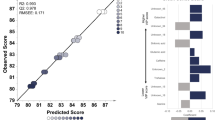
Abstract
Introduction
The consumption of high quality coffee such as specialty Arabica and fine Robusta coffee is increasing steadily in recent years. Development of single origin coffee is an important strategy to maintain coffee quality, grade and high cupping score. Indonesia is a top exporting country for Arabica coffee with high variety of specialty coffees from different origins. Despite its long standing reputation in global coffee market, very few is known about the variability among Indonesian specialty coffees.
Objectives
This study aims to observe metabolite variability among Indonesian coffees from different species and geographical origins by means of non-targeted GC/MS metabolite profiling.
Methods
Sixty-four compounds were tentatively identified from 16 green and roasted coffee beans from different species and cultivation areas in Indonesia and were subjected to principal component analysis (PCA). Ten Specialty Arabica coffee and five Fine Robusta representing all important high quality coffees of Indonesia were also analyzed independently to further classify Indonesian coffee according to their origin.
Results
PCA results of 16 green and roasted coffee beans of different species and cultivation areas showed that samples were separated along PC1 based on different roasting condition (green and roasted) with 52.9% variance and were separated along PC2 based on different species with 19.3% variance. The result from this study showed the clustering of samples based on three major cultivation areas in Indonesia (western, central, eastern part). Metabolites showing higher concentration in Sulawesi, Papua, Flores and Sumatra samples were glycerol, glucuno-1,5-lactone, gluconic acid and sorbitol. A clear distinction in galactitol and galactinol concentration between all samples from eastern part of Indonesia and western and middle part of Indonesia was also observed.
Conclusions
Our results showed that each region (western, central and eastern part of Indonesia) has signature compounds that may serve as discriminant markers for coffee authentication. This is the first report on the classification of Indonesian specialty coffee based on their metabolic profiles and can act as a basis for marker identification for routine procedures in industry.



Similar content being viewed by others
References
Alcázar, A., Fernández-Cáceres, P. L., Martı́n, M. J., Pablos, F., & González, A. G. (2003). Ion chromatographic determination of some organic acids, chloride and phosphate in coffee and tea. Talanta, 61(2), 95–101. https://doi.org/10.1016/s0039-9140(03)00244-3.
Arana, V. A., Medina, J., Alarcon, R., Moreno, E., Heintz, L., Schäfer, H., et al. (2015). Coffee’s country of origin determined by NMR: The Colombian case. Food Chemistry, 175, 500–506. https://doi.org/10.1016/j.foodchem.2014.11.160.
Caporaso, N., Whitworth, M. B., Cui, C., & Fisk, I. D. (2018). Variability of single bean coffee volatile compounds of Arabica and robusta roasted coffees analysed by SPME–GC–MS. Food Research International, 108, 628–640. https://doi.org/10.1016/j.foodres.2018.03.077.
Cevallos-Cevallos, J. M., Reyes-De-Corcuera, J. I., Etxeberria, E., Danyluk, M. D., & Rodrick, G. E. (2009). Metabolomic analysis in food science: A review. Trends in Food Science and Technology, 20(11), 557–566. https://doi.org/10.1016/j.tifs.2009.07.002.
Choi, M.-Y., Choi, W., Park, J. H., Lim, J., & Kwon, S. W. (2010). Determination of coffee origins by integrated metabolomic approach of combining multiple analytical data. Food Chemistry, 121(4), 1260–1268. https://doi.org/10.1016/j.foodchem.2010.01.035.
Consonni, R., Cagliani, L. R., & Cogliati, C. (2012). NMR based geographical characterization of roasted coffee. Talanta, 88, 420–426. https://doi.org/10.1016/j.talanta.2011.11.010.
Creydt, M., & Fischer, M. (2018). Omics approaches for food authentication. Electrophoresis, 39(13), 1569–1581. https://doi.org/10.1002/elps.201800004.
Cubero-Leon, E., Peñalver, R., & Maquet, A. (2014). Review on metabolomics for food authentication. Food Research International, 60, 95–107. https://doi.org/10.1016/j.foodres.2013.11.041.
