Abstract
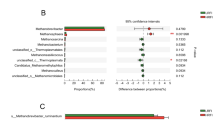
In order to increase productive yields, modulation of rumen fermentation has been a concern in economically relevant species. The ban of antibiotics has driven attention into alternative strategies to modulate ruminal fermentation. Among these, the use of probiotics appears as an interesting approach. The objective of this work was to assess the potential of native bacteria isolated from the rumen of a fed-on-template-pasture cow to modulate fermentation in vitro and to influence the microbiota structure. Seven native ruminal bacteria strains were used in an in vitro gas production experiment. Fermentation dynamics were evaluated, and volatile fatty acids (VFA) and methane were quantified by high-performance liquid chromatography (HPLC) and gas chromatography (GC), respectively. Microbiota structure was assessed by pyrosequencing and methanogens were quantified by quantitative PCR (qPCR). Added strains modulated fermentation dynamics and VFA synthesis. Neither the general structure of the fermenters microbiota, numbers of methanogenic microorganisms nor methane production were altered by added bacteria. However, addition of two strains reduced the volume of gas produced from soluble carbohydrates, while one of them reduced the ratio of gas production in this phase; this was supported by a VFA concentration diminution (4 h incubation) in almost every treatment. Gas produced by fermentation of non-soluble carbohydrates and its fermentation ratio were enhanced by several strains. Also, the abundances of Lachnospiraceae, Veillonellaceae, Rikenellaceae and Succinivibrionaceae were affected by strain supplementation. Modulation of fermentation by selected ruminal native bacteria was achieved, probably enhancing the fermentation of non-soluble carbohydrates. This study represents a new approach in the knowledge related to the use of probiotics in ruminants.

Similar content being viewed by others
References
Allen HK, Levine UY, Looft T, Bandric M, Casey TA (2013) Treatment, promotion, commotion: antibiotic alternatives in food-producing animals. Trends Microbiol 21:114–119
Altschul S, Madden T, Schäfer A, Zhang J, Miller W, Lipman D (1997) Grapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res 25:3389–3402
Calsamiglia S, Castillejos L, Busquet M (2006) Alternatives to antimicrobial growth promoters in cattle. In: Garnsworthy PC, Wiseman J (eds) Recent advances in animal nutrition. University Press Nottingham, Nottingham, pp 129–167
Caporaso JG, Kuczynski J, Stombaugh J, Bittinger K, Bushman FD, Costello EK, Fierer N, González Peña A, Goodrich JK, Gordon JI, Huttley GA, Kelley ST, Knights D, Koenig JE, Ley RE, Lozupone CA, McDonald D, Muegge BD, Pirrung M, Reeder J, Sevinsky JR, Turnbaugh PJ, Walters WA, Widmann J, Yatsunenko T, Zaneveld J, Knight R (2010) QIIME allows analysis of high-throughput community sequencing data. Nat Methods 7:335–336
Castro-Carrera T, Toral PG, Frutos P, McEwan NR, Hervás G, Abecia L, Pinloche E, Girdwood SE, Belenguer A (2014) Rumen bacterial community evaluated by 454 pyrosequencing and terminal restriction fragment length polymorphism analyses in dairy sheep fed marine algae. J Dairy Sci 97:1–9
Chaucheyras-Durand F, Durand H (2010) Probiotics in animal nutrition and health. Benefic Microbes 1:3–9
Cherdthong A, Wanapat M (2013) Manipulation of in vitro ruminal fermentation and digestibility by dried rumen digesta. Livest Sci 153:94–100
Chiquette J, Allison MJ, Rasmussen M (2012) Use of Prevotella bryantii 25A and a commercial probiotic during subacute acidosis challenge in midlactation dairy cows. J Dairy Sci 95:1–11
Denman SE, Tomkins NW, McSweeney CS (2007) Quantitation and diversity analysis of ruminal methanogenic populations in response to the antimethanogenic compound bromochloromethane. FEMS Microbiol Ecol 62:313–322
DeSantis TZ, Hugenholtz P, Larsen N, Rojas M, Brodie EL, Keller K, Huber T, Dalevi D, Hu P, Andersen GL (2006) Greengenes, a Chimera-checked 16S rRNA gene database and workbench compatible with ARB. Appl Environ Microbiol 72:5069–5072
Edgar RC (2010) Search and clustering orders of magnitude faster than BLAST. Bioinformatics 26:2460–2461
Edwards U, Rogall T, Blocker H, Emde M, Bottger EC (1989) Isolation and direct complete nucleotide determination of entire genes. Characterization of a gene coding for 16S ribosomal RNA. Nucleic Acids Res 17:7843–7853
Fierer N, Hamady M, Lauber CL, Knight R (2008) The influence of sex, handedness, and washing on the diversity of hand surface bacteria. Proc Natl Acad Sci U S A 105:17994–17999
Food and Agriculture Organization/World Health Organization (FAO/WHO) (2002) Guidelines for the evaluation of probiotics in food. Report of a Joint FAO/WHO Working Group on Drafting Guidelines for the Evaluation of Probiotics in Food. London. Ontario. Canada. ftp://ftp.fao.org/es/esn/food/wgreport2.pdf. Accessed 23 September 2014
