Abstract
Influenza A viruses (IAV) are members of the Orthomyxoviridae family of negative-sense RNA viruses. The greatest diversity of IAV strains is found in aquatic birds, but a subset of strains infects other avian as well as mammalian species, including humans. In aquatic birds, infection is largely restricted to the gastrointestinal tract and spread is through feces, while in humans and other mammals, respiratory epithelial cells are the primary sites supporting productive replication and transmission. IAV triggers the death of most cell types in which it replicates, both in culture and in vivo. When well controlled, such cell death is considered an effective host defense mechanism that eliminates infected cells and limits virus spread. Unchecked or inopportune cell death also results in immunopathology. In this chapter, we discuss the impact of cell death in restricting virus spread, supporting the adaptive immune response and driving pathogenesis in the mammalian respiratory tract. Recent studies have begun to shed light on the signaling pathways underlying IAV-activated cell death. These pathways, initiated by the pathogen sensor protein ZBP1 (also called DAI and DLM1), cause infected cells to undergo apoptosis, necroptosis, and pyroptosis. We outline mechanisms of ZBP1-mediated cell death signaling following IAV infection.
You have full access to this open access chapter, Download chapter PDF
Similar content being viewed by others
Keywords
1 Introduction
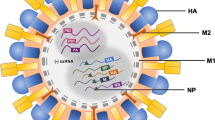
Influenza A virus (IAV) is a segmented negative-sense RNA virus of the family Orthomyxoviridae. Eight genome segments are packaged in ribonucleoprotein (RNP) complexes to form the capsid that is enclosed within a lipid bilayer envelope derived from host cell membranes. The virion envelope contains two virus-encoded glycoproteins, hemagglutinin (HA or H), and neuraminidase (NA or N). Eighteen HA and eleven NA variants have been serologically distinguished and used to identify IAV subtypes (e.g., H1N1, H5N1, and H7N9). IAV is unique in that these subtypes undergo antigenic drift over time. In addition, co-infection of cells with two different subtypes can result in exchange of genome segments between subtypes and generation of reassortants bearing new combinations of genome segments. Such reassortment can result in antigenic shift. IAV enjoys a wide species distribution, with the broadest subset of viral strains infecting aquatic birds, the primary reservoir of IAV diversity. In aquatic birds, gastrointestinal infection predominates, with few, if any, symptoms occurring during infection with common subtypes. Transmission to other avian species, such as poultry, often results in host adaptation and specialization, including replication in the respiratory tract, producing new strains that lose their ability to infect or transmit within the original avian species. This partitioning creates strains that become adapted to specific hosts (Yoon et al. 2014).
Fewer IAV subtypes infect mammals than avian species; however, these also display the same extreme partitioning, such that canine, equine, feline, porcine, and human viruses are characterized by distinct subtypes that each exhibits reduced fitness in other species (Yoon et al. 2014). This species restriction results from the presence in respiratory epithelium of enzymes that are capable of HA cleavage and activation (Bottcher-Friebertshauser et al. 2013). HA mediates virus binding to specific sialic acid moieties on the surface of target cells, leading to receptor-mediated endocytosis (Lakadamyali et al. 2004). Entry is completed from within endosomes, where acidification triggers a conformational change in HA (this is the step that requires HA to be appropriately cleaved), resulting in fusion of the virion envelope with the endosomal membrane and the deposition of viral RNPs into the cell cytoplasm. From here, RNPs are trafficked to the nucleus, where the viral RNA-dependent RNA-polymerase complex produces positive-sense RNAs that not only serve as mRNA precursors for synthesis of new virus proteins, but also as template RNAs for production of negative-sense viral genomes (te Velthuis and Fodor 2016).
The newly produced viral proteins and genomes assemble into progeny virions at the plasma membrane (Rossman and Lamb 2011). As the nascent virions bud from the cell, the envelope NA protein cleaves sialic acids to which the budding virus is bound, releasing free virions. The frontline anti-IAV drug oseltamivir is an NA inhibitor that arrests virus budding prior to release from the cell surface (Air 2012). Either inside the respiratory epithelial cell or on the cell surface, host proteases carry out HA cleavage/activation, thereby rendering progeny virus competent for successive rounds of infection.
In humans and other mammals, IAV infects epithelial cells lining the respiratory tract, from the nasal passages to the soft palette and the lung airways down to the alveolar epithelial cells (AECs) (Sanders et al. 2011). Transmission occurs primarily as a result of viral replication in the upper respiratory tract, particularly the soft palette (Lakdawala et al. 2015). Severe disease, in contrast, is often associated with infection in the lower respiratory tract, such that the extent of lung involvement correlates with disease outcomes in both humans and animals (Sanders et al. 2013; Soto-Abraham et al. 2009; Beigel et al. 2005; Watanabe et al. 2011; Belser et al. 2010; Shieh et al. 2010; Fujita et al. 2014). Type I and type II AECs are the primary replicative niches for IAV in mammalian lungs, although this depends on both the virus strain and the host species (Cardani et al. 2017; Sanders et al. 2013; Weinheimer et al. 2012; Rosenberger et al. 2014). Goblet cells, club cells, and ciliated cells are also capable of being infected by IAV (Heaton et al. 2014; Edinger et al. 2014). A schematic representation of the lower respiratory tract, showing terminal bronchioles, alveoli, and their primary structural cell types, is shown in Fig. 1. Sialic acid moieties on glycoproteins on the surface of these respiratory cell types dictate their capacity to become infected by distinct strains of IAV. Among mammals, pigs and ferrets share similar patterns of sialic acid distribution to humans, making them appropriate models for studies of IAV transmission (Kuiken et al. 2010; Thangavel and Bouvier 2014; and Wasik et al. 2017). In contrast, IAV does not transmit efficiently in the mouse and typically requires some adaptation for efficient replication. Despite these limitations, immune response parameters and pathogenicity profiles in this model are largely consistent with human disease (Samet and Tompkins 2017). Moreover, the vast repertoire of genetic and immunological tools available in the mouse makes it by far the most extensively used model for IAV studies in vivo.
Structural cell types of the lower respiratory tract. Terminal bronchioles comprise of a mix of ciliated cuboidal epithelial cells and non-ciliated club cells, which are secretory cells that protect, detoxify, and repair airways. These bronchioles divide into alveolar ducts that terminate in alveoli, comprised primarily of type I and type II AECs. Type I AECs are very thin epithelial cells constituting the majority (>95%) of the alveolar epithelium and are responsible for gas exchange during respiration. Type II AECs make up the rest of the alveolar structure and are interspersed among type I AECs, typically at junctions of alveolar septal walls. These secretory cells produce surfactant and are required for maintenance of alveolar structure and integrity
In this chapter, we will discuss the mechanisms of cell death in the respiratory tract that occur during mammalian IAV infections, with a focus on contributions of cell death to disease outcome.
2 Role of Cell Death in IAV Immunity and Pathogenesis
2.1 Cell Death in Innate Control of Initial Replication and Spread
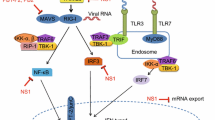
Laboratory studies of IAV infections in naïve mice lacking any adaptive immune memory toward IAV reveal that peak viral titers (in the whole lung) occur three to five days after infection (Fig. 2). Virus titers decrease after this time, despite a lack of any detectable adaptive immune response to IAV. Indeed, the earliest emergence of adaptive immune effectors in the lung of naïve animals, including CD4+ T cells, CD8+ T cells, and antiviral antibodies, takes place around five days after infection (Doherty et al. 2006). Even in immune-deficient Rag2−/− mice, a slight but measurable reduction in virus burden is apparent following peak titers three days after infection, although these mice never resolve the infection and eventually succumb to IAV (Wu et al. 2010). The initial control of virus load is, in large part, mediated by type I (predominantly α/β) and type III (λ) interferons (IFNs). These cytokines not only reduce the amount of virus produced per infected cell, but together limit spread of infection to neighboring, uninfected cells (Mordstein et al. 2008, 2010). Type I and III IFNs are chiefly produced by RIG-I-like receptor (RLR) and Toll-like receptor (TLR)-dependent signaling pathways, and exert their impact via the induction of hundreds of IFN-stimulated genes (ISGs). Among these ISGs are those encoding key components of the host cell death response to IAV, including ZBP1, MLKL, and TRAIL (discussed in more detail later), indicating that promoting programmed cell death of the infected cell is a key mechanism by which IFNs exert their antiviral properties. Indeed, mathematical simulation (i.e., so-called target cell limited models) of human and mouse IAV infection requires simple elimination of available target cells to account for in vivo virus titer data, without need to invoke other innate or adaptive antiviral clearance mechanisms (Baccam et al. 2006). Thus, cell death likely constitutes the dominant mechanism by which innate immune host defense defines the peak carrying capacity of virus titer, limits virus spread, and brings IAV infection under control.
