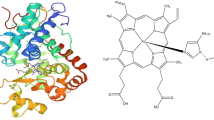
Abstract
Functional analyses of peroxidases are a major challenge. In silico analysis appears to be a powerful tool to overcome at least some of the problems that arose from (1) the numerous possible functions of peroxidases, (2) their low substrate specificity, and (3) the compensation of knockout mutants by other isoenzymes. Amino acid sequences and crystal structures of peroxidases were used for the prediction of tertiary structures, posttranslational modifications, ligand and substrate binding sites, and so on of uncharacterized peroxidases. This protocol presents tools and their applications for an in silico analysis of soluble and membrane-bound peroxidases, but it may be used for other proteins, too.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Welinder KG, Justesen AF, Kjaersgard IV et al (2002) Structural diversity and transcription of class III peroxidases from Arabidopsis thaliana. Eur J Biochem 269:6063–6081
Zámocký M, Furtmüller PG, Obinger C (2010) Evolution of structure and function of class I peroxidases. Arch Biochem Biophys 500:45–57
Hiraga S, Sasaki K, Ito H et al (2001) A large family of class III plant peroxidases. Plant Cell Physiol 42:462–468
Passardi F, Cosio C, Penel C et al (2005) Peroxidases have more functions than a Swiss army knife. Plant Cell Rep 24:255–265
Cosio C, Dunand C (2009) Specific functions of individual class III peroxidase genes. J Exp Bot 60:391–409
Lüthje S, Meisrimler CN, Hopff D et al (2011) Phylogeny, topology, structure and functions of membrane-bound class III peroxidases in vascular plants. Phytochemistry 72:1124–1135
Herrero J, Esteban-Carrasco A, Zapata JM (2013a) Looking for Arabidopsis thaliana peroxidases involved in lignin biosynthesis. Plant Physiol Biochem 67:77–86
Herrero J, Fernández-Pérez F, Yebra T et al (2013b) Bioinformatic and functional characterization of the basic peroxidase 72 from Arabidopsis thaliana involved in lignin biosynthesis. Planta 237:1599–1612
Shigeto J, Nagano M, Fujita K et al (2014) Catalytic profile of Arabidopsis peroxidases, AtPrx-2, 25 and 71, contributing to stem lignification. PLoS One 9:e105332
Lüthje S, Martinez-Cortes T (2018) Membrane-bound class III peroxidases: unexpected enzymes with exciting functions. Int J Mol Sci 19:E2876
Artimo P, Jonnalagedda M, Arnold K et al (2012) ExPASy: SIB bioinformatics resource portal. Nucleic Acids Res 40:W597–W603
UniProt Consortium T (2018) UniProt: the universal protein knowledgebase. Nucleic Acids Res 46:2699
Sayers EW, Agarwala R, Bolton EE et al (2019) Database resources of the National Center for Biotechnology Information. Nucleic Acids Res 47:D23–D28
Schwacke R, Schneider A, Van Der Graaff E et al (2003) ARAMEMNON, a novel database for Arabidopsis integral membrane proteins. Plant Physiol 131:16–26
Fawal N, Li Q, Savelli B et al (2013) PeroxiBase: a database for large-scale evolutionary analysis of peroxidases. Nucleic Acids Res 41:D441–D444
Berman HM, Westbrook J, Feng Z et al (2000) The Protein Data Bank. Nucleic Acids Res 28:235–242
Lipman DJ, Pearson WR (1985) Rapid and sensitive protein similarity searches. Science 227:1435–1441
Gasteiger E, Hoogland C, Gattiker A et al (2005) Protein identification and analysis tools on the ExPASy server. In: Walker JM (ed) The proteomics protocols handbook. Humana Press, New York, pp 571–607
Ren J, Wen L, Gao X et al (2008) CSS-Palm 2.0: an updated software for palmitoylation sites prediction. Protein Eng Des Sel 21:639–644
Sigrist CJA, de Castro E, Cerutti L et al (2012) New and continuing developments at PROSITE. Nucleic Acids Res 21:D344–D347
Blom N, Sicheritz-Ponten T, Gupta R et al (2004) Prediction of post-translational glycosylation and phosphorylation of proteins from the amino acid sequence. Proteomics 4:1633–1649
Fankhauser N, Mäser P (2005) Identification of GPI anchor attachment signals by a Kohonen self-organizing map. Bioinformatics 21:1846–1852
Krogh A, Larsson B, von Heijne G et al (2001) Predicting transmembrane protein topology with a hidden Markov model: application to complete genomes. J Mol Biol 305:567–580
