Abstract
Objective
Osteoporosis is hard to detect before it manifests symptoms and complications. In this study, we evaluated machine learning models for identifying individuals with abnormal bone mineral density (BMD) through an analysis of spine X-ray features extracted by deep learning to alert high-risk osteoporosis populations.
Materials and methods
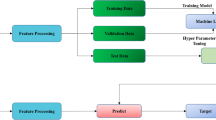
We retrospectively used data obtained from health check-ups including spine X-ray and dual-energy X-ray absorptiometry (DXA). Consecutively, we selected people with normal and abnormal bone mineral density. From the regions of interest of X-ray images, deep convolutional networks were used to generate image features. We designed prediction models for abnormal BMD using the image features trained by machine learning classification algorithms. The performances of each model were evaluated.
Results
From 334 participants, 170 images of abnormal (T scores < − 1.0 standard deviations (SD)) and 164 of normal BMD (T scores > = − 1.0 SD) were used for analysis. We found that a combination of feature extraction by VGGnet and classification by random forest based on the maximum balanced classification rate (BCR) yielded the best performance in terms of the area under the curve (AUC) (0.74), accuracy (0.71), sensitivity (0.81), specificity (0.60), BCR (0.70), and F1-score (0.73).
Conclusion
In this study, we explored various machine learning algorithms for the prediction of BMD using simple spine X-ray image features extracted by three deep learning algorithms. We identified the combination for the best performance in predicting high-risk populations with abnormal BMD.

Similar content being viewed by others
References
Yang J, Pham SM, Crabbe DL. Effects of oestrogen deficiency on rat mandibular and tibial microarchitecture. Dentomaxillofac Radiol. 2003;32(4):247–51.
Lee JJ, Aghdassi E, Cheung AM, et al. Ten-year absolute fracture risk and hip bone strength in Canadian women with systemic lupus erythematosus. J Rheumatol. 2012;39(7):1378–84.
Kanis JA. Diagnosis of osteoporosis and assessment of fracture risk. Lancet. 2002;359(9321):1929–36.
Demontiero O, Vidal C, Duque G. Aging and bone loss: new insights for the clinician. Ther Adv Musculoskelet Dis. 2012;4(2):61–76.
Lee EY, Kim HC, Rhee Y, et al. The Korean urban rural elderly cohort study: study design and protocol. BMC Geriatr. 2014;14:33.
Ha YC, Kim HY, Jang S, Lee YK, Kim TY. Economic burden of osteoporosis in South Korea: claim data of the national health insurance service from 2008 to 2011. Calcif Tissue Int. 2017;101(6):623–30.
Rehman DE, Qureshi S, Haq AA. Early detection of osteoporosis from incisure depth of human mandible in an orthopantomogram. J Pak Med Assoc. 2014;64(7):766–9.
Choi YJ, Oh HJ, Kim DJ, et al. The prevalence of osteoporosis in Korean adults aged 50 years or older and the higher diagnosis rates in women who were beneficiaries of a national screening program: the Korea National Health and Nutrition Examination Survey 2008–2009. J Bone Miner Res. 2012;27(9):1879–86.
US Preventive Services Task Force, Curry SJ, Krist AH, et al. Screening for osteoporosis to prevent fractures: US preventive services task force recommendation statement. JAMA. 2018;319(24):2521–31.
Brown C. Osteoporosis: staying strong. Nature. 2017;550(7674):S15–7.
Lee CH, Chung CK, Kim CH, et al. Health care burden of spinal diseases in the Republic of Korea: analysis of a nationwide database from 2012 through 2016. Neurospine. 2018;15(1):66–76.
Tannor AY. Lumbar spine X-Ray as a standard investigation for all low back pain in Ghana: is it evidence based? Ghana Med J. 2017;51(1):24–9.
Lee C, Choe EK, Choi JM, et al. Health and prevention enhancement (H-PEACE): a retrospective, population-based cohort study conducted at the Seoul national university hospital Gangnam center. Korea BMJ Open. 2018;8(4):e019327.
Kanis JA. Assessment of fracture risk and its application to screening for postmenopausal osteoporosis: synopsis of a WHO report. WHO Study Group Osteoporos Int. 1994;4(6):368–81.