Virus and host response dynamics during influenza infection. Influenza infection proceeds in infected naïve mice and humans through three phases of host responses, beginning with the early stages of virus growth and the ensuing innate immune response that initially controls the infection (Phase 1). Viral titer peaks around 3–5 days after infection depending on the virus strain and host, at which point adaptive immune effectors, including virus-specific CD8+ T cells and antibodies start to accumulate in the infected lung (Phase 2). Although these effectors first appear relatively early (starting around day 5), their accumulation is required to facilitate the clearance of the virus, which is usually complete by two weeks after infection. Effectors linger through the recovery phase, at which point titratable virus is undetectable but some viral antigen may remain (Phase 3). This phase is characterized by lung remodeling and repair in an effort to restore lung function to pre-infection levels
Although programmed cell death mechanisms restrict virus spread within the lung airways, the death of epithelium is also a dominant driver of IAV infection-associated morbidity and mortality (Davidson et al. 2014). In particular, the loss of type I AECs (which are essential for gas exchange) above a threshold level of ~10% correlates with mortality of IAV-infected mice (Sanders et al. 2013). In humans, lesions in the lower respiratory tract are a consistent finding in autopsies of fatal cases of the 2009 pandemic H1N1 virus (Mauad et al. 2010; Shieh et al. 2010). Dying cells can overwhelm cell clearance and tissue repair mechanisms, and reduce pulmonary function by promoting the recruitment of inflammatory cell types (e.g., monocytes, neutrophils) and by causing the leakage of fluid (edema) into the lung. Indeed, the generalized inflammatory environment triggered by cytokine production from infected epithelial cells and responding professional inflammatory cells correlates with the severity of lung damage. More virulent, rapidly growing influenza strains, such as avian H5N1 and H7N9 subtypes associated with epizootic infections of humans, are more likely to cause such “cytokine storms”. Variations have been noted between subtypes, with H5N1 inducing higher levels of inflammatory cytokines than H7N9 (Meliopoulos et al. 2014; de Wit et al. 2014; Chan et al. 2013). The reconstructed 1918 pandemic H1N1 virus also displayed a pattern of rapid growth in macaques, accompanied by elevated levels of cytokines, severe inflammation, and extensive alveolar cell death (Kobasa et al. 2007). This rapid growth was dependent on both HA and polymerase complex function (Watanabe et al. 2013). Despite several features of exacerbated inflammation in infected animals, the type I IFN response to the 1918 H1N1 strain was remarkably blunted, compared to a conventional H1 IAV infection, suggesting that the severe outcomes observed in experimental animals and seen during the worldwide pandemic caused by this virus may have resulted from a profoundly dysregulated immune signature that drove excess alveolar cell death. Thus, when metered appropriately, cell death represents a host defense mechanism that limits virus spread as well as immunopathology early in an infection. But when uncontrolled (e.g., during infection with highly-virulent strains of IAV), severe damage to airway epithelia and consequent host mortality may occur, despite resolution of infection. Such severe pathology is observed in mouse models, where destruction of airway epithelium is a common feature of lethal IAV infection (Sanders et al. 2011, 2013; Kash et al. 2006; Brandes et al. 2013; Hogner et al. 2013), and in humans, where death of lower pulmonary tract epithelium, marked by areas of bronchioalveolar necrosis, is a classic feature of IAV-induced acute respiratory distress syndrome (ARDS) (Korteweg and Gu 2008; Mauad et al. 2010).
2.2 Cell Death During the Adaptive Immune Response to IAV
The earliest cell-mediated immune responses against IAV are NK cell-dependent which, through the production of IFN-γ and other cytokines, recruit additional inflammatory cells to sites of infection (Zamora et al. 2017) (Fig. 2). NK cells can also directly kill infected cells via secretion of toxic granules (Jost and Altfeld 2013). The adaptive immune response to IAV is largely driven by surface glycoprotein-directed antibodies (e.g., HA, NA) that mediate clearance of virus and dictate immune memory. Virus-specific CD8+ T cells also contribute to the elimination of infected epithelial cells and the ultimate clearance of replicating virus. CD8+ T cells are primary mediators of cell death during the adaptive phase of the anti-IAV immune response, although non-CD8+ T cell killing by antibody-dependent cellular cytotoxicity is emerging as a contributor to this process, with particular implications for the development of “universal” influenza vaccines (Von Holle and Moody 2019). In mouse models of infection with attenuated strains of IAV, either anti-IAV antibodies or virus-specific CD8+ T cells are competent to promote viral clearance. In more severe infections, however, CD8+ T cells are generally required for recovery (Doherty et al. 2006).
In naïve animals or humans, the CD8+ T cell response is first detected around five days post-infection and IAV-specific CD8+ T cells accumulate to peak levels by approximately ten days after infection (Fig. 2). Secondary recall responses in previously exposed individuals are rapid, with lung resident or circulating virus-specific T cells capable of responding within hours after infection and peripheral cells recruited back to the lung within two days post-infection. In mice, the peak of secondary T cell responses are also typically earlier, resulting in more rapid virus clearance. Considering that the majority of humans will have had their first influenza infection by the age of two, older children, adults, and certainly the elderly, predominantly mount secondary-, tertiary-, or higher-level recall responses to the annual cycles of IAV infection (Oshansky and Thomas 2012).
The mechanisms of antiviral activity by CD8+ T cells include cytokine production, particularly IFN-γ and TNF-α, each of which are made in large amounts. Both these cytokines can prime target cells for apoptosis and necroptosis (Shinbori et al. 2004; Kalliolias and Ivashkiv 2016; Thapa et al. 2013). The exquisite specificity of CD8+ T cells, though, lies in their ability to selectively target infected cells by recognition of virus-specific antigenic peptide-MHC complexes on the surface of these cells, although collateral damage to nearby uninfected cells may occur (van de Sandt et al. 2017). CD8+ T cells deploy a number of killing mechanisms, including the secretion of granules containing perforin and granzymes as well as the engagement of death receptor pathways, including Fas/FasL and TRAIL/TRAIL-R (Herold et al. 2008; Hufford et al. 2015). The relative importance of these pathways is dependent on host and viral factors, such that they can be either protective or pathological depending on the strain of virus and overall virus load (Duan and Thomas 2016). Endogenous mechanisms of restraining CD8+ T cells during the effector response, such as via expression of PD1 or other immune checkpoints, are enhanced by severe infection and a highly inflamed environment. In these scenarios, blocking PD1-PDL1 signaling results in increased CD8+ T cell effector function, and is often accompanied by increased pathology, morbidity, and/or mortality (Rutigliano et al. 2014; Erickson et al. 2012; McNally et al. 2013).
Thus, as in the case of the early innate response, the effectiveness of the adaptive response depends on the ability of T cells to clear infected cells without compromising the ability of the lung to function or the capacity of the host to restore normal function to areas where damage has occurred. The large number of infiltrating T cells and their tremendous potency, however, makes this balance somewhat more precarious, and so it is perhaps not surprising that severe morbidity and mortality often occurs after viral clearance when CD8+ T cells remain in circulation in high numbers. One study suggests that the ultimate success or failure of adaptive immunity in clearing virus without irretrievably compromising host lung function can be determined by a viral “tipping point” at the onset of adaptive immune effector infiltration (Hatta et al. 2010). Viruses that have successfully evaded innate immune control, and/or that have proliferated aggressively and generated tissue damage as they have spread through the lung, will necessarily elicit a significantly more robust adaptive response than will viruses that do not manifest these behaviors. Above a critical tipping point, while the adaptive response may be capable of eliminating all infected cells, such killing would only come at the cost of irrevocable damage to the lung. Thus, the dynamic interplay of virus titers, the innate response, induction of cell death and inflammation, and the ensuing adaptive immune response combine to dictate eventual disease outcome.
3 Molecular Mechanisms of IAV-Induced Cell Death
It has been known for decades that IAV is a lytic virus. Epithelial cell lysis represents a major cause of influenza disease progression and pathogenesis, as well as of susceptibility to secondary bacterial infections in the lower respiratory tract. Indeed, the ability of IAV to kill infected cells underlies the standard plaque assay, an established method of quantifying virus by counting localized areas of dead cells, called plaques, in monolayers of infected cells. Before recent studies, this death was attributed to apoptosis, autophagy, or simply to passive, unprogrammed lysis of the infected cell (Yatim and Albert 2011; Herold et al. 2012). Of the pathways of programmed cell death, apoptosis was the first to be implicated in mediating the killing of IAV-infected cells (Herold et al. 2012). Over the years, various studies have revealed roles for both extrinsic (i.e., mediated by death receptors of the TNFR superfamily) and intrinsic (i.e., mitochondrial) pathways of apoptosis in IAV-associated death of epithelial and immune cells in culture, and of lung cell types in vivo (Herold et al. 2012). Type I IFNs and ISGs (e.g., PKR, TRAIL) produced during active IAV infection potentiate IAV-induced apoptosis (Balachandran et al. 2000a, b; Herold et al. 2012; Takizawa et al. 1996; Hogner et al. 2013). Several viral proteins, including NS1, PB1-F2, and M2, modulate apoptotic death outcomes (Herold et al. 2012; Yatim and Albert 2011). As IFNs exacerbate IAV-induced apoptosis (Balachandran et al. 2000b; Hogner et al. 2013), and as virus-encoded proteins such as NS1 delay or prevent IFN-mediated and other apoptotic pathways (Krug 2015), apoptosis is primarily considered a cell-intrinsic host defense mechanism that eliminates infected cells to prevent such cells from becoming virus factories. Interestingly, IAV may take advantage of the apoptosis machinery to boost replication, as caspase-3 deficiency compromises IAV replication (Wurzer et al. 2003). In line with these findings, the IAV protein PB1-F2 has pro-apoptotic activity in certain settings (Kosik et al. 2013). IAV also activates an autophagic response in infected cells, subject to modulation by M2 (Beale et al. 2014; Gannage et al. 2009). Collectively, these observations demonstrate a complex interplay between IAV and the host apoptotic and autophagic machinery, and suggest that, early in the virus life cycle, IAV impedes induction of cell death, via blockade of both apoptosis and autophagy, to prolong cell viability and allow the virus time to replicate. Later in the replicative cycle, IAV may promote cell death to not only eliminate immune and barrier cells that would otherwise limit viral spread, but also to potentially render death of the infected cell as immunologically “silent” as possible.