Danezis, G. P., Tsagkaris, A. S., Camin, F., Brusic, V., & Georgiou, C. A. (2016). Food authentication: Techniques, trends and emerging approaches. Trends in Analytical Chemistry, 85, 123–132. https://doi.org/10.1016/j.trac.2016.02.026.
Dawidowicz, A. L., & Typek, R. (2017). Transformation of chlorogenic acids during the coffee beans roasting process. European Food Research and Technology, 243(3), 379–390. https://doi.org/10.1007/s00217-016-2751-8.
Defernez, M., Wren, E., Watson, A. D., Gunning, Y., Colquhoun, I. J., Le Gall, G., et al. (2017). Low-field 1H NMR spectroscopy for distinguishing between arabica and robusta ground roast coffees. Food Chemistry, 216, 106–113. https://doi.org/10.1016/j.foodchem.2016.08.028.
Edmands, W. M., Ferrari, P., Rothwell, J. A., Rinaldi, S., Slimani, N., Barupal, D. K., et al. (2015). Polyphenol metabolome in human urine and its association with intake of polyphenol-rich foods across European countries. The American Journal of Clinical Nutrition, 102(4), 905–913. https://doi.org/10.3945/ajcn.114.101881.
El-Abassy, R. M., Donfack, P., & Materny, A. (2011). Discrimination between Arabica and Robusta green coffee using visible micro Raman spectroscopy and chemometric analysis. Food Chemistry, 126(3), 1443–1448. https://doi.org/10.1016/j.foodchem.2010.11.132.
Fiehn, O. (2002). Metabolomics—The link between genotypes and phenotypes. Plant Molecular Biology, 48(1), 155–171. https://doi.org/10.1023/a:1013713905833.
Galli, V., & Barbas, C. (2004). Capillary electrophoresis for the analysis of short-chain organic acids in coffee. Journal of Chromatography A, 1032(1), 299–304. https://doi.org/10.1016/j.chroma.2003.09.028.
Garrett, R., Schmidt, E. M., Pereira, L. F. P., Kitzberger, C. S. G., Scholz, M. B. S., Eberlin, M. N., et al. (2013). Discrimination of arabica coffee cultivars by electrospray ionization Fourier transform ion cyclotron resonance mass spectrometry and chemometrics. LWT—Food Science and Technology, 50(2), 496–502. https://doi.org/10.1016/j.lwt.2012.08.016.
Ginz, M., Balzer, H. H., Bradbury, A. G. W., & Maier, H. G. (2000). Formation of aliphatic acids by carbohydrate degradation during roasting of coffee. European Food Research and Technology, 211(6), 404–410. https://doi.org/10.1007/s002170000215.
Hetzel, A., & Coffee Quality Institute. (n.d.). Fine robusta coffee standards and protocols. Coffee Quality Institute. Retrieved December 6, 2018, from https://www.coffeestrategies.com/wp-content/uploads/2015/04/compiled-standards-distribute1.1.pdf.
International Trade Centre, ITC. (2012). Niche markets for coffee: Specialty, environment and social aspects, 52.
Jeszka-Skowron, M., Zgoła-Grześkowiak, A., & Grześkowiak, T. (2015). Analytical methods applied for the characterization and the determination of bioactive compounds in coffee. European Food Research and Technology, 240(1), 19–31. https://doi.org/10.1007/s00217-014-2356-z.
Jumhawan, U., Putri, S. P., Yusianto, Bamba, T., & Fukusaki, E. (2015). Application of gas chromatography/flame ionization detector-based metabolite fingerprinting for authentication of Asian palm civet coffee (Kopi Luwak). Journal of Bioscience and Bioengineering, 120(5), 555–561. https://doi.org/10.1016/j.jbiosc.2015.03.005.
Jumhawan, U., Putri, S. P., Yusianto, Marwani, E., Bamba, T., & Fukusaki, E. (2013). Selection of discriminant markers for authentication of Asian palm civet coffee (Kopi Luwak): A metabolomics approach. Journal of Agricultural and Food Chemistry, 61(33), 7994–8001. https://doi.org/10.1021/jf401819s.