Krizhevsky A, Sutskever I, Hinton GE. ImageNet classification with deep convolutional neural networks. In: NIPS’12 proceedings of the 25th international conference on neural information processing systems. New York, NY: ACM; 2012. p. 1097–105.
Simonyan K, Zisserman A. Very deep convolutional networks for large-scale image recognition. In: The 3rd international conference on learning representations 2015 (ICLR2015). San Diego, CA, USA; 2015. p. 1–14.
Szegedy C, Liu W, Jia Y, et al. Going deeper with convolutions. In: IEEE conference on computer vision and pattern recognition. Boston, MA: IEEE; 2015. p. 1–9.
Szegedy C, Vanhoucke V, Ioffe S, Shlens J. Rethinking the inception architecture for computer vision. In: 2016 conference on Computer Vision and Pattern Recognition (CVPR). Las Vegas, NV, USA; 2016. p. 2818–26.
He K, Zhang X, Ren S, Sun J. Deep residual learning for image recognition. In: 2016 IEEE conference on Computer Vision and Pattern Recognition (CVPR). Las Vegas, NV: IEEE; 2016. p. 770–8.
Jorrissen RN, Gilson MK. Virtual screening of molecular databases using a support vector machine. J Chem Inf Model. 2005;45(3):549–61.
Wu X, Kumar V, Quinlan JR, et al. Top 10 algorithms in data mining. Knowl Inf Syst. 2008;14(1):1–37.
Plewczynski D, von Grotthuss M, Rychlewski L, Ginalski K. Virtual high throughput screening using combined random forest and flexible docking. Comb Chem High Throughput Screen. 2009;12(5):484–9.
Ho TK. Random decision forests. In: Proceedings of 3rd international conference on document analysis and recognition. Montreal, Quebec: IEEE; 1995. p. 278–82.
Ho TK. The random subspace method for constructing decision forests. IEEE Trans Pattern Anal Mach Intell. 1998;20(8):832–44.
Menze BH, Kelm BM, Masuch R, et al. A comparison of random forest and its gini importance with standard chemometric methods for the feature selection and classification of spectral data. BMC Bioinformatics. 2009;10:213.
Calle ML, Urrea V, Boulesteix AL, Malats N. AUC-RF: a new strategy for genomic profiling with random forest. Hum Hered. 2011;72(2):121–32.
Chen X, Wang MH, Zhang HP. The use of classification trees for bioinformatics. Wiley Interdiscip Rev Data Min Knowl Discov. 2011;1(1):55–63.
Casanova R, Saldana S, Chew EY, Danis RP, Greven CM, Ambrosius WT. Application of random forests methods to diabetic retinopathy classification analyses. PLoS One. 2014;9(6):e98587.
Kahn S, Rahmani H, Shah SAA, Bennamoun M, Medioni G, Dickinson S. A guide to convolutional neural networks for computer vision. In: Medioni G, Dickinson S, editors. Synthesis lectures on computer vision. San Rafael, California: Morgan & Claypool; 2018. p. 104.
Lim HK, Ha HI, Park SY, Lee K. Comparison of the diagnostic performance of CT hounsfield unit histogram analysis and dual-energy X-ray absorptiometry in predicting osteoporosis of the femur. Eur Radiol. 2019;29(4):1831–40.
https://www.iscd.org/official-positions/2019-iscdofficial-positions-adult/
Yu TY, Cho H, Kim TY, et al. Utilization of osteoporosis-related health services: use of data from the Korean National Health Insurance Database 2008-2012. J Korean Med Sci. 2018;33(3):e20.
Funding
This study received funding from the Seoul National University Hospital Research Fund, grant number 0420170720.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
The study was conducted in accordance with the Declaration of Helsinki.
Conflict of interest
The authors declare that they have no conflicts of interest.
Informed consent
Informed consent was waived by the IRB.
Ethical approval
The Institutional Review Board (IRB) of the Seoul National University Hospital approved the study protocol (IRB number 1808-008-962).
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Lee, S., Choe, E.K., Kang, H.Y. et al. The exploration of feature extraction and machine learning for predicting bone density from simple spine X-ray images in a Korean population. Skeletal Radiol 49, 613–618 (2020). https://doi.org/10.1007/s00256-019-03342-6
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00256-019-03342-6