The mechanisms by which IAV regulates death receptor and mitochondrial apoptosis, as well as the significance of autophagy and apoptotic death modalities to IAV replication and pathogenesis have been comprehensively reviewed elsewhere (Herold et al. 2012; Yatim and Albert 2011; Peteranderl and Herold 2017). Here, we will focus on previously unrecognized pathways of IAV-activated cell death mediated by the host sensor protein ZBP1 and its downstream effector receptor interacting protein (RIP) kinase (RIPK)3.
3.1 Role of RIPK3 in IAV-Initiated Cell Death
RIPK3 is the mediator of a form of programmed necrotic cell death known as necroptosis (Pasparakis and Vandenabeele 2015). Necroptosis is activated by virus and microbial infections, and by RIP homotypic interaction motif (RHIM)-dependent innate immune signaling pathways, most notably those initiated by TNF, IFNs, and TLRs (Vanden Berghe et al. 2016). Once activated, RIPK3 assembles a cytosolic signaling complex that contains, at a minimum, the additional proteins RIPK1, FADD, caspase-8, and MLKL (Li et al. 2012; Moquin et al. 2013; Zhang et al. 2016). Under conditions where FADD or caspase-8 activity is compromised, RIPK3 phosphorylates and activates MLKL, which then oligomerizes and traffics to the plasma membrane where it mediates pore formation, triggering cell swelling and lysis (Zhang et al. 2016). Necroptosis causes the extensive release of DAMPs into the extracellular space and is considered highly inflammatory and immunogenic (Pasparakis and Vandenabeele 2015; Silke et al. 2015; Aaes et al. 2016). Notably, necrotic pathology is observed in mice during lethal IAV infection (Sanders et al. 2011, 2013; Kash et al. 2006; Brandes et al. 2013) and in human patients with severe IAV disease and ARDS (Korteweg and Gu 2008; Mauad et al. 2010).
Saleh and colleagues first implicated RIPK3 in IAV-activated cell death responses when they observed that mice lacking the E3 ubiquitin ligase cIAP2 were hypersusceptible to IAV-triggered lethality (Rodrigue-Gervais et al. 2014). IAV infection induces severe bronchiolar degradation via a RIPK3-dependent death pathway in cIAP2-deficient mice. Crossing cIAP2-deficient mice onto a RIPK3-null genetic background ameliorated pulmonary tissue damage and reversed hypersusceptibility to IAV lethality. Eliminating expression of TRAIL and Fas ligand in the hematopoetic compartment also ameliorated disease susceptibility, implicating these death receptor ligands in activation of necroptosis during IAV infections. Interestingly, neither the kinetics of virus clearance nor the adaptive immune response to IAV were significantly altered in the absence of cIAP2, suggesting that cIAP2 promoted disease primarily by lowering the threshold for RIPK3 activation by Fas and TRAIL receptors. The authors speculated that activation of these death signals in epithelial cells was likely a “bystander” effect, arising more from Fas ligand/TRAIL-expressing immune cells in the vicinity of the infection, than from virus replication in the infected cells themselves (Rodrigue-Gervais et al. 2014). Thus, whether IAV directly activates RIPK3 in infected cells remained unresolved.
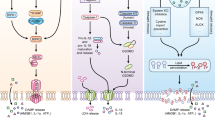
We reported that RIPK3 is directly activated by IAV and is required for IAV-induced death of fibroblasts as well as AECs (Nogusa et al. 2016). Activation of RIPK3 required replication competent virus and resulted in robust formation of a RIPK3 complex containing the FADD, RIPK1, and MLKL. From this death complex, RIPK3 mediated not only necroptosis (via MLKL), but also a parallel pathway of apoptosis, via RIPK1, FADD, and caspase-8 (Nogusa et al. 2016) (Fig. 3). Although RIPK3 has been previously shown to activate apoptosis via a RIPK1-FADD-caspase-8 axis (Cook et al. 2014; Mandal et al. 2014; Newton et al. 2014; Sun et al. 1999), those results were obtained primarily from overexpression studies, when certain active site mutations were introduced into RIPK3, or when RIPK3 kinase activity was blunted through the use of chemical inhibitors. Whether RIPK3-activated apoptosis occurs in physiological scenarios, for example, during an active virus infection, was unknown. Our finding that IAV triggers RIPK3-dependent apoptosis in infected cells demonstrates that RIPK3 does indeed induce apoptosis in physiological settings and positions this virus as the first pathogen that activates both apoptosis and necroptosis downstream of RIPK3. Remarkably, apoptosis and necroptosis are fully redundant mechanisms driving infected cell death. Ablation of MLKL does not much alter the kinetics or the magnitude of cell death. Instead, the form of death switches to apoptosis. Similarly, deleting either RIPK1 or FADD switches death to pure necroptosis, without significantly altering the degree or timing of death. Only when both apoptosis and necroptosis downstream of RIPK3 are simultaneously inhibited is cell death blocked. Notably, activation of necroptosis requires the protein kinase activity of RIPK3, whereas induction of apoptosis depends on RHIM signaling and proceeds independently of kinase function. Collectively, these findings demonstrate that IAV triggers necroptosis without need for concurrent suppression of apoptosis and call into question the prevalent idea in the field that necroptosis is a backup form of cell death that is only activated when apoptosis pathways are disabled. Importantly, RIPK3 is not expressed in most cell lines commonly used in IAV studies, including HeLa cells and A549 cells (He et al. 2009; Koo et al. 2015), explaining why these pathways of cell death were overlooked in earlier studies.
Model of ZBP1-induced cell death following IAV infection. Replicating IAV produces viral RNAs that are sensed by ZBP1 via its Zα domains. Once activated, ZBP1 associates with RIPK3 via homotypic interactions between the RHIMs of both proteins. Downstream of RIPK3, parallel pathways of MLKL-driven necroptosis and RIPK1/FADD/caspase-8-mediated apoptosis cooperate to eliminate the infected cell. ZBP1 also activates pyroptosis in macrophages, via RIPK3 and NLRP3. “ASC,” apoptosis-associated speck-like protein containing a CARD; “KD,” serine/threonine kinase domain; “DD,” death domain; “DED,” death effector domain; and “Casp,” caspase
The results detailed above (Nogusa et al. 2016) were obtained from viability analyses of cell populations in culture and do not provide insight into cell death events at the single-cell level. It was therefore unclear whether RIPK3 activates both apoptosis and necroptosis in the same cell, resulting in death with mixed apoptotic and necroptotic features, or whether RIPK3 can activate only one pathway at a time (on a per-cell basis), and activation of one pathway (e.g., apoptosis) precludes activation of the other (in this example, necroptosis). Our recent unpublished data support the idea that apoptosis and necroptosis are mutually exclusive fates that do not occur in the same cell. These results, obtained from examination of IAV-infected cells for evidence of apoptosis (i.e., the presence of cleaved caspases) or necroptosis (i.e., phosphorylated MLKL) at the single-cell level by immunofluorescence, showed that the infected cell is positive for either active caspases or phosphorylated MLKL; very rarely are both signals observed in the same cell. Thus, once a particular cell fate downstream of RIPK3 is “selected,” the other likely becomes unavailable. Post-translational modifications of RIPK3 or its binding partners may represent one mechanism by which the cell commits to one fate over the other; for example, the results from the Saleh laboratory suggest that ubiquitylation of RIPK3 may favor downstream execution of apoptosis over necroptosis (Rodrigue-Gervais et al. 2014). Such a scenario is not without precendent: in TNF signaling, ubiquitylation of RIPK1 is a chief determinant of whether a cell activates NF-κB survival signaling or undergoes cell death (Weinlich and Green 2014). The relative abundance of effector proteins available to RIPK3, both stochastically on a per-cell level, as well as between individual pulmonary cell types, is another factor that may determine both mode and magnitude of cell death. Vaux and colleagues have shown that simple availability of apoptosis verus necroptosis effectors can determine if RIPK3 will activate one form of death or the other (Cook et al. 2014). Cells in which levels of FADD, RIPK1, and/or caspase-8 are more abundant than MLKL may therefore preferentially undergo apoptosis when RIPK3 is activated; conversely, necroptosis may occur when MLKL availability exceeds those of the apoptosis effectors. Given the higher immunogenic and inflammatory potential of necroptosis over apoptosis (Yatim et al. 2015), it will be interesting to determine whether individual lung cell types are primed to undergo one form of death versus the other upon IAV infection. Intriguingly, neither pathway of cell death may operate in certain cell types: tenOever and colleagues have shown that a subset of club cells survive IAV infection in vivo and contribute to lung inflammation even after virus has been cleared (Heaton et al. 2014). Similarly, activation of RIPK3 in alveolar macrophages does not trigger cell death. In this setting, RIPK3 is required for optimal production of type I IFNs by RLRs (Downey et al. 2017).