Tusnády GE, Simon I (2001) The HMMTOP transmembrane topology prediction server. Bioinformatics 17:849–850
Möller S, Croning MDR, Apweiler R (2001) Evaluation of methods for the prediction of membrane spanning regions. Bioinformatics 17:646–653
Nielsen H, Krogh A (1998) Prediction of signal peptides and signal anchors by a hidden Markov model. Proc Int Conf Intell Syst Mol Biol 6:122–130
Nakai K, Horton P (1999) PSORT: a program for detecting the sorting signals of proteins and predicting their subcellular localization. Trends Biochem Sci 24:34–35
Waterhouse A, Bertoni M, Bienert S et al (2018) SWISS-MODEL: homology modelling of protein structures and complexes. Nucleic Acids Res 46:W296–W303
Kelley LA, Mezulis S, Yates CM et al (2015) The Phyre2 web portal for protein modeling, prediction and analysis. Nat Protoc 10:845–858
Wass MN, Kelley LA, Sternberg MJ (2010) 3DLigandSite: predicting ligand-binding sites using similar structures. Nucleic Acids Res 38(Suppl):W469–W473
Krissinel E, Henrick K (2007) Inference of macromolecular assemblies from crystalline state. J Mol Biol 372:774–797
Moural TW, Lewis KM, Barnaba C et al (2017) Characterization of class III peroxidases from Switchgrass. Plant Physiol 173:417–433
Pettersen EF, Goddard TD, Huang CC et al (2004) UCSF chimera a visualization system for exploratory research and analysis. J Comput Chem 25:1605–1612
Grosdidier A, Zoete V, Michielin O (2011) SwissDock, a protein-small molecule docking web service based on EADock DSS. Nucleic Acids Res 39:W270–W277
Morris GM, Huey R, Lindstrom W et al (2009) Autodock4 and AutoDockTools4: automated docking with selective receptor flexiblity. J Comput Chem 16:2785–2791
Seeliger D, de Groot BL (2010) Ligand docking and binding site analysis with PyMOL and autodock/Vina. J Comput Aided Mol Des 24:417–422
Nanda T, Tripathy K, Ashwin P (2011) Integration of Bioinformatics Tools for Proteomics Research. J Comput Sci Syst Biol S13. https://doi.org/10.4172/jcsb.S13-002
Hetenyi C, van der Spoel D (2011) Toward prediction of functional protein pockets using blind docking and pocket search algorithms. Protein Sci 20:880–893
Andrusier N, Nussinov R, Wolfson HJ (2007) FireDock: fast interaction refinement in molecular docking. Proteins 69:139–159
Mashiach E, Schneidman-Duhovny D, Andrusier N et al (2008) FireDock: a web server for fast interaction refinement in molecular docking. Nucleic Acids Res 36:W229–W232
Schneidman-Duhovny D, Inbar Y, Nussinov R et al (2005) PatchDock and SymmDock: servers for rigid and symmetric docking. Nucleic Acids Res 33:W363–W367
Irwin JJ, Sterling T, Mysinger MM et al (2012) ZINC: a free tool to discover chemistry for biology. J Chem Inf Model 52:1757–1768
Lee Y, Rubio MC, Alassimone J et al (2013) A mechanism for localized lignin deposition in the endodermis. Cell 153:402–412
Watanabe L, de Moura PR, Bleicher L et al (2010) Crystal structure and statistical coupling analysis of highly glycosylated peroxidase from royal palm tree (Roystonea regia). J Struct Biol 169:226–242
Berglund GI, Carlsson GH, Smith AT et al (2002) The catalytic pathway of horseradish peroxidase at high resolution. Nature 417:463
Acknowledgments
This work was supported by a PhD student grant to K. R. from the Dr. Elisabeth Appuhn Foundation.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2020 Springer Science+Business Media, LLC, part of Springer Nature
About this protocol
Cite this protocol
Lüthje, S., Ramanathan, K. (2020). In Silico Analysis of Class III Peroxidases: Hypothetical Structure, Ligand Binding Sites, Posttranslational Modifications, and Interaction with Substrates. In: Jorrin-Novo, J., Valledor, L., Castillejo, M., Rey, MD. (eds) Plant Proteomics. Methods in Molecular Biology, vol 2139. Humana, New York, NY. https://doi.org/10.1007/978-1-0716-0528-8_24
Download citation
DOI: https://doi.org/10.1007/978-1-0716-0528-8_24
Published:
Publisher Name: Humana, New York, NY
Print ISBN: 978-1-0716-0527-1
Online ISBN: 978-1-0716-0528-8
eBook Packages: Springer Protocols