Abstract
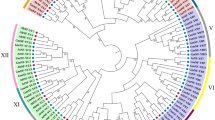
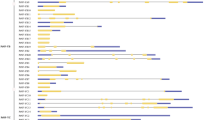
Some members of the nuclear transcription factor Y subunit B (NF-YB) have been newly reported to regulate a variety of functions in annual plant development and drought tolerance. The overall goal of this study was to identify and characterize the NF-YB genes in the model tree Populus trichocarpa, with the expectation of determining which orthologs in P. euphratica, naturally distributed in semiarid regions, were involved in drought response. A total of 20 NF-YB genes in the P. trichocarpa genome were identified and annotated. The examination on their phylogenetic relationships and ortholog predictions showed that Populus has an expanded set of NF-YB genes within unknown functional groups in comparison to Arabidopsis. Four members of the PeNF-YB gene family in P. euphratica were found to be up-regulated in expression during PEG-6000 (polyethylene glycol 6000) drought treatment based on RT-qPCR analyses. The increased knowledge on the phylogenetic relationships, predicted orthologs, and expression patterns of the poplar NF-YB genes during drought stress can be expected to promote the future study of woody plants.



Similar content being viewed by others
References
Bogeat-Triboulot M-B, Brosché M, Renaut J, Jouve L, Le Thiec D, Fayyaz P, Vinocur B, Witters E, Laukens K, Teichmann T, Altman A, Hausman J-F, Polle A, Kangasjärvi J, Dreyer E (2007) Gradual soil water depletion results in reversible changes of gene expression, protein profiles, ecophysiology, and growth performance in Populus euphratica, a poplar growing in arid regions. Plant Physiol 143(2):876–892. doi:10.1104/pp.106.088708
Brosche M, Vinocur B, Alatalo E, Lamminmaki A, Teichmann T, Ottow E, Djilianov D, Afif D, Bogeat-Triboulot M-B, Altman A, Polle A, Dreyer E, Rudd S, Paulin L, Auvinen P, Kangasjarvi J (2005) Gene expression and metabolite profiling of Populus euphratica growing in the Negev desert. Genome Biol 6(12):R101
Cao S, Kumimoto RW, Siriwardana CL, Risinger JR, Holt BF III (2011) Identification and characterization of NF-Y transcription factor families in the monocot model plant Brachypodium distachyon. PLoS One 6(6):e21805
Caruso A, Morabito D, Delmotte F, Kahlem G, Carpin S (2002) Dehydrin induction during drought and osmotic stress in Populus. Plant Physiol Biochem 40:1033–1042
Caruso A, Chefdor F, Carpin S, Depierreux C, Delmotte FM, Kahlem G, Morabito D (2008) Physiological characterization and identification of genes differentially expressed in response to drought induced by PEG 6000 in Populus canadensis leaves. J Plant Physiol 165(9):932–941. doi:10.1016/j.jplph.2007.04.006
Ceribelli M, Dolfini D, Merico D, Gatta R, Viganò AM, Pavesi G, Mantovani R (2008) The histone-like NF-Y is a bifunctional transcription factor. Mol Cell Biol 28(6):2047–2058. doi:10.1128/mcb.01861-07
Chang S, Puryear J, Cairney J (1993) A simple and efficient method for isolating RNA from pine trees. Plant Mol Biol Rep 11(2):113–116. doi:10.1007/bf02670468
Dolfini D, Zambelli F, Pavesi G, Mantovani R (2009) A perspective of promoter architecture from the CCAAT box. Cell Cycle 8(24):4127–4137
Dolfini D, Gatta R, Mantovani R (2012) NF-Y and the transcriptional activation of CCAAT promoters. Crit Rev Biochem Mol Biol 47(1):29–49. doi:10.3109/10409238.2011.628970
Ferreira S, Hjerno K, Larsen M, Wingsle G, Larsen P, Fey S, Roepstorff P, Salomea Pais M (2006) Proteome profiling of Populus euphratica Oliv. upon heat stress. Ann Bot 98(2):361–377. doi:10.1093/aob/mcl106
Gao A, Zhu Q, Chen X, Luo J (2007) A gene structure display server. Yi Chuan 29(8):1023–1026. doi:10.1371/journal.pone.0021805
Gray J, Bevan M, Brutnell T, Buell CR, Cone K, Hake S, Jackson D, Kellogg E, Lawrence C, McCouch S, Mockler T, Moose S, Paterson A, Peterson T, Rokshar D, Souza GM, Springer N, Stein N, Timmermans M, Wang G-L, Grotewold E (2009) A Recommendation for naming transcription factor proteins in the grasses. Plant Physiol 149(1):4–6. doi:10.1104/pp.108.128504
