Abstract
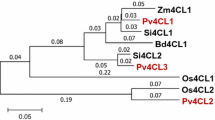
Sugarcane (Saccharum spp. hybrids) is a major feedstock for commercial bioethanol production. The recent integration of conversion technologies that utilize lignocellulosic sugarcane residues as well as sucrose from stem internodes has elevated bioethanol yields. RNAi suppression of lignin biosynthetic enzymes is a successful strategy to improve the saccharification of lignocellulosic biomass. 4-coumarate:coenzyme A ligase (4CL) is a key enzyme in the biosynthesis of phenylpropanoid metabolites, such as lignin and flavonoids. Identifying a major 4CL involved in lignin biosynthesis among multiple isoforms with functional divergence is key to manipulate lignin biosynthesis. In this study, two full length 4CL genes (Sh4CL1 and Sh4CL2) were isolated and characterized in sugarcane. Phylogenetic, expression and RNA interference (RNAi) analysis confirmed that Sh4CL1 is a major lignin biosynthetic gene. An intragenic precision breeding strategy may facilitate the regulatory approval of the genetically improved events and was used for RNAi suppression of Sh4CL1. Both, the RNAi inducing cassette and the expression cassette for the mutated ALS selection marker consisted entirely of DNA sequences from sugarcane or the sexually compatible species Sorghum bicolor. Field grown sugarcane with intragenic RNAi suppression of Sh4CL1 resulted in reduction of the total lignin content by up to 16.5 % along with altered monolignol ratios without reduction in biomass yield. Mature, field grown, intragenic sugarcane events displayed 52–76 % improved saccharification efficiency of lignocellulosic biomass compared to wild type (WT) controls. This demonstrates for the first time that an intragenic approach can add significant value to lignocellulosic feedstocks for biofuel and biochemical production.







Similar content being viewed by others
References
Altpeter F, Oraby H (2010) Sugarcane. In: Kempken F, Jung C (eds) Genetic Modification of Plants. Springer, Heidelberg, pp 453–472
Altpeter F, Sandhu S (2010) Genetic transformation–biolistics. In: Davey MR, Anthony P (eds) Plant Cell Culture: Essential Methods. Scion Publishing Ltd, Oxfordshire, UK, pp 217–240
Baxter HL, Mazarei M, Labbe N, Kline LM, Cheng Q, Windham MT, Mann DGJ, Chunxiang F, Ziebell A, Sykes RW, Rodriguez M, Davis MF, Mielenz JR, Dixon RA, Wang Z-Y, Stewart CN (2014) Two-year field analysis of reduced recalcitrance transgenic switchgrass. Plant Biotech J 12:914–924
Bonawitz ND, Chapple C (2010) The genetics of lignin biosynthesis: connecting genotype to phenotype. Annu Rev Genet 44:337–363
Bottcher A, Cesarino I, Santos AB, Vicentini R, Mayer JL, Vanholme R, Morreel K, Goeminne G, Moura JC, Nobile PM, Carmello-Guerreiro SM, Anjos IA, Creste S, Boerjan W, Landell MG, Mazzafera P (2013) Lignification in sugarcane: biochemical characterization, gene discovery, and expression analysis in two genotypes contrasting for lignin content. Plant Physiol 163:1539–1557
Chen F, Dixon RA (2007) Lignin modification improves fermentable sugar yields for biofuel production. Nat Biotech 25:759–761
Dal-Bianco M, Carneiro MS, Hotta CT, Chapola RG, Hoffmann HP, Garcia AAF, Souza GM (2012) Sugarcane improvement: how far can we go? Curr Opin Biotechnol 23:265–270
de Vetten N, Wolters AM, Raemakers K, van der Meer I, ter Stege R, Heeres E, Heeres P, Visser R (2003) A transformation method for obtaining marker-free plants of a cross-pollinating and vegetatively propagated crop. Nat Biotech 21:439–442
