Abstract
The transcription factors, such as activators and repressors, can interact with the promoter of gene either in a competitive or non-competitive way. In this paper, we construct a stochastic model with non-competitive transcriptional regulatory architecture and develop an analytical theory that re-establishes the experimental results with an improved data fitting. The analytical expressions in the theory allow us to study the nature of the system corresponding to any of its parameters and hence, enable us to find out the factors that govern the regulation of gene expression for that architecture. We notice that, along with transcriptional reinitiation and repressors, there are other parameters that can control the noisiness of this network. We also observe that, the Fano factor (at mRNA level) varies from sub-Poissonian regime to super-Poissonian regime. In addition to the aforementioned properties, we observe some anomalous characteristics of the Fano factor (at mRNA level) and that of the variance of protein at lower activator concentrations in the presence of repressor molecules. This model is useful to understand the architecture of interactions which may buffer the stochasticity inherent to gene transcription.
Graphical abstract



















Similar content being viewed by others
Data availability
This manuscript has associated data in a data repository. [Authors’ comment: The datasets used and generated for this study are available on request to the corresponding author.]
Notes
The Fano factor is a measure of noise. The Fano factor and noise strength are synonymous throughout the paper. For more, refer to glossary.
Where \(C_{i}\) is a constant and the factor e appearing here eventually controls the transitions from \(G_{a}\) to \(G_{ar}\) via \(k_{5}\) and from \(G_{r}\) to \(G_{ar}\) via \(k_{8}=ek_{1}\). Hence, it affects the noise strength. We will explore it later in Sect. 2.4.
Refer to Appendix-A.
Refer to Appendix-C for the corresponding analytical expressions.
References
M.B. Elowitz, A.J. Levine, E.D. Siggia, P.S. Swain, Stochastic gene expression in a single cell. Science 297, 1183 (2002)
E.M. Ozbudak, M. Thattai, I. Kurtser, A.D. Grossman, A.V. Oudenaarden, Regulation of noise in the expression of single gene. Nature Genet. 31, 69 (2002)
W.J. Blake, M. Kaern, C.R. Cantor, J.J. Collins, Noise in eukaryotic gene expression. Nature 422, 633 (2003)
J.M. Raser, E.K. O’Shea, Noise in gene expression: origins, consequences, and control. Science 309, 2010 (2005)
I. Golding, J. Paulsson, S.M. Zawilski, E.C. Cox, Real-time kinetics of gene activity in individual bacteria. Cell 123, 1025 (2005)
W.J. Blake, G. Balazsi, M.A. Kohanski, F.J. Isaas, K.F. Murphy, Y. Kuang, C.R. Cantor, D.R. Walt, J.J. Collins, Phenotypic consequences of promoter-mediated transcriptional noise. Mol. Cell 24, 853 (2006)
A. Raj, C.S. Peskin, D. Tranchina, D.Y. Vargas, S. Tyagi, Stochastic mRNA synthesis in mammalian cells. PLOS Biol. 4, e309/1707 (2006)
D.M. Suter, M. Molina, D. Gatfield, K. Schneider, U. Schibler, F. Naef, Mammalian genes are transcribed with widely different bursting kinetics. Science 332, 472 (2011)
C.R. Bartman, N. Hamagami, C.A. Keller, B. Giardine, R.C. Hardison, G.A. Blobel, A. Raj, Transcriptional burst initiation and polymerase pause release are key control points of transcriptional regulation. Mol. Cell 79, 519 (2019)
F.M.V. Rossi, A.M. Kringstein, A. Spicher, O.M. Guicherit, H.M. Blau, Transcriptional control: rheostat converted to on/off switch. Mol. Cell 6, 723 (2000)