3.2 ZBP1 as a RIPK3-Activating Sensor of IAV
Activation of RIPK3 by IAV was found to proceed independently of known RNA virus-sensing pathways, including PKR, RLRs, or TLRs, suggesting that an as-yet undiscovered sensing mechanism was responsible for stimulating RIPK3 in IAV-infected cells. In 2016, two groups, the Kanneganti laboratory and ours, reported that the protein ZBP1 was essential for IAV-induced cell death in fibroblasts, airway epithelial cells, and macrophages (Kuriakose et al. 2016; Thapa et al. 2016). ZBP1 is a 411 a.a. (in mice) protein containing two tandem Zα domains (Zα1 and Zα2) toward its N-terminus followed by a centrally positioned RHIM that mediates signal transduction and at least two additional RHIM-like sequences (Kaiser et al. 2008; Rebsamen et al. 2009) that do not contribute to cell death signaling during IAV infection (Thapa et al. 2016). ZBP1 was initially recognized as an IFN-inducible protein capable of binding Z-form double-stranded nucleic acid via its Zα domains (Schwartz et al. 2001; Fu et al. 1999). ZBP1 was subsequently proposed to function as a sensor of cytosolic DNA, initiating an innate signaling pathway leading to induction of NF-κB (Kaiser et al. 2008; Rebsamen et al. 2009) and IFNs (Takaoka et al. 2007). Subsequent studies, however, demonstrated that ZBP1-deficient cells and mice retained the ability to respond to cytosolic DNA (Ishii et al. 2008). The cGAS–STING pathway is now recognized as a dominant sensor of cytoplasmic DNA leading to type I IFN production (Li et al. 2013; Sun et al. 2013). ZBP1, instead, was found to mediate necroptosis following infection with murine cytomegalovirus (MCMV), a herpesvirus with a DNA genome (Upton et al. 2010, 2012). Thus, ZBP1 is an instigator of cell death—not IFN production—during virus infections. The identification of ZBP1 as necessary for IAV-induced cell death is the first time this sensor has been shown to respond to an RNA virus. Following IAV infection, ZBP1 complexes with RIPK3 and triggers association of RIPK3 with both RIPK1 (for apoptosis) and MLKL (for necroptosis) (Fig. 3). Consequently, ZBP1 was essential for activation of both caspase-8 and MLKL downstream of RIPK3 (Thapa et al. 2016). In addition, ZBP1 activates a RIPK3-independent pathway of apoptosis by direct RHIM-dependent recruitment of RIPK1 and subsequent DD-dependent recruitment of FADD, leading to the activation of caspase-8 (Thapa et al. 2016). Moreover, Kanneganti and colleagues showed that ZBP1 activates the NLRP3 inflammasome and induces pyroptosis in IAV-infected macrophages (Kuriakose et al. 2016) (Fig. 3). Of note, NLRP3 inflammasome activation by other negative strand RNA viruses involves RIPK3 (Wang et al. 2014; Kuriakose et al. 2016), but the requirement for ZBP1 in inflammasome activation appears unique to IAV (Kuriakose et al. 2016). In sum, ZBP1 appears to be central to all major pathways of IAV-activated cell death in primary cells; consequently, cells lacking ZBP1 are remarkably resistant to IAV-triggered death, even more so than cells lacking RIPK3 (Thapa et al. 2016).
Through systematic mutagenesis, we found that the Zα1 domain and C-terminal third of ZBP1 are not required for function, but that Zα2 is essential. Mutations in just two Zα2 domain amino acids (N122 and Y126), shown previously to mediate contact with DNA (Ha et al. 2008), abolished death signaling in IAV-infected cells (Thapa et al. 2016). Studies with the ZBP1 Zα2 mutant strongly suggested that ZBP1 functions in cell death signaling by directly sensing IAV RNA (Thapa et al. 2016). Although ZBP1 was initially characterized as a DNA-binding protein (Takaoka et al. 2007), Zα domains from related proteins (e.g., ADAR-1) can associate with the left-handed (Z-form) double helical form of RNA in vitro (Athanasiadis 2012; Placido et al. 2007). Indeed, our in silico simulation predicts that a putative complex between ZBP1 Zα2 dimers and Z-RNA is almost identical to the published ZBP1:Z-DNA crystal structure (Ha et al. 2008), and that the two residues shown to contact Z-DNA, N122 and Y126, are also properly positioned to make contacts with Z-RNA. Thus, ZBP1 may be a bifunctional protein capable of binding both DNA and RNA. In IAV-infected cells, ZBP1 robustly and preferentially associated with IAV genomic (i.e., negative polarity) RNAs (Thapa et al. 2016). These RNAs fell into two classes: (1) smaller (~500 bp) RNA species, mapping to the very 3′ and 5′ ends of the polymerase gene segments; and (2) somewhat longer RNAs (1000–1500 bp) representing full-length IAV gene segments, most notably NA and NP. The Class I (500 bp) RNAs are internally deleted subgenomic versions of the longer polymerase gene segments. These truncated RNAs are formed when the IAV polymerase falls off its template RNAs but re-engages further downstream along the same RNA strands. Subgenomic vRNAs produced in this manner nonetheless retain their 3′ and 5′ packaging signals and can be sorted into replication-incompetent defective interfering (DI) particles (Nayak et al. 1985; Saira et al. 2013). It is noteworthy that the spectrum of IAV vRNAs associated with ZBP1 bears striking similarity to those bound by the RNA sensor RIG-I (Baum et al. 2010) indicating that these RNAs, especially if packaged into DI particles (i.e., Class I), are likely improperly encapsidated and thus perhaps more accessible to the host innate machinery. Indeed, Lopez and colleagues have demonstrated that subgenomic vRNAs found in DI particles are naturally produced during the course of virus infection in vivo and are primary instigators of immune responses in infected lungs (Tapia et al. 2013). Whether some of these RNAs adopt a Z-form double-stranded conformation remains to be determined.
The findings described above suggest a model of ZBP1–RIPK3 cell death signaling in which ZBP1 first senses genomic and subgenomic RNAs via its Zα2 domain and dimerizes to provide a platform for RHIM-dependent recruitment of RIPK3. ZBP1 then clusters RIPK3, which initiates cell death signaling (Thapa et al. 2016) (Fig. 3). ZBP1 is ubiquitylated upon IAV infection (Kesavardhana et al. 2017), but how ubiquitylation regulates ZBP1 function and RIPK3 activation remains to be determined. Interestingly, evidence has implicated ZBP1 sensing of newly synthesized RNA, rather than DNA, during infection with the DNA viruses MCMV and HSV1, suggesting that nascent viral RNAs may represent ligands for ZBP1 even for DNA viruses (Maelfait et al. 2017; Sridharan et al. 2017; Guo et al. 2018)
In vivo, loss of ZBP1 rendered mice highly susceptible to intranasal infection by IAV. ZBP1-deficient mice were unable to control virus replication or prevent virus spread through the lung (Thapa et al. 2016; Momota et al. 2019). Similarly, loss of RIPK3, or combined loss of apoptosis and necroptosis pathways (i.e., Mlkl−/−Fadd−/− double-knockouts) downstream of RIPK3, rendered mice incapable of controlling IAV replication in lungs, resulting in lethal infection (Nogusa et al. 2016). Strikingly, mice deficient only in MLKL resolved infection in a manner largely indistinguishable from WT mice (Nogusa et al. 2016). Similarly, our unpublished results show that mice harboring an inactivating mutation in caspase-8, and therefore selectively incapable of supporting IAV-activated apoptosis, also mount robust antiviral responses that are comparable to wild-type animals. These results demonstrate that necroptosis and apoptosis are redundant with each other for anti-IAV host defense. Only when both pathways are neutralized is antiviral defense compromised. In Ripk3−/− mice, recruitment of T cells to the infected lung was diminished, as were anti-IAV adaptive immune responses, despite higher virus production in these animals (Nogusa et al. 2016). Notably, phosphorylated MLKL, indicative of active necroptosis, is observed not only in lung structural cells (such as AECs), but also in infiltrating immune cells, in vivo (Wang et al. 2019). Thus, RIPK3-mediated cell death may promote virus clearance not only by preventing infected cells from becoming “factories” for virus replication, but also by promoting adaptive immunity to IAV, such as via release of immunogenic DAMPs from necroptotic cells (Yatim et al. 2017).
The NLRP3 inflammasome has also been implicated in protection against IAV (Thomas et al. 2009; Allen et al. 2009; Ichinohe et al. 2009), so ZBP1-RIPK3-mediated activation of this inflammasome (Wang et al. 2014; Kuriakose et al. 2016) may co-operate with apoptosis and necroptosis to mediate virus clearance, alter the adaptive immune response, and promote healing of lung tissue post-infection. Although NLRP3 inflammasome activation can be protective during IAV infection, it may also contribute to pathology, depending on virus subtype, severity of disease, and stage of infection (Tate and Mansell 2018; Kuriakose and Kanneganti 2017). For example, activation of RIPK3 (Xu et al. 2017) and the NLRP3 inflammasome (Ren et al. 2017) are detrimental to animal survival following infection with virulent strains of H7N9 IAV, where they drive hyperinflammation that is ultimately fatal to the host. Similarly, ZBP1 promotes pathology and lethality in an intratracheal model of IAV infection, where severe disease results from infection of the lower airways (Momota et al. 2019). Consistent with the idea that NLRP3 inflammasome activity is required for antiviral host defense during sublethal infection, but can mediate pathological inflammation in severe or lethal disease, administration of an NLRP3-selective small molecule inhibitor decreased animal survival when administered early (1–3 d.p.i.) upon infection with a non-lethal dose of IAV, but improved survival outcomes when given later (7–9 d.p.i) in the course of disease following a lethal dose of H1N1 IAV, when pathogenic neutrophil-mediated hyperinflammation drives morality (Brandes et al. 2013; Camp and Jonsson 2017; Narasaraju et al. 2011; Pillai et al. 2016; Bradley et al. 2012). How the ZBP1–RIPK3 axis triggers the NLRP3 inflammasome during IAV infections remains an area of active investigation. Caspase-8 signaling, rather than MLKL activity, has been proposed to link RIPK3 to the NLRP3 inflammasome in IAV-infected cells (Kuriakose et al. 2016; Kuriakose and Kanneganti 2018), as has been shown in other settings of NLRP3 activation by RIPK3 (Lawlor et al. 2015), but the mechanistic underpinnings of this pathway still need to be identified.