Fraga M, Perelmuter K, Valencia MJ, Cajarville C, Zunino P (2013) Caracterización de la microbiota bacteriana ruminal de un bovino a pastoreo mediante técnicas clásicas e independientes del cultivo. Veterinaria (Montevideo) 49:40–55
Fraga M, Perelmuter K, Valencia MJ, Martínez M, Abin-Carriquiry JA, Cajarville C, Zunino P (2014) Evaluation of native potential probiotic bacteria using an in vitro ruminal fermentation system. Ann Microbiol 64:1149–1156
Hammer Ø, Harper DAT, Ryan PD (2001) PAST: paleontological statistics software package for education and data analysis. Palaeontol Electron 4:9pp
Hungate RE (1950) The anaerobic mesophilic cellulolytic bacteria. Bacteriol Rev 14:1–49
Jami E, Mizrahi I (2012) Composition and similarity of bovine rumen microbiota across individual animals. PLoS One 7:e33306
Jami E, White BA, Mizrahi I (2014) Potential role of the bovine rumen microbiome in modulating milk composition and feed efficiency. PLoS One 9:e85423
Kobayashi Y (2006) Inclusion of novel bacteria in rumen microbiology: need for basic and applied science. Anim Sci 77:375–385
Kung L, Hession AO (1995) Preventing in vitro lactate accumulation in ruminal fermentations by inoculation with Megasphaera elsdenii. J Anim Sci 73:250–256
Lavrenčič A, Levart A, Košir IJ, Cerenak A (2014) In vitro gas production kinetics and short-chain fatty acid production from rumen incubation of diets supplemented with hop cones (Humulua lupulus L.). Animal. doi:10.1017/S1751731114002936, Available on CJO2014
Mauricio RM, Mould FL, Dhanoa MS, Owen E, Channa KS, Theodorou MK (1999) A semi-automated in vitro gas production technique for ruminant feedstuff evaluation. Anim Feed Sci Technol 79:321–330
Mutsvangwa T, Walton JP, Plaisier JC, Duffield TF, Bagg R, Dick P, Vessie G, McBride BW (2002) Effects of a monensin controlled-release capsule or premix on attenuation of subacute ruminal acidosis in dairy cows. J Dairy Sci 85:3454–3461
Nagaraja TG, Avery TB, Bartley EE, Roof SK, Dayton AD (1982) Effect of lasalocid, monensin or thiopeptin on lactic acidosis in cattle. J Anim Sci 54:649–658
Oeztuerk H, Schroeder B, Beyerbach M, Breves G (2005) Influence of living and autoclaved yeasts of Saccharomyces boulardii on in vitro ruminal microbial metabolism. J Dairy Sci 88:2594–2600
Rymer C, Huntington JA, Williams BA, Givens DI (2005) In vitro cumulative gas production techniques: history, methodological considerations and challenges. Anim Feed Sci Technol 123–124:9–30
Schofield P, Pitt RE, Pell AN (1994) Kinetics of fiber digestion from in vitro gas production. J Anim Sci 72:2980–2991
Seo JK, Kim S-W, Kim MH, Upadhaya SD, Kam DK, Ha JK (2010) Direct-fed microbials for ruminant animals. Asian Aust J Anim 23:1657–16679
Shinkai T, Ueki T, Kobayashi Y (2010) Detection and identification of rumen bacteria constituting a fibrolytic consortium dominated by Fibrobacter succinogenes. Anim Sci J 81:72–79
Soccol CR, Vandenberghe LPS, Spier MR, Pedroni Medeiros AB, Tiemi C, De Dea Lindner J, Pandey A, Thomaz-Soccol V (2010) The potential of probiotics: a review. Food Technol Biotechnol 48:413–434
Soto EC, Molina-Alcaide E, Khelil H, Yáñez-Ruiz DR (2013) Ruminal microbiota developing in different in vitro simulation systems inoculated with goats’ rumen liquor. Anim Feed Sci Technol 185:9–18
Stahl DA, Flesher B, Mansfield HR, Montgomery L (1988) Use of phylogenetically based hybridization probes for studies of ruminal microbial ecology. Appl Environ Microbiol 54:1079–1084
Theodorou MK, Williams BA, Dhanoa MS, Mcallan AB, France J (1994) A simple gas production method using a pressure tranducer to determine the fermentation kinetics or rumiant feeds. Anim Feed Sci Technol 48:185–197
Wu S, Baldwin RL 6th, Li W, Li C, Connor EE, Li RW (2012) The bacterial community composition of the bovine rumen detected using pyrosequencing of 16S rRNA genes. Metagenomics 1:1–11
Zened A, Combes S, Cauquil L, Mariette J, Klopp C, Bouchez O, Troegeler-Meynadier A, Enjalbert F (2013) Microbial ecology of the rumen evaluated by 454 GS FLX pyrosequencing is affected by starch and oil supplementation of diets. FEMS Microbiol Ecol 83:504–514
Zhou J, Bruns MA, Tiedje JM (1996) DNA recovery from soils of diverse composition. Appl Environ Microbiol 62:316–322
Acknowledgments
This work was supported by the Agencia Nacional de Investigación e Innovación (ANII), Uruguay, through the projects: ANII FCE 2493 and ANII FMV 2799. Also, Martín Fraga received a PhD fellowship from ANII, Uruguay. Authors wish to thank Luis Vázquez for his assistance during the in vitro experiment.
Conflict of interest
The authors declare no conflict of interests.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Fraga, M., Fernández, S., Cajarville, C. et al. In vitro modulation of rumen microbiota and fermentation by native microorganisms isolated from the rumen of a fed-exclusively-on-pasture bovine. Ann Microbiol 65, 2355–2362 (2015). https://doi.org/10.1007/s13213-015-1077-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13213-015-1077-2