://doi.org/10.15252/embj.2020107270
Yamamoto T, Yamazaki T, Hirose T (2020) Triblock copolymer micelle model of spherical paraspeckles. bioRxiv. https://doi.org/10.1101/2020.11.01.364190
Halperin A, Alexander S (1989) Polymer micelles: their relaxation kinetics. Macromolecules 22:2403–2412
Zhulina EB, Adam M, LaRue I, Sheiko SS, Rubinstein M (2005) Diblock copolymer micelles in a dilute solution. Macromolecules 38:5330–5535
Yamamoto T, Yamazaki T, Hirose T (2020) Phase separation driven by production of architectural RNA transcripts. Soft Matter 16(19):4692–4698
Sasaki YT et al (2009) MENepsilon/beta noncoding RNAs are essential for structural integrity of nuclear paraspeckles. Proc Natl Acad Sci U S A 106(8):2525–2530
Sunwoo H et al (2009) MEN epsilon/beta nuclear-retained non-coding RNAs are up-regulated upon muscle differentiation and are essential components of paraspeckles. Genome Res 19(3):347–359
Chen LL, Carmichael GG (2009) Altered nuclear retention of mRNAs containing inverted repeats in human embryonic stem cells: functional role of a nuclear noncoding RNA. Mol Cell 35(4):467–478
Clemson CM et al (2009) An architectural role for a nuclear noncoding RNA: NEAT1 RNA is essential for the structure of paraspeckles. Mol Cell 33(6):717–726
Yamazaki T et al (2018) Functional domains of NEAT1 architectural lncRNA induce Paraspeckle assembly through phase separation. Mol Cell 70(6):1038–1053 e7
Mao YS et al (2011) Direct visualization of the co-transcriptional assembly of a nuclear body by noncoding RNAs. Nat Cell Biol 13(1):95–101
Naganuma T et al (2012) Alternative 3′-end processing of long noncoding RNA initiates construction of nuclear paraspeckles. EMBO J 31(20):4020–4034
Souquere S et al (2010) Highly ordered spatial organization of the structural long noncoding NEAT1 RNAs within paraspeckle nuclear bodies. Mol Biol Cell 21(22):4020–4027
West JA et al (2016) Structural, super-resolution microscopy analysis of paraspeckle nuclear body organization. J Cell Biol 214(7):817–830
Souquere S, Pierron G (2015) Ultrastructural analysis of nuclear bodies using electron microscopy. Methods Mol Biol 1262:105–118
Mito M et al (2016) Simultaneous multicolor detection of RNA and proteins using super-resolution microscopy. Methods 98:158–165
Yamazaki T, Hirose T (2021) CRISPR-mediated mutagenesis of long noncoding RNAs. Methods Mol Biol 2254:283–303
Acknowledgments
This research was supported by KAKENHI grants from the Ministry of Education, Culture, Sports, Science, and Technology (MEXT) of Japan [to T. Yamazaki (19K06479, 19H05250, 21H00253), T. Yamamoto (18K03558, 19H05259, 20H05934, 21K03479, 21H00241)], JST, PRESTO Grant Number JPMJPR18KA (to T. Yamamoto), the Mochida Memorial Foundation for Medical and Pharmaceutical Research (to T. Yamazaki), the Naito Foundation (to T. Yamazaki), and the Takeda Science Foundation (to T. Yamazaki). T. Yamamoto acknowledges S. A. Safran (Weizmann Institute of Science) and Takahiro Sakaue (Aoyama Gakuin University) for their critical reading of the manuscript.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2022 The Author(s), under exclusive license to Springer Science+Business Media, LLC, part of Springer Nature
About this protocol
Cite this protocol
Yamazaki, T., Yamamoto, T. (2022). Statistical Thermodynamics Approach for Intracellular Phase Separation. In: Parrish, N.F., Iwasaki, Y.W. (eds) piRNA. Methods in Molecular Biology, vol 2509. Humana, New York, NY. https://doi.org/10.1007/978-1-0716-2380-0_22
Download citation
DOI: https://doi.org/10.1007/978-1-0716-2380-0_22
Published:
Publisher Name: Humana, New York, NY
Print ISBN: 978-1-0716-2379-4
Online ISBN: 978-1-0716-2380-0
eBook Packages: Springer Protocols