4 Concluding Perspectives
Is ZBP1–RIPK3 signaling a determinant of host species restriction? The RIPK3-driven cell death machinery, restricted to vertebrates, is poorly conserved across the classes of this subphylum (Dondelinger et al. 2016). Among mammals, ZBP1, RIPK3, and MLKL are absent in marsupials and MLKL appears missing in carnivorous placentals (Dondelinger et al. 2016). Curiously, neither ZBP1 nor RIPK3 are found in birds (Dondelinger et al. 2016). IAV infection in its natural host, aquatic birds, is often asymptomatic or only associated with mild symptoms (Yoon et al. 2014); it is therefore possible that the lack of ZBP1–RIPK3 signaling in birds dampens immunogenic cell death responses and allows the maintenance of gut epithelial integrity at levels that prevent severe disease during IAV infections. When these viruses jump from birds to other species, they are frequently unfit for replication or, in rare instances, cause extremely severe disease. Future studies will need to examine whether ZBP1-RIPK3 death signaling in non-avian species is a determinant of virus fitness in these species. Hypothetically, activation of the ZBP1–RIPK3 signaling axis during cross-species infections may prevent establishment of productive infection by prematurely eliminating infected cells. Alternatively, in sporadic cases of severe disease that is sometimes observed when IAV strains jump from one species to another, sub-optimal activation of ZBP1–RIPK3-mediated cell death may contribute to the rapid virus growth that drives pathology in these settings.
Does necroptosis represent a new therapeutic entrypoint for IAV disease? We have found that RIPK3-driven apoptosis can largely compensate for loss of necroptosis in clearing IAV from the infected lung (Nogusa et al. 2016). As necroptosis, but not apoptosis, requires the kinase activity of RIPK3, it can be selectively targeted with RIPK3 kinase inhibitors. The redundancy of necroptosis with apoptosis for IAV clearance therefore represents an unexpected therapeutic opportunity in IAV-induced diseases, including viral pneumonia and ARDS, in which necrotic death is implicated in pathogenesis. A selective RIPK3 inhibitor will be expected to ameliorate the deleterious effects of necroptosis, underscored by the results of Saleh and colleagues (Rodrigue-Gervais et al. 2014), without impeding virus clearance via apoptosis. Current RIPK3 inhibitors either trigger toxic conformational changes in RIPK3 (Mandal et al. 2014) or are not selective for this kinase (e.g., ponatinib (Najjar et al. 2015; Fauster et al. 2015)), highlighting a need for additional approaches targeting this kinase. We expect that a selective, non-toxic RIPK3 inhibitor will offer therapeutic benefit in IAV disease without issues of drug resistance affecting many current antivirals.
References
Aaes TL, Kaczmarek A, Delvaeye T, De Craene B, De Koker S, Heyndrickx L, Delrue I, Taminau J, Wiernicki B, De Groote P, Garg AD, Leybaert L, Grooten J, Bertrand MJ, Agostinis P, Berx G, Declercq W, Vandenabeele P, Krysko DV (2016) Vaccination with necroptotic cancer cells induces efficient anti-tumor immunity. Cell Rep 15(2):274–287
Air GM (2012) Influenza neuraminidase. Influenza Other Respir Viruses 6(4):245–256
Allen IC, Scull MA, Moore CB, Holl EK, McElvania-TeKippe E, Taxman DJ, Guthrie EH, Pickles RJ, Ting JP (2009) The NLRP3 inflammasome mediates in vivo innate immunity to influenza A virus through recognition of viral RNA. Immunity 30(4):556–565
Athanasiadis A (2012) Zalpha-domains: at the intersection between RNA editing and innate immunity. Sem Cell Dev Biol 23(3):275–280
Baccam P, Beauchemin C, Macken CA, Hayden FG, Perelson AS (2006) Kinetics of influenza A virus infection in humans. J Virol 80(15):7590–7599
Balachandran S, Roberts PC, Brown LE, Truong H, Pattnaik AK, Archer DR, Barber GN (2000a) Essential role for the dsRNA-dependent protein kinase PKR in innate immunity to viral infection. Immunity 13:129–141
Balachandran S, Roberts PC, Kipperman T, Bhalla KN, Compans RW, Archer DR, Barber GN (2000b) Alpha/beta interferons potentiate virus-induced apoptosis through activation of the FADD/caspase-8 death signaling pathway. J Virol 74(3):1513–1523
Baum A, Sachidanandam R, Garcia-Sastre A (2010) Preference of RIG-I for short viral RNA molecules in infected cells revealed by next-generation sequencing. Proc Natl Acad Sci USA 107(37):16303–16308
Beale R, Wise H, Stuart A, Ravenhill BJ, Digard P, Randow F (2014) A LC3-interacting motif in the influenza A virus M2 protein is required to subvert autophagy and maintain virion stability. Cell Host Microbe 15(2):239–247
Beigel JH, Farrar J, Han AM, Hayden FG, Hyer R, de Jong MD, Lochindarat S, Nguyen TK, Nguyen TH, Tran TH, Nicoll A, Touch S, Yuen KY (2005) Avian influenza A (H5N1) infection in humans. N Engl J Med 353(13):1374–1385
Belser JA, Wadford DA, Pappas C, Gustin KM, Maines TR, Pearce MB, Zeng H, Swayne DE, Pantin-Jackwood M, Katz JM, Tumpey TM (2010) Pathogenesis of pandemic influenza A (H1N1) and triple-reassortant swine influenza A (H1) viruses in mice. J Virol 84(9):4194–4203
Bottcher-Friebertshauser E, Klenk HD, Garten W (2013) Activation of influenza viruses by proteases from host cells and bacteria in the human airway epithelium. Pathog Dis 69(2):87–100
Bradley LM, Douglass MF, Chatterjee D, Akira S, Baaten BJ (2012) Matrix metalloprotease 9 mediates neutrophil migration into the airways in response to influenza virus-induced toll-like receptor signaling. PLoS Pathog 8(4):e1002641
Brandes M, Klauschen F, Kuchen S, Germain RN (2013) A systems analysis identifies a feedforward inflammatory circuit leading to lethal influenza infection. Cell 154(1):197–212
Camp JV, Jonsson CB (2017) A role for neutrophils in viral respiratory disease. Front Immunol 8:550
Cardani A, Boulton A, Kim TS, Braciale TJ (2017) Alveolar macrophages prevent lethal influenza pneumonia by inhibiting infection of type-1 alveolar epithelial cells. PLoS Pathog 13(1):e1006140
Chan MC, Chan RW, Chan LL, Mok CK, Hui KP, Fong JH, Tao KP, Poon LL, Nicholls JM, Guan Y, Peiris JS (2013) Tropism and innate host responses of a novel avian influenza A H7N9 virus: an analysis of ex-vivo and in-vitro cultures of the human respiratory tract. Lancet Respir Med 1(7):534–542
Cook WD, Moujalled DM, Ralph TJ, Lock P, Young SN, Murphy JM, Vaux DL (2014) RIPK1- and RIPK3-induced cell death mode is determined by target availability. Cell Death Differ 21(10):1600–1612
Davidson S, Crotta S, McCabe TM, Wack A (2014) Pathogenic potential of interferon alphabeta in acute influenza infection. Nat Commun 5:3864
de Wit E, Rasmussen AL, Feldmann F, Bushmaker T, Martellaro C, Haddock E, Okumura A, Proll SC, Chang J, Gardner D, Katze MG, Munster VJ, Feldmann H (2014) Influenza virus A/Anhui/1/2013 (H7N9) replicates efficiently in the upper and lower respiratory tracts of cynomolgus macaques. mBio 5(4):e01331-14
Doherty PC, Turner SJ, Webby RG, Thomas PG (2006) Influenza and the challenge for immunology. Nat Immunol 7(5):449–455
Dondelinger Y, Hulpiau P, Saeys Y, Bertrand MJ, Vandenabeele P (2016) An evolutionary perspective on the necroptotic pathway. Trends Cell Biol 26(10):721–732
Downey J, Pernet E, Coulombe F, Allard B, Meunier I, Jaworska J, Qureshi S, Vinh DC, Martin JG, Joubert P, Divangahi M (2017) RIPK3 interacts with MAVS to regulate type I IFN-mediated immunity to Influenza A virus infection. PLoS Pathog 13(4):e1006326
Duan S, Thomas PG (2016) Balancing immune protection and immune pathology by CD8(+) T-cell responses to influenza infection. Front Immunol 7:25
Edinger TO, Pohl MO, Stertz S (2014) Entry of influenza A virus: host factors and antiviral targets. J Gen Virol 95(Pt 2):263–277
