Abstract
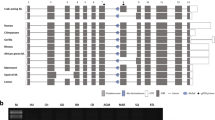
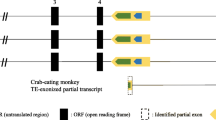
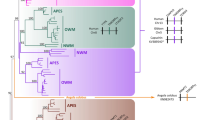
The leptin receptor (LEPR) is a crucial regulatory protein that interacts with Leptin. In our analysis of LEPR, novel AluJb-derived alternative transcripts were identified in the genome of the rhesus monkey. In order to investigate the occurrence of AluJb-derived alternative transcripts and the mechanism underlying exonization events, we conducted analyses using a number of primate genomic DNAs and adipose RNAs of tissue and primary cells derived from the crab-eating monkey. Our results demonstrate that the AluJb element has been integrated into our common ancestor genome prior to the divergence of simians and prosimians. The lineage-specific exonization event of the LEPR gene in chimpanzees, orangutans, and Old World monkeys appear to have been accomplished via transition mutations of the 5′ splicing site (second position of C to T). However, in New World monkeys and prosimians, the AluJb-related LEPR transcript should be silenced by the additional transversion mutation (fourth position of T to G). The AluJb-related transcript of human LEPR should also be silenced by a mutation of the 5′ splicing site (first position of G to A) and the insertion of one nucleotide sequence (minus fourth position of A). Our data suggests that lineage-specific exonization events should be determined by the combination event of the formation of splicing sites and protection against site-specific mutation pressures. These evolutionary mechanisms could be major sources for primate diversification.
Similar content being viewed by others
References
Ast, G. (2004). How did alternative splicing evolve? Nat. Rev. Genet. 5, 773–782.
Batzer, M.A., and Deininger, P.L. (2002). Alu repeats and human genomic diversity. Nat. Rev. Genet. 3, 370–379.
Costas, J. (2003). Molecular characterization of the recent intragenomic spread of the murine endogenous retrovirus MuERV-L. J. Mol. Evol. 56, 181–186.
Damert, A., Lower, J., and Lower, R. (2004). Leptin receptor isoform 219.1: an example of protein evolution by LINE-1-mediated human-specific retrotransposition of a coding SVA element. Mol. Biol. Evol. 21, 647–651.
Friedman, J.M., and Halaas, J.L. (1998). Leptin and the regulation of body weight in mammals. Nature 395, 763–770.
Fruhbeck, G. (2006). Intracellular signalling pathways activated by leptin. Biochem. J. 393, 7–20.
Han, K., Konkel, M.K., Xing, J., Wang, H., Lee, J., Meyer, T.J., Huang, C.T., Sandifer, E., Hebert, K., Barnes, E.W., et al. (2007). Mobile DNA in Old World monkeys: a glimpse through the rhesus macaque genome. Science 316, 238–240.
Huh, J.W., Kim, T.H., Yi, J.M., Park, E.S., Kim, W.Y., Sin, H.S., Kim, D.S., Min, D.S., Kim, S.S., Kim, C.B., et al. (2006). Molecular evolution of the periphilin gene in relation to human endogenous retrovirus m element. J. Mol. Evol. 62, 730–737.
Huh, J.W., Ha, H.S., Kim, D.S., and Kim, H.S. (2008). Placenta-restricted expression of LTR-derived NOS3. Placenta 29, 602–608.
International Human Genome Sequencing Consortium (2001). Initial sequencing and analysis of the human genome. Nature 409, 860–921.
Jurka, J. (2000). Repbase update: a database and an electronic journal of repetitive elements. Trends Genet. 16, 418–420.
Kreahling, J., and Graveley, B.R. (2004). The origins and implications of alternative splicing. Trends Genet. 20, 1–4.
Krull, M., Brosius, J., and Schmitz, J. (2005). Alu-SINE exonization: en route to protein-coding function. Mol. Biol. Evol. 22, 1702–1711.
Lev-Maor, G., Sorek, R., and Ast, G. (2003). The birth of an alternatively spliced exon: 3′ splice-site selection in Alu exons. Science 300, 1288–1291.
Li, W.H., Gu, Z., Wang, H., and Nekrutenko, A. (2001). Evolutionary analyses of the human genome. Nature 409, 847–849.
Lin, L., Shen, S., Tye, A., Cai, J.J., Jiang, P., Davidson, B.L., and Xing, Y. (2008). Diverse splicing patterns of exonized Alu elements in human tissues. PLoS Genet. 17, e1000225.
Lord, G.M., Matarese, G., Howard, J.K., Baker, R.J., Bloom, S.R., and Lechler, R.I. (1998). Leptin modulates the T-cell immune response and reverses starvation-induced immunosuppression. Nature 394, 897–901.
Piriyapongsa, J., Polavarapu, N., Borodovsky, M., and McDonald, J. (2007). Exonization of the LTR transposable elements in human genome. BMC Genomics 8, 291.
Ram, O., Schwartz, S., and Ast, G. (2008). Multifactorial interplay controls the splicing profile of Alu-derived exons. Mol. Cell. Biol. 28, 3513–3525.
Sela, N., Mersch, B., Gal-Mark, N., Lev-Maor, G., Hots-Wagenblatt, A., and Ast, G. (2007). Comparative analysis of transposed element insertion within human and mouse genes reveals Alu’s unique role in shaping the human transcriptome. Genome Biol. 8, R127.
Singer, S.S., Männel, D.N., Hehlgans, T., Brosius, J., and Schmitz, J. (2004). From “junk” to gene: curriculum vitae of a primate receptor isoform gene. J. Mol. Biol. 20, 883–886.
Sorek, R., Ast, G., and Graur, D. (2002). Alu-containing exons are alternatively spliced. Genome Res. 12, 1060–1067.
Sorek, R., Lev-Maor, G., Reznik, M., Dagan, T., Belinky, F., Graur, D., and Ast, G. (2004). Minimal conditions for exonization of intronic sequences: 5′ splice site formation in alu exons. Mol. Cell 14, 221–231.
Sverdlov, E.D. (1998). Perpetually mobile footprints of ancient infections in human genome. FEBS Lett. 428, 1–6.
The Chimpanzee Sequencing and Analysis Consortium (2005). Initial sequencing of the chimpanzee genome and comparison with the human genome. Nature 437, 69–87.
van de Lagemaat, L.N., Landry, J.R., Mager, D.L., and Medstrand, P. (2003). Transposable elements in mammals promote regulatory variation and diversification of genes with specialized functions. Trends Genet. 19, 530–536.
Van Vugt, D.A., Lujan, M.E., Froats, M., Krzemien, A., Couceyro, P.R., and Reid, R.L. (2006). Effect of fasting on cocaine-amphetamine-regulated transcript, neuropeptide Y, and leptin receptor expression in the non-human primate hypothalamus. Neuroendocrinology 84, 83–93.
Xiao, R., Park, K., Oh, Y., Kim, J., and Park, C. (2008). Structural characterization of the genome of BERV gamma4, the most abundant endogenous retrovirus family in cattle. Mol. Cells 26, 404–408.
Author information
Authors and Affiliations
Corresponding author
Additional information
These authors contributed equally to this work.
About this article
Cite this article
Huh, JW., Kim, YH., Kim, DS. et al. Alu-derived old world monkeys exonization event and experimental validation of the LEPR gene. Mol Cells 30, 201–207 (2010). https://doi.org/10.1007/s10059-010-0108-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10059-010-0108-x