Hackenberg D, Wu Y, Voigt A, Adams R, Schramm P, Grimm B (2011) Studies on differential nuclear translocation mechanism and assembly of the three subunits of the Arabidopsis thaliana transcription factor NF-Y. Mol Plant. doi:10.1093/mp/ssr107
Hackenberg D, Keetman U, Grimm B (2012) Homologous NF-YC2 subunit from Arabidopsis and tobacco is activated by photooxidative stress and induces flowering. Int J Mol Sci 13(3):3458–3477. doi:10.3390/ijms13033458
Han XJ, He G, Zhao ST, Guo CH, Lu MZ (2012) Expression analysis of two NAC transcription factors PtNAC068 and PtNAC154 from poplar. Plant Mol Biol Rep 30(2):370–378. doi:10.1007/s11105-011-0350-1
Ito Y, Thirumurugan T, Serizawa A, Hiratsu K, Ohme-Takagi M, Kurata N (2011) Aberrant vegetative and reproductive development by overexpression and lethality by silencing of OsHAP3E in rice. Plant Sci 181(2):105–110. doi:10.1016/j.plantsci.2011.04.009
Kumimoto RW, Adam L, Hymus GJ, Repetti PP, Reuber TL, Marion CM, Hempel FD, Ratcliffe OJ (2008) The nuclear factor Y subunits NF-YB2 and NF-YB3 play additive roles in the promotion of flowering by inductive long-day photoperiods in Arabidopsis. Planta 228(5):709–723. doi:10.1007/s00425-008-0773-6
Kumimoto RW, Zhang Y, Siefers N, Holt BF (2010) NF-YC3, NF-YC4 and NF-YC9 are required for CONSTANS-mediated, photoperiod-dependent flowering in Arabidopsis thaliana. Plant J 63(3):379–391. doi:10.1111/j.1365-313X.2010.04247.x
Kwong RW, Bui AQ, Lee H, Kwong LW, Fischer RL, Goldberg RB, Harada JJ (2003) Leafy cotyledon1-like defines a class of regulators essential for embryo development. Plant Cell 15(1):5–18. doi:10.1105/tpc.006973
Li X-Y, Mantovani R, Hooft van Huijsduijnen R, Andre I, Benoist C, Mathis D (1992) Evolutionary variation of the CCAAT-binding transcription factor NF-Y. Nucleic Acids Res 20(5):1087–1091. doi:10.1093/nar/20.5.1087
Li WX, Oono Y, Zhu JH, He XJ, Wu JM, Iida K, Lu XY, Cui XP, Jin HL, Zhu JK (2008) The Arabidopsis NFYA5 transcription factor is regulated transcriptionally and posttranscriptionally to promote drought resistance. Plant Cell 20(8):2238–2251. doi:10.1105/tpc.108.059444
Liu J-X, Howell SH (2010) bZIP28 and NF-Y transcription factors are activated by ER stress and assemble into a transcriptional complex to regulate stress response genes in Arabidopsis. The Plant Cell 22:782–796
Mantovani R (1998) A survey of 178 NF-Y binding CCAAT boxes. Nucleic Acids Res 26(5):1135–1143. doi:10.1093/nar/26.5.1135
Mantovani R (1999) The molecular biology of the CCAAT-binding factor NF-Y. Gene 239(1):15–27. doi:10.1016/s0378-1119(99)00368-6
Najafabadi MS (2012) Improving rice (Oryza sativa L.) drought tolerance by suppressing a NF-YA transcription factor. Iranian. J Biotechnol 10(1):40–48
Nakano T, Suzuki K, Fujimura T, Shinshi H (2006) Genome-wide analysis of the ERF gene family in Arabidopsis and rice. Plant Physiol 140(2):411–432. doi:10.1104/pp.105.073783
Nelson DE, Repetti PP, Adams TR, Creelman RA, Wu J, Warner DC, Anstrom DC, Bensen RJ, Castiglioni PP, Donnarummo MG, Hinchey BS, Kumimoto RW, Maszle DR, Canales RD, Krolikowski KA, Dotson SB, Gutterson N, Ratcliffe OJ, Heard JE (2007) Plant nuclear factor Y (NF-Y) B subunits confer drought tolerance and lead to improved corn yields on water-limited acres. Proc Natl Acad Sci U S A 104(42):16450–16455
Ottow EA, Brinker M, Teichmann T, Fritz E, Kaiser W, Brosché M, Kangasjärvi J, Jiang X, Polle A (2005) Populus euphratica displays apoplastic sodium accumulation, osmotic adjustment by decreases in calcium and soluble carbohydrates, and develops leaf succulence under salt stress. Plant Physiol 139(4):1762–1772. doi:10.1104/pp.105.069971
Prabu G, Kawar PG, Pagariya MC, Prasad DT (2011) Identification of water deficit stress upregulated genes in sugarcane. Plant Mol Biol Rep 29(2):291–304. doi:10.1007/s11105-010-0230-0
Qiu Q, Ma T, Hu Q, Liu B, Wu Y, Zhou H, Wang Q, Wang J, Liu J (2011) Genome-scale transcriptome analysis of the desert poplar, Populus euphratica. Tree Physiol. doi:10.1093/treephys/tpr015
Romier C, Cocchiarella F, Mantovani R, Moras D (2003) The NF-YB/NF-YC structure gives insight into DNA binding and transcription regulation by CCAAT factor NF-Y. J Biol Chem 278(2):1336–1345. doi:10.1074/jbc.M209635200