Erickson JJ, Gilchuk P, Hastings AK, Tollefson SJ, Johnson M, Downing MB, Boyd KL, Johnson JE, Kim AS, Joyce S, Williams JV (2012) Viral acute lower respiratory infections impair CD8(+) T cells through PD-1. J Clin Invest 122(8):2967–2982
Fauster A, Rebsamen M, Huber KV, Bigenzahn JW, Stukalov A, Lardeau CH, Scorzoni S, Bruckner M, Gridling M, Parapatics K, Colinge J, Bennett KL, Kubicek S, Krautwald S, Linkermann A, Superti-Furga G (2015) A cellular screen identifies ponatinib and pazopanib as inhibitors of necroptosis. Cell Death Dis 6:e1767
Fu Y, Comella N, Tognazzi K, Brown LF, Dvorak HF, Kocher O (1999) Cloning of DLM-1, a novel gene that is up-regulated in activated macrophages, using RNA differential display. Gene 240(1):157–163
Fujita J, Ohtsuki Y, Higa H, Azuma M, Yoshinouchi T, Haranaga S, Higa F, Tateyama M (2014) Clinicopathological findings of four cases of pure influenza virus A pneumonia. Intern Med (Tokyo, Japan) 53(12):1333–1342
Gannage M, Dormann D, Albrecht R, Dengjel J, Torossi T, Ramer PC, Lee M, Strowig T, Arrey F, Conenello G, Pypaert M, Andersen J, Garcia-Sastre A, Munz C (2009) Matrix protein 2 of influenza A virus blocks autophagosome fusion with lysosomes. Cell Host Microbe 6(4):367–380
Guo H, Gilley RP, Fisher A, Lane R, Landsteiner VJ, Ragan KB, Dovey CM, Carette JE, Upton JW, Mocarski ES, Kaiser WJ (2018) Species-independent contribution of ZBP1/DAI/DLM-1-triggered necroptosis in host defense against HSV1. Cell Death Dis 9(8):816
Ha SC, Kim D, Hwang HY, Rich A, Kim YG, Kim KK (2008) The crystal structure of the second Z-DNA binding domain of human DAI (ZBP1) in complex with Z-DNA reveals an unusual binding mode to Z-DNA. Proc Natl Acad Sci USA 105(52):20671–20676
Hatta Y, Hershberger K, Shinya K, Proll SC, Dubielzig RR, Hatta M, Katze MG, Kawaoka Y, Suresh M (2010) Viral replication rate regulates clinical outcome and CD8 T cell responses during highly pathogenic H5N1 influenza virus infection in mice. PLoS Pathog 6(10):e1001139
He S, Wang L, Miao L, Wang T, Du F, Zhao L, Wang X (2009) Receptor interacting protein kinase-3 determines cellular necrotic response to TNF-alpha. Cell 137(6):1100–1111
Heaton NS, Langlois RA, Sachs D, Lim JK, Palese P, tenOever BR (2014) Long-term survival of influenza virus infected club cells drives immunopathology. J Exp Med 211(9):1707–1714
Herold S, Ludwig S, Pleschka S, Wolff T (2012) Apoptosis signaling in influenza virus propagation, innate host defense, and lung injury. J Leukoc Biol 92(1):75–82
Herold S, Steinmueller M, von Wulffen W, Cakarova L, Pinto R, Pleschka S, Mack M, Kuziel WA, Corazza N, Brunner T, Seeger W, Lohmeyer J (2008) Lung epithelial apoptosis in influenza virus pneumonia: the role of macrophage-expressed TNF-related apoptosis-inducing ligand. J Exp Med 205(13):3065–3077
Hogner K, Wolff T, Pleschka S, Plog S, Gruber AD, Kalinke U, Walmrath HD, Bodner J, Gattenlohner S, Lewe-Schlosser P, Matrosovich M, Seeger W, Lohmeyer J, Herold S (2013) Macrophage-expressed IFN-beta contributes to apoptotic alveolar epithelial cell injury in severe influenza virus pneumonia. PLoS Pathog 9(2):e1003188
Hufford MM, Kim TS, Sun J, Braciale TJ (2015) The effector T cell response to influenza infection. Curr Top Microbiol Immunol 386:423–455
Ichinohe T, Lee HK, Ogura Y, Flavell R, Iwasaki A (2009) Inflammasome recognition of influenza virus is essential for adaptive immune responses. J Exp Med 206(1):79–87
Ishii KJ, Kawagoe T, Koyama S, Matsui K, Kumar H, Kawai T, Uematsu S, Takeuchi O, Takeshita F, Coban C, Akira S (2008) TANK-binding kinase-1 delineates innate and adaptive immune responses to DNA vaccines. Nature 451(7179):725–729
Jost S, Altfeld M (2013) Control of human viral infections by natural killer cells. Annual Review Immunol 31:163–194
Kaiser WJ, Upton JW, Mocarski ES (2008) Receptor-interacting protein homotypic interaction motif-dependent control of NF-kappa B activation via the DNA-dependent activator of IFN regulatory factors. J Immunol 181(9):6427–6434
Kalliolias GD, Ivashkiv LB (2016) TNF biology, pathogenic mechanisms and emerging therapeutic strategies. Nat Rev Rheumatol 12(1):49–62
Kash JC, Tumpey TM, Proll SC, Carter V, Perwitasari O, Thomas MJ, Basler CF, Palese P, Taubenberger JK, Garcia-Sastre A, Swayne DE, Katze MG (2006) Genomic analysis of increased host immune and cell death responses induced by 1918 influenza virus. Nature 443(7111):578–581
Kesavardhana S, Kuriakose T, Guy CS, Samir P, Malireddi RKS, Mishra A, Kanneganti TD (2017) ZBP1/DAI ubiquitination and sensing of influenza vRNPs activate programmed cell death. J Exp Med 214(8):2217–2229
Kobasa D, Jones SM, Shinya K, Kash JC, Copps J, Ebihara H, Hatta Y, Kim JH, Halfmann P, Hatta M, Feldmann F, Alimonti JB, Fernando L, Li Y, Katze MG, Feldmann H, Kawaoka Y (2007) Aberrant innate immune response in lethal infection of macaques with the 1918 influenza virus. Nature 445(7125):319–323
Koo GB, Morgan MJ, Lee DG, Kim WJ, Yoon JH, Koo JS, Kim SI, Kim SJ, Son MK, Hong SS, Levy JM, Pollyea DA, Jordan CT, Yan P, Frankhouser D, Nicolet D, Maharry K, Marcucci G, Choi KS, Cho H, Thorburn A, Kim YS (2015) Methylation-dependent loss of RIP3 expression in cancer represses programmed necrosis in response to chemotherapeutics. Cell Res 25(6):707–725
Korteweg C, Gu J (2008) Pathology, molecular biology, and pathogenesis of avian influenza A (H5N1) infection in humans. Am J Pathol 172(5):1155–1170
Kosik I, Holly J, Russ G (2013) PB1-F2 expedition from the whole protein through the domain to aa residue function. Acta Virol 57(2):138–148
Krug RM (2015) Functions of the influenza A virus NS1 protein in antiviral defense. Curr Opin Virol 12:1–6
Kuiken T, van den Brand J, van Riel D, Pantin-Jackwood M, Swayne DE (2010) Comparative pathology of select agent influenza A virus infections. Vet Pathol 47(5):893–914
Kuriakose T, Kanneganti TD (2017) Regulation and functions of NLRP3 inflammasome during influenza virus infection. Mol Immunol 86:56–64
Kuriakose T, Kanneganti TD (2018) ZBP1: innate sensor regulating cell death and inflammation. Trends Immunol 39(2):123–134
Kuriakose T, Man SM, Malireddi RKS, Karki R, Kesavardana S, Place DE, Neale G, Vogel P, Kanneganti TD (2016) ZBP1/DAI is an innate sensor of influenza virus triggering the NLRP3 inflammasome and programmed cell death pathways. Sci Immunol 1:aag2045
Lakadamyali M, Rust MJ, Zhuang X (2004) Endocytosis of influenza viruses. Microbes Infect 6(10):929–936
Lakdawala SS, Jayaraman A, Halpin RA, Lamirande EW, Shih AR, Stockwell TB, Lin X, Simenauer A, Hanson CT, Vogel L, Paskel M, Minai M, Moore I, Orandle M, Das SR, Wentworth DE, Sasisekharan R, Subbarao K (2015) The soft palate is an important site of adaptation for transmissible influenza viruses. Nature 526(7571):122–125
Lawlor KE, Khan N, Mildenhall A, Gerlic M, Croker BA, D’Cruz AA, Hall C, Kaur Spall S, Anderton H, Masters SL, Rashidi M, Wicks IP, Alexander WS, Mitsuuchi Y, Benetatos CA, Condon SM, Wong WW, Silke J, Vaux DL, Vince JE (2015) RIPK3 promotes cell death and NLRP3 inflammasome activation in the absence of MLKL. Nat Commun 6:6282
Li J, McQuade T, Siemer AB, Napetschnig J, Moriwaki K, Hsiao YS, Damko E, Moquin D, Walz T, McDermott A, Chan FK, Wu H (2012) The RIP1/RIP3 necrosome forms a functional amyloid signaling complex required for programmed necrosis. Cell 150(2):339–350
Li XD, Wu J, Gao D, Wang H, Sun L, Chen ZJ (2013) Pivotal roles of cGAS-cGAMP signaling in antiviral defense and immune adjuvant effects. Science 341(6152):1390–1394
Maelfait J, Liverpool L, Bridgeman A, Ragan KB, Upton JW, Rehwinkel J (2017) Sensing of viral and endogenous RNA by ZBP1/DAI induces necroptosis. EMBO J 36(17):2529–2543
Mandal P, Berger SB, Pillay S, Moriwaki K, Huang C, Guo H, Lich JD, Finger J, Kasparcova V, Votta B, Ouellette M, King BW, Wisnoski D, Lakdawala AS, DeMartino MP, Casillas LN, Haile PA, Sehon CA, Marquis RW, Upton J, Daley-Bauer LP, Roback L, Ramia N, Dovey CM, Carette JE, Chan FK, Bertin J, Gough PJ, Mocarski ES, Kaiser WJ (2014) RIP3 induces apoptosis independent of pronecrotic kinase activity. Mol Cell 56(4):481–495