Keidel, A., von Stetten, D., Rodrigues, C., Máguas, C., & Hildebrandt, P. (2010). Discrimination of green arabica and robusta coffee beans by Raman spectroscopy. Journal of Agricultural and Food Chemistry, 58(21), 11187–11192. https://doi.org/10.1021/jf101999c.
Kurzrock, T., & Speer, K. (2001). Identification of kahweol fatty acid esters in Arabica coffee by means of LC/MS. Journal of Separation Science, 24(10–11), 843–848. https://doi.org/10.1002/1615-9314(20011101)24:10/11%3c843:aid-jssc843%3e3.0.co;2-9.
Kwon, D.-J., Jeong, H.-J., Moon, H., Kim, H.-N., Cho, J.-H., Lee, J.-E., et al. (2015). Assessment of green coffee bean metabolites dependent on coffee quality using a 1H NMR-based metabolomics approach. Food Research International, 67, 175–182. https://doi.org/10.1016/j.foodres.2014.11.010.
Lommen, A. (2009). MetAlign: Interface-driven, versatile metabolomics tool for hyphenated full-scan mass spectrometry data preprocessing. Analytical Chemistry, 81(8), 3079–3086. https://doi.org/10.1021/ac900036d.
Martı́n, M. J., Pablos, F., & González, A. G. (1998). Discrimination between arabica and robusta green coffee varieties according to their chemical composition. Talanta, 46(6), 1259–1264. https://doi.org/10.1016/s0039-9140(97)00409-8.
Monakhova, Y. B., Ruge, W., Kuballa, T., Ilse, M., Winkelmann, O., Diehl, B., et al. (2015). Rapid approach to identify the presence of Arabica and Robusta species in coffee using 1H NMR spectroscopy. Food Chemistry, 182, 178–184. https://doi.org/10.1016/j.foodchem.2015.02.132.
Putri, S., & Fukusaki, E. (2018). Metabolomics as a powerful tool for coffee authentication (pp. 337–357). https://doi.org/10.19103/as.2017.0022.19.
Putri, S. P., Nakayama, Y., Matsuda, F., Uchikata, T., Kobayashi, S., Matsubara, A., et al. (2013a). Current metabolomics: Practical applications. Journal of Bioscience and Bioengineering, 115(6), 579–589. https://doi.org/10.1016/j.jbiosc.2012.12.007.
Putri, S. P., Yamamoto, S., Tsugawa, H., & Fukusaki, E. (2013b). Current metabolomics: Technological advances. Journal of Bioscience and Bioengineering, 116(1), 9–16. https://doi.org/10.1016/j.jbiosc.2013.01.004.
Rodrigues, C. I., Maia, R., Miranda, M., Ribeirinho, M., Nogueira, J. M. F., & Máguas, C. (2009). Stable isotope analysis for green coffee bean: A possible method for geographic origin discrimination. Journal of Food Composition and Analysis, 22(5), 463–471. https://doi.org/10.1016/j.jfca.2008.06.010.
Rodrigues, C. I., Marta, L., Maia, R., Miranda, M., Ribeirinho, M., & Máguas, C. (2007). Application of solid-phase extraction to brewed coffee caffeine and organic acid determination by UV/HPLC. Journal of Food Composition and Analysis, 20(5), 440–448. https://doi.org/10.1016/j.jfca.2006.08.005.
Rogers, W. J., Michaux, S., Bastin, M., & Bucheli, P. (1999). Changes to the content of sugars, sugar alcohols, myo-inositol, carboxylic acids and inorganic anions in developing grains from different varieties of Robusta (Coffea canephora) and Arabica (C. arabica) coffees. Plant Science, 149(2), 115–123. https://doi.org/10.1016/s0168-9452(99)00147-8.
Rothwell, J. A., Fillâtre, Y., Martin, J.-F., Lyan, B., Pujos-Guillot, E., Fezeu, L., et al. (2014). New biomarkers of coffee consumption identified by the non-targeted metabolomic profiling of cohort study subjects. PLoS ONE, 9(4), e93474. https://doi.org/10.1371/journal.pone.0093474.