Rubin EM (2008) Genomics of cellulosic biofuels. Nature 454(7206):841–845
Sengupta D, Ramesh G, Mudalkar S, Kumar KRR, Kirti PB, Reddy AR (2012) Molecular cloning and characterization of gamma-glutamyl cysteine synthetase (Vr gamma ECS) from roots of Vigna radiata (L.) Wilczek under progressive drought stress and recovery. Plant Mol Biol Rep 30(4):894–903. doi:10.1007/s11105-011-0398-y
Siefers N, Dang KK, Kumimoto RW, Bynum WE, Tayrose G, Holt BF (2009) Tissue-specific expression patterns of Arabidopsis NF-Y transcription factors suggest potential for extensive combinatorial complexity. Plant Physiol 149(2):625–641. doi:10.1104/pp.108.130591
Stephenson TJ, McIntyre CL, Collet C, Xue GP (2007) Genome-wide identification and expression analysis of the NF-Y family of transcription factors in Triticum aestivum. Plant Mol Biol 65(1–2):77–92. doi:10.1007/s11103-007-9200-9
Stephenson T, McIntyre C, Collet C, Xue G-P (2010) TaNF-YC11, one of the light-upregulated NF-YC members in Triticum aestivum, is co-regulated with photosynthesis-related genes. Funct Integr Genomics 10(2):265–276. doi:10.1007/s10142-010-0158-3
Stephenson T, McIntyre C, Collet C, Xue G-P (2011) TaNF-YB3 is involved in the regulation of photosynthesis genes in Triticum aestivum. Funct Integr Genomics 1–14. doi:10.1007/s10142-011-0212-9
Testa A, Donati G, Yan P, Romani F, Huang TH-M, Viganò MA, Mantovani R (2005) Chromatin immunoprecipitation (ChIP) on chip experiments uncover a widespread distribution of NF-Y binding CCAAT sites outside of core promoters. J Biol Chem 280(14):13606–13615. doi:10.1074/jbc.M414039200
Thirumurugan T, Ito Y, Kubo T, Serizawa A, Kurata N (2008) Identification, characterization and interaction of HAP family genes in rice. Mol Genet Genomics 279(3):279–289. doi:10.1007/s00438-007-0312-3
Tiwari SB, Shen Y, Chang H-C, Hou Y, Harris A, Ma SF, McPartland M, Hymus GJ, Adam L, Marion C, Belachew A, Repetti PP, Reuber TL, Ratcliffe OJ (2010) The flowering time regulator constans is recruited to the flowering locus T promoter via a unique cis-element. New Phytol 187(1):57–66. doi:10.1111/j.1469-8137.2010.03251.x
Tuskan GA, DiFazio SP, Teichmann T (2004) Poplar genomics is getting popular: the impact of the poplar genome project on tree research. Plant Biol (Stuttg) 7(01):2,4. doi:10.1055/s-2003-44715
Warpeha KM, Upadhyay S, Yeh J, Adamiak J, Hawkins SI, Lapik YR, Anderson MB, Kaufman LS (2007) The GCR1, GPA1, PRN1, NF-Y signal chain mediates both blue light and abscisic acid responses in Arabidopsis. Plant Physiol 143(4):1590–1600. doi:10.1104/pp.106.089904
Yamamoto A, Kagaya Y, Toyoshima R, Kagaya M, Takeda S, Hattori T (2009) Arabidopsis NF-YB subunits LEC1 and LEC1-LIKE activate transcription by interacting with seed-specific ABRE-binding factors. Plant J 58(5):843–856. doi:10.1111/j.1365-313X.2009.03817.x
Yang J, Xie ZY, Glover BJ (2005) Asymmetric evolution of duplicate genes encoding the CCAAT-binding factor NF-Y in plant genomes. New Phytol 165(2):623–631. doi:10.1111/j.1469-8137.2004.01260.x
Zhu Z, Shendure J, Church GM (2005) Discovering functional transcription-factor combinations in the human cell cycle. Genome Res 15(6):848–855. doi:10.1101/gr.3394405
Acknowledgments
The research was supported by grants from the Ministry of Science and Technology of China (2011BAD38B01, 2009CB119101), and the National Natural Science Foundation of China (30972339, 31070597, 30730077).
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
ESM 1
Primers designed based on the 20 NF-YB genes in Populus genome for amplicon of P. euphratica NF-YB genes by RT-qPCR. (XLSX 14 kb)
Rights and permissions
About this article
Cite this article
Yan, DH., Xia, X. & Yin, W. NF-YB Family Genes Identified in a Poplar Genome-wide Analysis and Expressed in Populus euphratica Are Responsive to Drought Stress. Plant Mol Biol Rep 31, 363–370 (2013). https://doi.org/10.1007/s11105-012-0508-5
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11105-012-0508-5