Mauad T, Hajjar LA, Callegari GD, da Silva LF, Schout D, Galas FR, Alves VA, Malheiros DM, Auler JO Jr, Ferreira AF, Borsato MR, Bezerra SM, Gutierrez PS, Caldini ET, Pasqualucci CA, Dolhnikoff M, Saldiva PH (2010) Lung pathology in fatal novel human influenza A (H1N1) infection. Am J Respir Crit Care Med 181(1):72–79
McNally B, Ye F, Willette M, Flano E (2013) Local blockade of epithelial PDL-1 in the airways enhances T cell function and viral clearance during influenza virus infection. J Virol 87(23):12916–12924
Meliopoulos VA, Karlsson EA, Kercher L, Cline T, Freiden P, Duan S, Vogel P, Webby RJ, Guan Y, Peiris M, Thomas PG, Schultz-Cherry S (2014) Human H7N9 and H5N1 influenza viruses differ in induction of cytokines and tissue tropism. J Virol 88(22):12982–12991
Momota M, Lelliott P, Kubo A, Kusakabe T, Kobiyama K, Kuroda E, Imai Y, Akira S, Coban C, Ishii KJ (2019) ZBP1 governs the inflammasome-independent IL-1alpha and neutrophil inflammation that play a dual role in anti-influenza virus immunity. Int Immunol 32(3):203–212
Moquin DM, McQuade T, Chan FK (2013) CYLD deubiquitinates RIP1 in the TNFalpha-induced necrosome to facilitate kinase activation and programmed necrosis. PLoS ONE 8(10):e76841
Mordstein M, Kochs G, Dumoutier L, Renauld JC, Paludan SR, Klucher K, Staeheli P (2008) Interferon-lambda contributes to innate immunity of mice against influenza A virus but not against hepatotropic viruses. PLoS Pathog 4(9):e1000151
Mordstein M, Neugebauer E, Ditt V, Jessen B, Rieger T, Falcone V, Sorgeloos F, Ehl S, Mayer D, Kochs G, Schwemmle M, Gunther S, Drosten C, Michiels T, Staeheli P (2010) Lambda interferon renders epithelial cells of the respiratory and gastrointestinal tracts resistant to viral infections. J Virol 84(11):5670–5677
Najjar M, Suebsuwong C, Ray SS, Thapa RJ, Maki JL, Nogusa S, Shah S, Saleh D, Gough PJ, Bertin J, Yuan J, Balachandran S, Cuny GD, Degterev A (2015) Structure guided design of potent and selective ponatinib-based hybrid inhibitors for RIPK1. Cell Rep 10(11):1850–1860
Narasaraju T, Yang E, Samy RP, Ng HH, Poh WP, Liew AA, Phoon MC, van Rooijen N, Chow VT (2011) Excessive neutrophils and neutrophil extracellular traps contribute to acute lung injury of influenza pneumonitis. Am J Pathol 179(1):199–210
Nayak DP, Chambers TM, Akkina RK (1985) Defective-interfering (DI) RNAs of influenza viruses: origin, structure, expression, and interference. Curr Top Microbiol Immunol 114:103–151
Newton K, Dugger DL, Wickliffe KE, Kapoor N, de Almagro MC, Vucic D, Komuves L, Ferrando RE, French DM, Webster J, Roose-Girma M, Warming S, Dixit VM (2014) Activity of protein kinase RIPK3 determines whether cells die by necroptosis or apoptosis. Science 343(6177):1357–1360
Nogusa S, Thapa RJ, Dillon CP, Liedmann S, Oguin TH 3rd, Ingram JP, Rodriguez DA, Kosoff R, Sharma S, Sturm O, Verbist K, Gough PJ, Bertin J, Hartmann BM, Sealfon SC, Kaiser WJ, Mocarski ES, Lopez CB, Thomas PG, Oberst A, Green DR, Balachandran S (2016) RIPK3 activates parallel pathways of MLKL-driven necroptosis and FADD-mediated apoptosis to protect against influenza A virus. Cell Host Microbe 20(1):13–24
Oshansky CM, Thomas PG (2012) The human side of influenza. J Leukoc Biol 92(1):83–96
Pasparakis M, Vandenabeele P (2015) Necroptosis and its role in inflammation. Nature 517(7534):311–320
Peteranderl C, Herold S (2017) The impact of the interferon/TNF-related apoptosis-inducing ligand signaling axis on disease progression in respiratory viral infection and beyond. Front Immunol 8:313
Pillai PS, Molony RD, Martinod K, Dong H, Pang IK, Tal MC, Solis AG, Bielecki P, Mohanty S, Trentalange M, Homer RJ, Flavell RA, Wagner DD, Montgomery RR, Shaw AC, Staeheli P, Iwasaki A (2016) Mx1 reveals innate pathways to antiviral resistance and lethal influenza disease. Science 352(6284):463–466
Placido D, Brown BA 2nd, Lowenhaupt K, Rich A, Athanasiadis A (2007) A left-handed RNA double helix bound by the Z alpha domain of the RNA-editing enzyme ADAR1. Structure 15(4):395–404
Rebsamen M, Heinz LX, Meylan E, Michallet MC, Schroder K, Hofmann K, Vazquez J, Benedict CA, Tschopp J (2009) DAI/ZBP1 recruits RIP1 and RIP3 through RIP homotypic interaction motifs to activate NF-kappaB. EMBO Rep 10(8):916–922
Ren R, Wu S, Cai J, Yang Y, Ren X, Feng Y, Chen L, Qin B, Xu C, Yang H, Song Z, Tian D, Hu Y, Zhou X, Meng G (2017) The H7N9 influenza A virus infection results in lethal inflammation in the mammalian host via the NLRP3-caspase-1 inflammasome. Sci Rep 7(1):7625
Rodrigue-Gervais IG, Labbe K, Dagenais M, Dupaul-Chicoine J, Champagne C, Morizot A, Skeldon A, Brincks EL, Vidal SM, Griffith TS, Saleh M (2014) Cellular inhibitor of apoptosis protein cIAP2 protects against pulmonary tissue necrosis during influenza virus infection to promote host survival. Cell Host Microbe 15(1):23–35
Rosenberger CM, Podyminogin RL, Askovich PS, Navarro G, Kaiser SM, Sanders CJ, McClaren JL, Tam VC, Dash P, Noonan JG, Jones BG, Surman SL, Peschon JJ, Diercks AH, Hurwitz JL, Doherty PC, Thomas PG, Aderem A (2014) Characterization of innate responses to influenza virus infection in a novel lung type I epithelial cell model. J Gen Virol 95(Pt 2):350–362
Rossman JS, Lamb RA (2011) Influenza virus assembly and budding. Virology 411(2):229–236
Rutigliano JA, Sharma S, Morris MY, Oguin TH, McClaren JL, Doherty PC, Thomas PG (2014) Highly pathological influenza A virus infection is associated with augmented expression of PD-1 by functionally compromised virus-specific CD8(+) T cells. J Virol 88(3):1636–1651
Saira K, Lin X, DePasse JV, Halpin R, Twaddle A, Stockwell T, Angus B, Cozzi-Lepri A, Delfino M, Dugan V, Dwyer DE, Freiberg M, Horban A, Losso M, Lynfield R, Wentworth DN, Holmes EC, Davey R, Wentworth DE, Ghedin E, Group IFS, Group IFS (2013) Sequence analysis of in vivo defective interfering-like RNA of influenza A H1N1 pandemic virus. J Virol 87 (14):8064–8074
Samet SJ, Tompkins SM (2017) Influenza pathogenesis in genetically defined resistant and susceptible murine strains. Yale J Biol Med 90(3):471–479
Sanders CJ, Doherty PC, Thomas PG (2011) Respiratory epithelial cells in innate immunity to influenza virus infection. Cell Tissue Res 343(1):13–21
Sanders CJ, Vogel P, McClaren JL, Bajracharya R, Doherty PC, Thomas PG (2013) Compromised respiratory function in lethal influenza infection is characterized by the depletion of type I alveolar epithelial cells beyond threshold levels. Am J Physiol Lung Cell Mol Physiol 304(7):L481–L488
Schwartz T, Behlke J, Lowenhaupt K, Heinemann U, Rich A (2001) Structure of the DLM-1–Z-DNA complex reveals a conserved family of Z-DNA-binding proteins. Nat Struct Biol 8(9):761–765
Shieh W-J, Blau DM, Denison AM, DeLeon-Carnes M, Adem P, Bhatnagar J, Sumner J, Liu L, Patel M, Batten B, Greer P, Jones T, Smith C, Bartlett J, Montague J, White E, Rollin D, Gao R, Seales C, Jost H, Metcalfe M, Goldsmith CS, Humphrey C, Schmitz A, Drew C, Paddock C, Uyeki TM, Zaki SR (2010) 2009 Pandemic influenza A (H1N1): pathology and pathogenesis of 100 fatal cases in the United States. Am J Pathol 177(1):166–175
Shinbori T, Walczak H, Krammer PH (2004) Activated T killer cells induce apoptosis in lung epithelial cells and the release of pro-inflammatory cytokine TNF-alpha. Eur J Immunol 34(6):1762–1770
Silke J, Rickard JA, Gerlic M (2015) The diverse role of RIP kinases in necroptosis and inflammation. Nat Immunol 16(7):689–697
Soto-Abraham MV, Soriano-Rosas J, Diaz-Quinonez A, Silva-Pereyra J, Vazquez-Hernandez P, Torres-Lopez O, Roldan A, Cruz-Gordillo A, Alonso-Viveros P, Navarro-Reynoso F (2009) Pathological changes associated with the 2009 H1N1 virus. N Eng J Med 361(20):2001–2003
