Abstract
Purpose
This study was conducted to determine the association between single-nucleotide polymorphisms (SNPs) in apoptosis-related genes and survival outcomes of patients with early-stage non-small-cell lung cancer (NSCLC).
Methods
Three hundred ten consecutive patients with surgically resected NSCLC were enrolled. Twenty-five SNPs in 17 apoptosis-related genes were genotyped by a sequenome mass spectrometry-based genotyping assay. The genotype associations with overall survival (OS) and disease-free survival (DFS) were analyzed.
Results
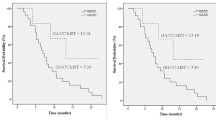
Three SNPs (TNFRSF10B rs1047266, TNFRSF1A rs4149570, and PPP1R13L rs1005165) were significantly associated with survival outcomes on multivariate analysis. When the three SNPs were combined, OS and DFS were decreased as the number of bad genotypes increased (P trend for OS and DFS = 7 × 10−5 and 1 × 10−4, respectively). Patients with one bad genotype, and patients with two or three bad genotypes had significantly worse OS and DFS compared with those with no bad genotypes [adjusted hazard ratio (aHR) for OS = 2.27, 95% confidence interval (CI) = 1.22–4.21, P = 0.01, aHR for DFS = 1.74, 95% CI = 1.08–2.81, P = 0.02; aHR for OS = 4.11, 95% CI = 2.03–8.29, P = 8 × 10−5; and aHR for DFS = 2.89, 95% CI = 1.64–5.11, P = 3 × 10−4, respectively].
Conclusion
Three SNPs in apoptosis-related genes were identified as possible prognostic markers of survival in patients with early-stage NSCLC. The SNPs, and particularly their combined genotypes, can be used to identify patients at high risk for poor disease outcome.


Similar content being viewed by others
References
Parkin DM, Bray F, Ferlay J, Pisani P. Global cancer statistics, 2002. CA Cancer J Clin. 2005;55:74–108.
Arriagada R, Bergman B, Dunant A, Le Chevalier T, Pignon JP, Vansteenkiste J. Cisplatin-based adjuvant chemotherapy in patients with completely resected non-small-cell lung cancer. N Engl J Med. 2004;350:351–60.
Coate LE, John T, Tsao MS, Shepherd FA. Molecular predictive and prognostic markers in non-small cell lung cancer. Lancet Oncol. 2009;10:1001–10.
Raff M. Cell suicide for beginners. Nature. 1998;396:119–22.
Lowe SW, Lin AW. Apoptosis in cancer. Carcinogenesis. 2000;21:485–95.
Evan GI, Vousden KH. Proliferation, cell cycle and apoptosis in cancer. Nature. 2001;411:342–8.
Shivapurkar N, Reddy J, Chaudhary PM, Gazdar AF. Apoptosis and lung cancer: a review. J Cell Biochem. 2003;88:885–98.
Schmitz A, Bayer J, Dechamps N, Thomas G. Intrinsci susceptibility to radiation-induced apoptosis of human lymphocyte subpopulations. Int J Radat Oncol Biol Phys. 2003;57:769–78.
Camplejohn RS, Hodgson S, Carter N, Kato BS, Spector TD. Heretability of DNA-damage-induced apoptosis and its relationship with age in lymphocytes from female twins. Br J Cancer. 2006;95:520–4.
Chen B, Liu S, Wang XL, Xu W, Li Y, Zhao WH, et al. TRAIL-R1 polymorphisms and cancer susceptibility: an evidence-based meta-analysis. Eur J Cancer. 2009;45:2598–2605.
Jang JS, Kim KM, Kang KH et al. Polymorphisms in the survivin gene and the risk of lung cancer. Lung Cancer. 2008;60:31–9.
Li Y, Qiu LX, Shen XK, Lv XJ, Qian XP, Song Y. A meta-analysis of TP53 codon 72 polymorphism and lung cancer risk: evidence from 15,857 subjects. Lung Cancer. 2009;66:15–21.
Gupta R, Sharma SC, Das SN. Association of TNF-alpha and TNFR1 promoters and 3’ UTR region of TNFR2 gene polymorphisms with genetic susceptibility to tobacco-related oral carcinoma in Asian Indians. Oral Oncol. 2008;44:455–63.
Yuan HY, Chiou JJ, Tseng WH et al. FASTSNP: an always up-to-date and extended service for SNP function analysis and prioritization. Nucleic Acids Res. 2006;34:W635–41.
Stephens M, Donnelly P. A comparison of Bayesian methods for haplotype reconstruction from population genotype data. Am J Hum Genet. 2003;73:1162–9.
Gabriel SB, Schaffner SF, Nguyen H et al. The structure of haplotype blocks in the human genome. Science. 2002;296:2225–9.
Locksley RM, Killeen N, Lenardo MJ. The TNF and TNF receptor superfamilies: integrating mammalian biology. Cell. 2001;104:487–501.