S.R. Biggar, G.R. Crabtree, Cell signaling can direct either binary or graded transcriptional responses. EMBO J. 20, 3167 (2001)
J. Paulsson, Models of stochastic gene expression. Phys. Life Rev. 2, 157 (2005)
A. Sanchez, J. Kondev, Transcriptional control of noise in gene expression. PNAS 105, 5081 (2008)
V. Shahrezaei, P.S. Swain, Analytical distributions for stochastic gene expression. PNAS 105, 17256 (2008)
R. Karmakar, I. Bose, Graded and binary responses in stochastic gene expression. Phys. Biol. 1, 197 (2004)
R. Karmakar, Conversion of graded to binary responses in an activator-repressor system. Phys. Rev. E. 81, 021905 (2010)
N. Kumar, T. Platini, R.V. Kulkarni, Exact distribution for stochastic gene expression models with bursting and feedback. Phys. Rev. Lett. 113, 268105/1 (2014)
L. Bintu et al., Transcriptional regulation by the numbers: models. Curr. Opin. Genet. Dev. 15, 116–124 (2005)
L. Bintu et al., Transcriptional regulation by the numbers: applications. Curr. Opin. Genet. Dev. 15, 125–135 (2005)
T. Kuhlman, Z. Zhang, M.H. Saier Jr., T. Hwa, Combinatorial transcriptional control of the lactose operon of Escherichia coli. Proc. Natl. Acad. Sci. USA. 104, 6043–6048 (2007)
J.M.G. Vilar, L. Saiz, DNA looping and physical constraints on transcription regulation. J. Mol. Biol. 331, 981–989 (2003)
J.M.G. Vilar, L. Saiz, DNA looping in gene regulation: from the assembly of macromolecular complexes to the control of transcriptional noise. Curr. Opin. Genet. Dev. 15, 136–144 (2005)
M. Kaern, T.C. Elston, W.J. Blake, J.J. Collins, Stochasticity in gene expression: from theories to phenotypes. Nat. Rev. Genet 6, 451–464 (2005)
A. Raj, A. van Oudenaarden, Nature, nurture, or chance: stochastic gene expression and its consequences. Cell 135, 723–728 (2008)
A. Sanchez, S. Choubey, J. Kondev, Regulation of noise in gene expression. Annu. Rev. Biophys 42, 469–491 (2013)
M. Ptashne, Regulation of transcription: from lambda to eukaryotes. TRENDS Biochem. Sci. 30, 275 (2005)
B. Alberts, A. Johnson, J. Lewis, M. Raff, K. Roberts, P. Walters, Molecular Biology of the Cell (Garland Science, UK, 2002)
K. Struhl, Fundamentally different logic of gene regulation in eukaryotes and prokaryotes. Cell 98, 1–4 (1999)
A. Raj, A.V. Oudenaarden, Single-molecule approaches to stochastic gene expression. Annu. Rev. Biophys. 38, 255–270 (2009)
M. Acar, J.T. Mettetal, A. van Oudenaarden, Stochastic switching as a survival strategy in fluctuating environments. Nat. Genet. 40, 471–475 (2008)
Z. Cao, T. Filatova, D.A. Oyarzun, R. Grima, A stochastic model of gene expression with polymerase recruitment and pause release. Biophys. J. 119, 1002 (2020)
S. Braichenko, J. Holehouse, R. Grima, Distinguishing between models of mammalian gene expression: telegraph-like models versus mechanistic models. Interface 18, 20210510 (2021)
B. Liu, Z. Yuan, K. Aihara, L. Chen, Reinitiation enhances reliable transcriptional responses in eukaryotes. J. R. Soc. Interface 11, 0326/1–11 (2014)
W. Shao, J. Zeitlinger, Paused RNA Polymerase II inhibits new transcriptional initiation. Nat. Genetics Advance online publication (2017). https://doi.org/10.1038/ng.3867
N. Yudkovsky, J.A. Ranish, S. Hahn, A transcription reinitiation intermediate that is stabilized by activator. Nature 408, 225 (2000)
C.R. Bratman, N. Hamagami, C.A. Keller, B. Giardine, R.C. Hardison, G.A. Blobel, A. Raj, Transcriptional burst initiation and polymerase pause release are key control points of transcriptional regulation. Mol. Cell 73, 519 (2019)
R. Karmakar, Control of noise in gene expression by transcriptional reinitiation. J. Stat. Mech.: Theory Exp. 20, 063402 (2020)