Sridharan H, Ragan KB, Guo H, Gilley RP, Landsteiner VJ, Kaiser WJ, Upton JW (2017) Murine cytomegalovirus IE3-dependent transcription is required for DAI/ZBP1-mediated necroptosis. EMBO Rep 18(8):1429–1441
Sun L, Wu J, Du F, Chen X, Chen ZJ (2013) Cyclic GMP-AMP synthase is a cytosolic DNA sensor that activates the type I interferon pathway. Science 339(6121):786–791
Sun X, Lee J, Navas T, Baldwin DT, Stewart TA, Dixit VM (1999) RIP3, a novel apoptosis-inducing kinase. J Biol Chem 274(24):16871–16875
Takaoka A, Wang Z, Choi MK, Yanai H, Negishi H, Ban T, Lu Y, Miyagishi M, Kodama T, Honda K, Ohba Y, Taniguchi T (2007) DAI (DLM-1/ZBP1) is a cytosolic DNA sensor and an activator of innate immune response. Nature 448(7152):501–505
Takizawa T, Ohashi K, Nakanishi Y (1996) Possible involvement of double-stranded RNA-activated protein kinase in cell death by influenza virus infection. J Virol 70(11):8128–8132
Tapia K, Kim WK, Sun Y, Mercado-Lopez X, Dunay E, Wise M, Adu M, Lopez CB (2013) Defective viral genomes arising in vivo provide critical danger signals for the triggering of lung antiviral immunity. PLoS Pathog 9(10):e1003703
Tate MD, Mansell A (2018) An update on the NLRP3 inflammasome and influenza: the road to redemption or perdition? Curr Opin Immunol 54:80–85
te Velthuis AJW, Fodor E (2016) Influenza virus RNA polymerase: insights into the mechanisms of viral RNA synthesis. Nat Rev Microbiol 14(8):479–493
Thangavel RR, Bouvier NM (2014) Animal models for influenza virus pathogenesis, transmission, and immunology. J Immunol Methods 410:60–79
Thapa RJ, Ingram JP, Ragan KB, Nogusa S, Boyd DF, Benitez AA, Sridharan H, Kosoff R, Shubina M, Landsteiner VJ, Andrake M, Vogel P, Sigal LJ, tenOever BR, Thomas PG, Upton JW, Balachandran S (2016) DAI senses influenza A virus genomic RNA and activates RIPK3-dependent cell death. Cell Host Microbe 20(5):674–681
Thapa RJ, Nogusa S, Chen P, Maki JL, Lerro A, Andrake M, Rall GF, Degterev A, Balachandran S (2013) Interferon-induced RIP1/RIP3-mediated necrosis requires PKR and is licensed by FADD and caspases. Proc Natl Acad Sci USA 110(33):E3109–3118
Thomas PG, Dash P, Aldridge JR Jr, Ellebedy AH, Reynolds C, Funk AJ, Martin WJ, Lamkanfi M, Webby RJ, Boyd KL, Doherty PC, Kanneganti TD (2009) The intracellular sensor NLRP3 mediates key innate and healing responses to influenza A virus via the regulation of caspase-1. Immunity 30(4):566–575
Upton JW, Kaiser WJ, Mocarski ES (2010) Virus inhibition of RIP3-dependent necrosis. Cell Host Microbe 7(4):302–313
Upton JW, Kaiser WJ, Mocarski ES (2012) DAI/ZBP1/DLM-1 complexes with RIP3 to mediate virus-induced programmed necrosis that is targeted by murine cytomegalovirus vIRA. Cell Host Microbe 11(3):290–297
van de Sandt CE, Barcena M, Koster AJ, Kasper J, Kirkpatrick CJ, Scott DP, de Vries RD, Herold S, Rimmelzwaan GF, Kuiken T, Short KR (2017) Human CD8 + T cells damage non-infected epithelial cells during influenza virus infection in vitro. Am J Respir Cell Mol Biol 57(5):536–546
Vanden Berghe T, Hassannia B, Vandenabeele P (2016) An outline of necrosome triggers. Cell Mol Life Sci 73(11–12):2137–2152
Von Holle TA, Moody MA (2019) Influenza and antibody-dependent cellular cytotoxicity. Front Immunol 10:1457
Wang X, Jiang W, Yan Y, Gong T, Han J, Tian Z, Zhou R (2014) RNA viruses promote activation of the NLRP3 inflammasome through a RIP1-RIP3-DRP1 signaling pathway. Nat Immunol 15(12):1126–1133
Wang Y, Hao Q, Florence JM, Jung BG, Kurdowska AK, Samten B, Idell S, Tang H (2019) Influenza virus infection induces ZBP1 expression and necroptosis in mouse lungs. Front Cell Infect Microbiol 9:286
Wasik BR, Barnard KN, Ossiboff RJ, Khedri Z, Feng KH, Yu H, Chen X, Perez DR, Varki A, Parrish CR (2017) Distribution of O-acetylated sialic acids among target host tissues for influenza virus. mSphere 2(5):e00379-16
Watanabe T, Shinya K, Watanabe S, Imai M, Hatta M, Li C, Wolter BF, Neumann G, Hanson A, Ozawa M, Yamada S, Imai H, Sakabe S, Takano R, Iwatsuki-Horimoto K, Kiso M, Ito M, Fukuyama S, Kawakami E, Gorai T, Simmons HA, Schenkman D, Brunner K, Capuano SV, Weinfurter JT, Nishio W, Maniwa Y, Igarashi T, Makino A, Travanty EA, Wang J, Kilander A, Dudman SG, Suresh M, Mason RJ, Hungnes O, Friedrich TC, Kawaoka Y (2011) Avian-type receptor-binding ability can increase influenza virus pathogenicity in macaques. J Virol 85(24):13195–13203
Watanabe T, Tisoncik-Go J, Tchitchek N, Watanabe S, Benecke AG, Katze MG, Kawaoka Y (2013) 1918 Influenza virus hemagglutinin (HA) and the viral RNA polymerase complex enhance viral pathogenicity, but only HA induces aberrant host responses in mice. J Virol 87(9):5239–5254
Weinheimer VK, Becher A, Tonnies M, Holland G, Knepper J, Bauer TT, Schneider P, Neudecker J, Ruckert JC, Szymanski K, Temmesfeld-Wollbrueck B, Gruber AD, Bannert N, Suttorp N, Hippenstiel S, Wolff T, Hocke AC (2012) Influenza A viruses target type II pneumocytes in the human lung. J Inf Dis 206(11):1685–1694
Weinlich R, Green DR (2014) The two faces of receptor interacting protein kinase-1. Mol Cell 56(4):469–480
Wu H, Haist V, Baumgärtner W, Schughart K (2010) Sustained viral load and late death in Rag2-/- mice after influenza A virus infection. Virol J 7:172
Wurzer WJ, Planz O, Ehrhardt C, Giner M, Silberzahn T, Pleschka S, Ludwig S (2003) Caspase 3 activation is essential for efficient influenza virus propagation. EMBO J 22(11):2717–2728
Xu YL, Tang HL, Peng HR, Zhao P, Qi ZT, Wang W (2017) RIP3 deficiency ameliorates inflammatory response in mice infected with influenza H7N9 virus infection. Oncotarget 8(17):27715–27724
Yatim N, Albert ML (2011) Dying to replicate: the orchestration of the viral life cycle, cell death pathways, and immunity. Immunity 35(4):478–490
Yatim N, Cullen S, Albert ML (2017) Dying cells actively regulate adaptive immune responses. Nat Rev Immunol 17(4):262–275
Yatim N, Jusforgues-Saklani H, Orozco S, Schulz O, da Silva R Barreira, e Sousa C Reis, Green DR, Oberst A, Albert ML (2015) RIPK1 and NF-kappaB signaling in dying cells determines cross-priming of CD8(+) T cells. Science 350(6258):328–334
Yoon SW, Webby RJ, Webster RG (2014) Evolution and ecology of influenza A viruses. Curr Top Microbiol Immunol 385:359–375
Zamora AE, Aguilar EG, Sungur CM, Khuat LT, Dunai C, Lochhead GR, Du J, Pomeroy C, Blazar BR, Longo DL, Venstrom JM, Baumgarth N, Murphy WJ (2017) Licensing delineates helper and effector NK cell subsets during viral infection. JCI Insight 2(10):e87032
Zhang J, Yang Y, He W, Sun L (2016) Necrosome core machinery: MLKL. Cell Mol Life Sci 73(11–12):2153–2163
Acknowledgements
Work in the SB laboratory is supported by NIH grants AI144400, AI135025, CA168621, CA190542, P30CA006927, and an appropriation from the Commonwealth of Pennsylvania. Work in the PGT laboratory is supported by ALSAC and the National Institute of Allergy and Infectious Diseases, the National Institutes of Health, under contract number HHSN272201400006C (St. Jude Center of Excellence for Influenza Research and Surveillance).
Author information
Authors and Affiliations
Corresponding authors
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2019 Springer Nature Switzerland AG
About this chapter
Cite this chapter
Thomas, P.G., Shubina, M., Balachandran, S. (2019). ZBP1/DAI-Dependent Cell Death Pathways in Influenza A Virus Immunity and Pathogenesis. In: Mocarski, E.S., Mandal, P. (eds) Alternate Programmed Cell Death Signaling in Antiviral Host Defense. Current Topics in Microbiology and Immunology, vol 442. Springer, Cham. https://doi.org/10.1007/82_2019_190
Download citation
DOI: https://doi.org/10.1007/82_2019_190
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-031-45277-2
Online ISBN: 978-3-031-45278-9
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)