Souard, F., Delporte, C., Stoffelen, P., Thévenot, E. A., Noret, N., Dauvergne, B., et al. (2018). Metabolomics fingerprint of coffee species determined by untargeted-profiling study using LC–HRMS. Food Chemistry, 245, 603–612. https://doi.org/10.1016/j.foodchem.2017.10.022.
Specialty Coffee Association of America. (2015, December 16). SCAA protocols–Cupping specialty coffee. Specialty Coffee Association of America. Retrieved December 6, 2018, from http://www.scaa.org/PDF/resources/cupping-protocols.pdf.
Speer, K., & Mischnick, P. (1989). 16-O-Methylcafestol - ein neues Diterpen im Kaffee Entdeckung und Identifizierung. Zeitschrift für Lebensmittel-Untersuchung und Forschung, 189(3), 219–222. https://doi.org/10.1007/bf01028067.
TPSA Project. (2018, January). Canada–Indonesia Trade and Private Sector Assistance Project. An analysis of the global value chain for Indonesian coffee exports. TPSA. Retrieved December 6, 2018, from http://www.tpsaproject.com/wp-content/uploads/2018-01-30-Report-1211.03c.pdf.
Tsugawa, H., Tsujimoto, Y., Arita, M., Bamba, T., & Fukusaki, E. (2011). GC/MS based metabolomics: Development of a data mining system for metabolite identification by using soft independent modeling of class analogy (SIMCA). BMC Bioinformatics, 12(1), 131. https://doi.org/10.1186/1471-2105-12-131.
van Boekel, M. A. J. S. (2006). Formation of flavour compounds in the Maillard reaction. Biotechnology Advances, 24(2), 230–233. https://doi.org/10.1016/j.biotechadv.2005.11.004.
Weckerle, B., Richling, E., Heinrich, S., & Schreier, P. (2002). Origin assessment of green coffee (Coffea arabica) by multi-element stable isotope analysis of caffeine. Analytical and Bioanalytical Chemistry, 374(5), 886–890. https://doi.org/10.1007/s00216-002-1560-z.
Wei, F., Furihata, K., Koda, M., Hu, F., Kato, R., Miyakawa, T., et al. (2012). 13C NMR-based metabolomics for the classification of green coffee beans according to variety and origin. Journal of Agricultural and Food Chemistry, 60(40), 10118–10125. https://doi.org/10.1021/jf3033057.
Wei, F., Furihata, K., Miyakawa, T., & Tanokura, M. (2014). A pilot study of NMR-based sensory prediction of roasted coffee bean extracts. Food Chemistry, 152, 363–369. https://doi.org/10.1016/j.foodchem.2013.11.161.
Wishart, D. S. (2008). Metabolomics: Applications to food science and nutrition research. Trends in Food Science and Technology, 19(9), 482–493. https://doi.org/10.1016/j.tifs.2008.03.003.
Zhang, C., Wang, C., Liu, F., & He, Y. (2016). Mid-infrared spectroscopy for coffee variety identification: Comparison of pattern recognition methods. Journal of Spectroscopy, 2016, 1–7. https://doi.org/10.1155/2016/7927286.
Acknowledgements
The authors thank Indonesian National Commission for UNESCO and L’Oreal Indonesia for the L’Oreal Award for Women in Science funding received by Sastia P. Putri and to Dr. Agung Wahyu Susilo and Dr. Sukrisno Widyotomo of Indonesian Coffee and Cocoa Research Institute for their kind support in this study. The authors also thank Mr. Abu Hanifah for his technical assistance in this study.
Author information
Authors and Affiliations
Contributions
Sastia Prama Putri and Eiichiro Fukusaki conceived and designed the study. Tomoya Irifune and Yusianto performed the experiments and analyzed the data. Sastia Prama Putri wrote the paper. Tomoya Irifune, Yusianto, and Eiichiro Fukusaki revised the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Research involving human and animal rights
This article does not contain any studies with human participants or animal performed by any of the authors.
Informed consent
This article does not involve human participants.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Putri, S.P., Irifune, T., Yusianto et al. GC/MS based metabolite profiling of Indonesian specialty coffee from different species and geographical origin. Metabolomics 15, 126 (2019). https://doi.org/10.1007/s11306-019-1591-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11306-019-1591-5