R. Karmakar, A.K. Das, Effect of transcription reinitiation in stochastic gene expression. J. Stat. Mech.: Theory Exp. 21, 033502 (2021)
K.F. Murphy, G. Balazsi, J.J. Collins, Combinatorial promoter design for engineering noisy gene expression. Proc. Natl. Acad. Sci. USA 104, 12726–12731 (2007)
Y. Setty, A.E. Mayo, M.G. Surette, U. Alon, Detailed map of a cis-regulatory input function. Proc. Natl. Acad. Sci. USA 100, 7702–7707 (2003)
H. Maamar, A. RaJ, D. Dubnau, Noise in gene expression determines cell fate in bacillus subtilis. Science 317, 526–529 (2007)
N.G. van Kampen, Stochastic Processes in Physics and Chemistry (North-Holland, Amsterdam, 1985)
D.T. Gillespie, Exact stochastic simulation of coupled chemical reactions. J. Phys. Chem. 81, 2340–2361 (1977)
T. Dingermann, U. Frank-Stoll, H. Werner, A. Wissmann, W. Hillen, M. Jacquet, R. Marschalek, RNA polymerase III catalysed transcription can be regulated in Saccharomyces cerevisiae by the bacterial tetracycline repressor-operator system. EMBO J. 11(4), 1487–1492 (1992)
M. Gossen, H. Bujard, Anhydrotetracycline, a novel effector for tetracycline controlled gene expression systems in eukaryotic cells. Nucleic Acids Res. 21(18), 4411–4412 (1993). https://doi.org/10.1093/nar/21.18.4411
J.A. Magee, S.A. Abdulkadir, J. Milbrandt, Haploinsufficiency at the Nkx 3.1 locus. A paradigm for stochastic, dose-sensitive gene regulation during tumor initiation. Cancer Cell 3, 273–283 (2003)
L.S. Weinberger, J.C. Burnett, J.E. Toettcher, A.P. Arkin, D.V. Schaffer, Stochastic gene expression in a lentiviral positive-feedback loop: HIV-1 tat fluctuations drive phenotypic diversity. Cell 122, 169–182 (2005)
R.D. Dar, N.N. Hosmane, M.R. Arkin, R.F. Siliciano, L.S. Weinberger, Screening for noise in gene expression identifies drug synergies. Science (New York, N.Y.) 344(6190), 1392–1396 (2014)
T.B. Kepler, T.C. Elston, Stochasticity in transcriptional regulation: origins, consequences, and mathematical representations. Biophys. J. 81, 3116–3136 (2001)
A K. Das, work in progress on a “competitive binding of activator-repressor system” (2021–2022)
F.M. Rossi, A.M. Kringstein, A. Spicher, O.M. Guicherit, H.M. Blau, Transcriptional control: rheostat converted to on/off switch. Mol. Cell 6(3), 723–728 (2000)
F. Jiao, C. Zhu, Regulation of gene activation by competitive cross talking pathways. Biophys. J. 119(6), 1204–1214 (2020). https://doi.org/10.1016/j.bpj.2020.08.011
L.H. Smith, P.K. Kitanidis, P.L. McCarty, Numerical modeling and uncertainties in rate coefficients for methane utilization and TCE co-metabolism by a methane-oxidizing mixed culture. Biotechnol. Bioeng. 53, 320–331 (1997)
Acknowledgements
The author would like to acknowledge the helpful suggestions of Dr. Rajesh Karmakar during the initial stage of the project, before he deceased on 09th June 2021. The author also thanks Dr. Indrani Bose and Dr. Arindam Lala for their valuable suggestions and discussions on the paper.
Author information
Authors and Affiliations
Corresponding author
Supplementary Information
Below is the link to the electronic supplementary material.
Appendices
Appendix A
In an attempt to deduce the expressions of mean and the Fano factors, we use a moment generating function which is defined as,
Here, \(i=1,2,......,6\).
we have,
In steady state, \(\frac{\partial F(z_{i},t)}{\partial t}=0\) and for total probability, \(F((z_{i}=1,0)=1\)
Now, by setting [\(\frac{\partial }{\partial z_{1}}(\frac{\partial F}{\partial t})]_{z_{i}=1}=0\), we get \(\frac{\partial F}{\partial z_{1}}=f_{1}(say)= <n_{1}> =\) average number of gene at state \(G_{a}\).
