Abstract
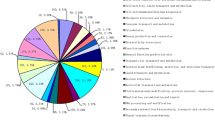
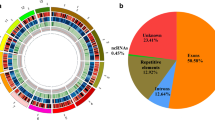
The first full-length cDNA library for lichenized fungi was constructed from cultured mycobiont of the arid desert lichen Endocarpon pusillum. Based on small-scale sequencing results, 111 genes of the lichenized fungi were identified for the first time, among which 11 genes shared no homology with any known fungal genes. Real-time PCR showed that the size of the mycobiont genome is 39.13 Mb and the copy number of ribosomal RNA gene repeat units is 43. The results of this study will be valuable for the ongoing lichen genome-sequencing project and the large-scale identification of functional genes from lichenized fungi.


Similar content being viewed by others
References
Armaleo D, May S (2009) Sizing the fungal and algal genomes of the lichen Cladonia grayi through quantitative PCR. Symbiosis 49:43–51
Belkhiri A, Klassen GR (1996) Diverged 5 s rRNA sequences adjacent to 5 s rRNA genes in the rDNA of Pythium pachycaule. Curr Genet 29:287–292
Brink JJ, Ahmadjian V, Fröberg L, Goldsmith S (1995) Time course of RNA accumulation in cultures of lichen fungi. Cryptogam Bot 5:55–59
Buschbom J (2007) Migration between continents: geographical structure and long-distance gene flow in Porpidia flavicunda (lichen-forming Ascomycota). Mol Ecol 16:1835–1846
Chooi YH, Stalker DM, Davis MA, Fujii I, Elix JA, Louwhoff SH, Lawrie AC (2008) Cloning and sequence characterization of a non-reducing polyketide synthase gene from the lichen Xanthoparmelia semiviridis. Mycol Res 112:147–161
Clarke JD (2009) Cetyltrimenthyl ammonium bromide (CTAB) DNA miniprep for plant DNA isolation. Cold Spring Harb Protoc doi:10.1101/pdb.prot5177
Corradi N, Croll D, Colard A, Kuhn G, Ehinger M, Sanders IR (2007) Gene copy number polymorphisms in an arbuscular mycorrhizal fungal population. Appl Environ Microb 73:366–369
DePriest PT (1994) Variation in the Cladonia chlorophaea complex II: ribosomal DNA variation in a Southern Appalachian population. Bryologist 97:117–126
Fournier P, Gaillardin C, Persuy MA, Klootwijk J, Heerikhuizen H (1986) Heterogeneity in the ribosomal family of the yeast Yarrowia lipolytica: genomic organization and segregation studies. Gene 42:273–282
Gagunashvili AN, Davidsson SP, Jonsson ZO, Andresson OS (2009) Cloning and heterologous transcription of a polyketide synthase gene from the lichen Solorina crocea. Mycol Res 113:354–363
Garber RC, Turgeon BG, Selker EU, Yoder OC (1988) Organization of ribosomal RNA genes in the fungus Cochliobolus heterostrophus. Curr Genet 14:573–582
Gutierrez G, Blanco O, Divakar PK, Lumbsch HT, Crespo A (2007) Patterns of group I intron presence in nuclear SSU rDNA of the lichen family Parmeliaceae. J Mol Evol 64:181–195
Hayes CK, Klemsdal S, Lorito M, Pietro AD, Peterbauer C, Nakas JP, Tronsmo A, Harman GE (1994) Isolation and sequence of an endochitinase-encoding gene from a cDNA library of Trichoderma harzianum. Gene 138:143–148
Herrera ML, Vallor AC, Gelfond JA, Patterson TF, Wickes BL (2009) Strain-dependent variation in 18S ribosomal DNA copy numbers in Aspergillus fumigatus. J Clin Microbiol 47:1325–1332
Hibbett DS, Binder M, Bischoff JF et al (2007) A higher-level phylogenetic classification of the fungi. Mycol Res 111:509–547
Honegger R (2003) The impact of different long-term storage conditions on the viability of lichen-forming Ascomycees and their green algal photobiont, Trebouxia spp. Plant Biol 5:324–330
Howlett BJ, Rolls BD, Cozijnsen AJ (1997) Organisation of ribosomal DNA in the ascomycete Leptosphaeria maculans. Microbiol Res 152:261–267
Ilse K, Richard B, Ayala H, Thomas H, Nash III (2008) Desiccation-tolerance in lichens: a review. Bryologist 111:576–593
James TY, Kauff F, Conrad L et al (2006) Reconstructing the early evolution of fungi using a six-gene phylogeny. Nature 443:818–822
Joneson S, Armaleo D, Lutzoni F (2011) Fungal and algal gene expression in early developmental stages of lichen-symbiosis. Mycologia 103:291–306
Junttila S, Lim KJ, Rudd S (2009) Optimization and comparison of different methods for RNA isolation for cDNA library construction from the reindeer lichen Cladonia rangiferina. BMC Res Notes 2:204–208
Lindblom L, Ekman S (2006) Genetic variation and population differentiation in the lichen-forming ascomycete Xanthoria parietina on the island Storfosna, central Norway. Mol Ecol 15:1545–1559