Aggarwal BB. Signaling pathways of the TNF superfamily: a double-edged sword. Nat Rev Immunol. 2003;3:745–56.
Han JY, Hong EK, Choi BG, et al. Death receptor 5 and Bcl-2 protein expression as predictors of tumor response to gemcitabine and cisplatin in patients with advanced non-small cell lung cancer. Med Oncol. 2003;20:355–62.
Jin H, Yang R, Fong S, et al. Apo2 ligand/tumor necrosis factor-related apoptosis-inducing ligand cooperates with chemotherapy to inhibit orthotopic lung growth and improve survival. Cancer Res. 2004;64:4900–5.
Leithner K, Stacher E, Wurm R, et al. Nuclear and cytoplasmic death receptor 5 as prognostic factors in patients with non-small cell lung cancer treated with chemotherapy. Lung Cancer. 2009;65:98–104.
Zhuang L, Lee CS, Scolyer RA, et al. Progression of melanoma is associated with decreased expression of death receptors for tumor necrosis factor-related apoptosis-inducing ligand. Hum Pathol. 2006;37:1286–94.
Camidge DR. The potential of death receptor 4- and 5-directed therapies in the treatment of lung cancer. Clin Lung Cancer. 2007;8:413–9.
Jin H, Yang R, Ross J, et al. Cooperation of the agonistic DR5 antibody apomab with chemotherapy to inhibit orthotopic lung tumor growth and improve survival. Clin Cancer Res. 2008;14:7733–40.
Ashkenazi A, Holland P, Eckhardt SG. Ligand-based targeting of apoptosis in cancer: the potential of recombinant human apoptosis ligand 2/tumor necrosis factor-related apoptosis-inducing ligand (rhApo2L/TRAIL). J Clin Oncol. 2008;26:3621–30.
Hymowitz SG, Christinger HW, Fuh G, et al. Triggering cell death: the crystal structure of Apo2L/TRAIL in a complex with death receptor 5. Mol Cell. 1999;4:563–71.
Balkwill F. Tumor necrosis factor or tumor promoting factor? Cytokine Growth Factor Rev. 2002;13:135–41.
Mocellin S, Rossi CR, Pilati P, Nitti D. Tumor necrosis factor, cancer and anticancer therapy. Cytokine Growth Factor Rev. 2005;16:35–53.
Kim S, Moon SM, Kim YS, et al. TNFR1 promoter -329G/T polymorphism results in allele specific repression in TNFR1 expression. Biochem Biophys Res Commun. 2008;368–401.
Yang JP, Hori M, Sande T, Okamoto T. Identification of a novel inhibitor of nuclear factor-кB, RelA-associated inhibitor. J Biol Chem. 1999;18:6938–47.
Bergamaschi D, Samuels Y, O’Neil NJ, et al. iASPP oncoprotein is a key inhibitor of p53 conserved from worm to human. Nat Genet. 2003;33:162–7.
Grabe N. Alibaba2: context specific identification of transcription factor binding sites. In Silico Biol. 2002;2:S1–S15.
Vogel U, Laros I, Jacobsen NR et al. Two regions in chromosome 19q13.2-3 are associated with risk of lung cancer. Mut Res. 2004;546:65–74.
Neumann AS, Sturgis EM, Wei Q. Nucleotide excision repair as a marker for susceptibility to tobacco-related cancers: a review of molecular epidemiological studies. Mol Carcinog. 2005;42:65–92.
Takenaka T, Yano T, Kiyohara C, et al. Effects of excision repair cross-complementation group 1 (ERCC1) single nucleotide polymorphisms on the prognosis of non-small cell lung cancer patients. Lung Cancer. 2009 (in press).
Chanock SJ, Manolio T, Boehnke M, et al. Replicating genotype-phenotype associations. Nature. 2007;447:655–60.
Taylor JMG, Ankerst DP, Andridge RR. Validation of biomark-based prediction models. Clin Cancer Res. 2008;14:5977–83.
Acknowledgment
This study was supported by the National R&D Program for Cancer Control Ministry of Health & Welfare (0720550-2) and the Regional Technology Innovation Program of the Ministry of Commerce, Industry and Energy (RTI04-01-01) of Republic of Korea.
Disclosures
The authors indicated no commercial interest.
Author information
Authors and Affiliations
Corresponding author
Additional information
Eung Bae Lee, Hyo-Sung Jeon have contributed equally to this study.
Rights and permissions
About this article
Cite this article
Lee, E.B., Jeon, HS., Yoo, S.S. et al. Polymorphisms in Apoptosis-Related Genes and Survival of Patients with Early-Stage Non-Small-Cell Lung Cancer. Ann Surg Oncol 17, 2608–2618 (2010). https://doi.org/10.1245/s10434-010-1082-4
Received:
Published:
Issue Date:
DOI: https://doi.org/10.1245/s10434-010-1082-4