similarly, by setting [\(\frac{\partial }{\partial z_{1}}(\frac{\partial ^{2}F}{\partial z_{1}\partial t})]_{z_{i}=1}=0\) will give \(\frac{\partial ^{2}F}{\partial z_{1}^{2}}=f_{11}(say)\) and so on. Proceeding in the same way, we obtain
and
Appendix B
The parameters used in Eqs. (2), (3) and (4) are given below
Appendix C
Expression for mean mRNA and protein levels for transcription without reinitiation is given by
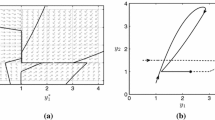
Variation of mean mRNA against GAL keeping aTc = 500 ng ml\(^{-1}\) as parameter in (a) and mean mRNA vs \(J_{m}\) with aTc = 40 ng ml\(^{-1}\) and GAL = 2% as parameter in (b) while other rate constants are chosen from Blake et al. [3]
where \(a_{6}=\left( a_{1}k_{1}+k_{6}k_{8}k_{10}\right) \), \(a_{5}=a_{1}k_{2}+a_{3}k_{2}+a_{2}k_{1}k_{5}+a_{4}\), \(a_{1}=k_{7}k_{9}+k_{6}(k_{8}+k_{9})\), \(a_{2}=k_{7}+k_{8}+k_{9}\), \(a_{3}=k_{6}k_{10}+k_{7}k_{10}+k_{8}k_{10}\), \(a_{4}=k_{5}k_{7}k_{10}+k_{5}k_{8}k_{10}+k_{5}k_{7}k_{9}\),
The expression for the Fano factor at mRNA levels is given by
where \(X=\frac{\begin{array}{ll} &{}g_{22}k_{8}J_{m}^{2}(g_{20}(2g_{19}(g_{1}+k_{m})(g_{3}(k_{m}+k_{1})\\ &{}\quad +k_{1}k_{10})-g_{18}(g_{15}(k_{m}+k_{1})\\ &{}\quad +g_{10}k_{10}))-g_{23}g_{24}) \end{array}}{\begin{array}{ll} &{}(g_{20}J_{m}(-g_{19}(g_{1}+k_{m})(g_{13}-2g_{4}k_{8}J_{m})\\ &{}\quad +(g_{18}J_{m}(-g_{10}(g_{2}+k_{m})-g_{15}k_{1}\\ &{}\quad +g_{25}k_{8})))+g_{20}g_{22}g_{24}k_{m}) \end{array}},\)
The expression of the Fano factor at protein levels is given by
where
Appendix D
In Fig. 17a, b, it is seen that mean mRNA and mean protein in case of transcriptional reinitiation-based network is greater than that of without reinitiation network. But the mean values against other variables like \(J_{m}\) shows that \(m^{WR}\) can be high or less than \(m^{WTR}\) as shown in figure below.
Relative Error for: a fitting of curves when aTc is fixed at 500 ng/ml. Blake et al. [3] model offers MSE = 7.108, while our proposed model fits quite nicely with the experimentally observed data with a minimize MSE = 0.873 b fitting of curves when aTc is fixed at 500 ng/ml. Blake et al. [3] model offers MSE = 2.918, while our proposed model fits better with the experimentally observed data with a reduced MSE = 2.105
Figures 17a and 19a show that \(m^{WR}\) is higher than \(m^{WTR}\) keeping GAL and aTc as parameter, respectively, but Fig. 19b shows a different scenario where we keep both GAL and aTc fixed. It is verified that, the slope of mean mRNA curve becomes high against \(J_{m}\) for a higher concentration of GAL keeping aTc fixed. On the other hand, if we increase aTc for a fixed GAL the slope of the curve goes high against \(J_{m}\). Imposing the condition \(m^{WR}=m^{WTR}\), we have found a critical value of \(J_{m}\) given by
where, \(B_{7}=a_{2}k_{1}k_{5}+a_{4}+b_{11}k_{2}\) and \(k_{r}=k_{4}+k_{11}\) other parameters are supplied earlier.
Appendix E
Model fitting parameters: We proposed an analytical model that fits very much to the experimental data as supplied by Blake et al. [3]. The parameters are chosen from the supplementary material of [3]. The form of parameters \(k_{1}\) and \(k_{2}\) as functions of GAL are determined from an exact analytical treatment as described in the main text. While for the best estimates of other parameters are revised through trial and error to minimize the sum of squared error (SSE) and the mean square error (MSE). We also show the relative percentile error (RE), in Fig. 20a, b for the fitting of each of the data points, which is given by
where, a stands for analytical and e stands for experimental.
Uncertainty in fitted parameters : The uncertainty of the parameter estimates, is generally expressed by the mean square errors, is proportional to the SSE and inversely proportional to the square of the sensitivity coefficient of the model parameters [53]. The mean square fitting error is
n is the number of observations and k is the number of parameters being determined.
The sensitivity (\(\mathscr {S}\)) of a function f(k) over the parameter k is given by
We obtain the sensitivity of the Fano factor (protein) over the fitting parameters and calculate MSE of each parameter keeping others as constant.
where the denominator is the coefficient of sensitivity, squared and summed over all observations.
The GAL is the most sensitive parameter and e is the least in order (see Fig. 21). We found that, the Fano factor is sensitive within a small range of values of these parameters and with the best parameter estimation and minimization of errors (see Fig. 22) support the robustness of our result. The square root of the MSE is the standard deviation, and the approximate 95% confidence interval for k is [53]
\(\kappa \) is the best estimate value of parameter k.
Rights and permissions
About this article
Cite this article
Das, A.K. Stochastic gene transcription with non-competitive transcription regulatory architecture. Eur. Phys. J. E 45, 61 (2022). https://doi.org/10.1140/epje/s10189-022-00213-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1140/epje/s10189-022-00213-2