Logemann J, Schell J, Willmitzer L (1987) Improved method for the isolation of RNA from plant tissues. Anal Biochem 163:16–20
Lucas MC, Jacobson JW, Giles NH (1977) Characterization and in vitro translation of polyadenylated messenger ribonucleic acid from Neurospora crassa. J Bacteriol 130:1192–1198
Maleszka R, Clark-Walker GD (1990) Magnification of the rDNA cluster in Kluyveromyces lactis. Mol Gen Genet 223:342–344
Maleszka R, Clark-Walker GD (1993) Yeasts have a four-fold variation in ribosomal DNA copy number. Yeast 9:53–58
McNeil M, Darvill AG, Fry SC, Albersheim P (1984) Structure and function of the primary cell walls of plants. Annu Rev Biochemistry 53:625–663
Michael A, Connolly, Peter A, Clausen, James G, Lazar (2006) Preparation of RNA from plant tissue using trizol. Cold Spring Harb Protoc doi:10.1101/pdb.prot4105
Palice Z, Schmitt I, Lumbsch HT (2005) Molecular data confirm that Omphalina foliacea is a lichen-forming basidiomycete. Mycol Res 109:447–451
Reeb V, Lutzoni F, Roux C (2004) Contribution of RPB2 to multilocus phylogenetic studies of the euascomycetes (Pezizomycotina, Fungi) with special emphasis on the lichen-forming Acarosporaceae and evolution of polyspory. Mol Phylogenet Evol 32:1036–1060
Scherrer S, Haisch A, Honegger R (2002) Characterization and expression of XPH1, the hydrophobin gene of the lichen-forming ascomycete Xanthoria parietina. New Phytol 154:175–184
Sinnemann SJ, Andrésson óS, Brown DW, Miao VPW (2000) Cloning and heterologous expression of Solorina crocea pyrG. Curr Genet 37:333–338
Staiger B, Kalb K, Grube M (2006) Phylogeny and phenotypic variation in the lichen family Graphidaceae (Ostropomycetidae, Ascomycota). Mycol Res 110:765–772
Sturani E, Costantini MG, Martegani E, Alberghina L (1979) Level and turnover of polyadenylate-containing ribonucleic acid in Neurospora crassa in different steady states of growth. Eur J Biochem 99:1–7
Valarmathi R, Hariharan GN, Venkataraman G, Parida A (2009) Characterization of a non-reducing polyketide synthase gene from lichen Dirinaria applanata. Phytochemistry 70:721–729
Walser JC, Holderegger R, Gugerli F, Hoebee SE, Scheidegger C (2005) Microsatellites reveal regional population differentiation and isolation in Lobaria pulmonaria, an epiphytic lichen. Mol Ecol 14:457–467
White TJ, Bruns T, Lee S, Taylor J (1990) Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. In: Innis M, Gelfand D, Sninsky J, White T (eds) PCR protocols: a guide to methods and applications. Academic Press, San Diego, CA, pp 315–322
Wilhelm J, Pingoud A, Hahn M (2003) Real-time PCR-based method for the estimation of genome sizes. Nucleic Acids Res 31:e56
Yoshimura I, Yamamoto Y, Nakano T, Finnie J (2002) Isolation and culture of lichen photobionts and mycobionts. In: Kranner I, Beckett RP, Varma AK (eds) Protocols in lichenology. Culturing, biochemistry, ecophysiology and use in biomonitoring. Springer, Berlin, pp 3–33
Zhang CS, Cao YQ, Wang ZK, Yin YQ, Peng GX, Xia YX (2007) A method to construct cDNA library of the entomopathogenic fungus, Metarhizium anisopliae, in the hemolymph of the infected locust. Mol Biotechnol 36:23–31
Zhou QM, Guo SY, Huang MR, Wei JC (2006) A study of the genetic variability of Rhizoplaca chrysoleuca using DNA sequences and secondary metabolic substances. Mycologia 98:57–67
Acknowledgments
We are indebted to Dr. Bin Liu and Miss Dan Guo (College of life science, Nankai University) for proving us the accurate concentration of nucleic acids using Qubit system. The authors would like to thank Professor Chuan-Peng Liu and an anonymous reviewer for their critical reading of this paper and valuable suggestions. The present study was supported in part by the Ministry of Science and Technology of the People’s Republic of China (MOST, No. 2007AA021405) and the Knowledge Innovation Program of Chinese Academy of Sciences (KSCX2-YW-Z-1020, 2010-Biols-CAS-0104).
Author information
Authors and Affiliations
Corresponding authors
Rights and permissions
About this article
Cite this article
Wang, YY., Zhang, T., Zhou, QM. et al. Construction and characterization of a full-length cDNA library from mycobiont of Endocarpon pusillum (lichen-forming Ascomycota). World J Microbiol Biotechnol 27, 2873–2884 (2011). https://doi.org/10.1007/s11274-011-0768-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11274-011-0768-5