Dermawan H, Karan R, Jung JH, Zhao Y, Parajuli S, Sanahuja G, Altpeter F (2016) Development of an intragenic gene transfer and selection protocol for sugarcane resulting in resistance to acetolactate synthase-inhibiting herbicide. Plant Cell Tissue Org (in press)
Dias MOS, Ensinas AV, Nebra SA, Filho RM, Rossell CEV, Maciel MRW (2009) Production of bioethanol and other bio-based materials from sugarcane bagasse: Integration to conventional bioethanol production process. Chem Eng Res Des 87:1206–1216
Dias MOS, Cavalett O, Filho RM, Bonomi A (2014) Integrated first and second generation ethanol production from sugarcane. Chem Eng Trans 37:445–450
Ehlting J, Büttner D, Wang Q, Douglas CJ, Somssich IE, Kombrink E (1999) Three 4-coumarate:coenzyme A ligases in Arabidopsis thaliana represent two evolutionarily divergent classes in angiosperms. Plant J 19:9–20
Gadaleta A, Giancaspro A, Blechl AE, Blanco A (2008) A transgenic durum wheat line that is free of marker genes and expresses 1Dy10. J Cereal Sci 48:439–445
Gray DJ, Li ZT, Grant TNL, Dean DA, Trigiano RN, Dhekney SA (2015) The application of precision breeding (PB) for crop improvement is fully consistent with the plant life cycle: the utility of PB for grapevine. Acta Hortic 1115:49–56
Gui J, Shen J, Li L (2011) Functional characterization of evolutionarily divergent 4-coumarate:coenzyme a ligases in rice. Plant Physiol 157:574–586
Han KM, Dharmawardhana P, Arias RS, Ma C, Busov V, Strauss SH (2011) Gibberellin-associated cisgenes modify growth, stature and wood properties in Populus. Plant Biotech J 9:162–178
Harrison MD, Geijskes RJ, Lloyd R, Miles S, Palupe A, Sainz MB, Dale JL (2014) Recombinant cellulase accumulation in the leaves of mature, vegetatively propagated transgenic sugarcane. Mol Biotechnol 56:795–802
Hatfield RD, Grabber J, Ralph J, Brei K (1999) Using the acetyl bromide assay to determine lignin concentrations in herbaceous plants: some cautionary notes. J Agricult Food Chem 47:628–632
Holme IB, Dionisio G, Brinch-Pedersen H, Wendt T, Madsen CK, Vincze E, Holm PB (2012) Cisgenic barley with improved phytase activity. Plant Biotech J 10:237–247
Holme IB, Wendt T, Holm PB (2013) Intragenesis and cisgenesis as alternatives to transgenic crop development. Plant Biotech J 11:395–407
Hu WJ, Kawaoka A, Tsai CJ, Lung J, Osakabe K, Ebinuma H, Chiang VL (1998) Compartmentalized expression of two structurally and functionally distinct 4-coumarate:CoA ligase genes in aspen (Populus tremuloides). Proc Natl Acad Sci USA 95:5407–5412
Hu Y, Gai Y, Yin L, Wang X, Feng C, Feng L, Li D, Jiang X-N, Wang D-C (2010) Crystal structures of a Populus tomentosa 4-coumarate:CoA ligase shed light on its enzymatic mechanisms. Plant Cell 22:3093–3104
Iskandar H, Simpson R, Casu R, Bonnett G, Maclean D, Manners J (2004) Comparison of reference genes for quantitative real-time polymerase chain reaction analysis of gene expression in sugarcane. Plant Mol Biol Rep 22:325–337
Jørgensen H, Kristensen JB, Felby C (2007) Enzymatic conversion of lignocellulose into fermentable sugars: challenges and opportunities. Biofuels Bioprod Biorefin 1:119–134
Joshi S, Schaart J, Groenwold R, Jacobsen E, Schouten H, Krens F (2011) Functional analysis and expression profiling of HcrVf1 and HcrVf2 for development of scab resistant cisgenic and intragenic apples. Plant Mol Biol 75:579–591
Jung JH, Altpeter F (2016) TALEN mediated targeted mutagenesis of the caffeic acid O-methyltransferase in highly polyploid sugarcane improves cell wall composition for production of bioethanol. Plant Mol Biol (in press)
Jung JH, Kim JY, Fouad M, Vermerris W, Gallo M, Altpeter F (2010) Isolation, characterization and RNAi-suppression of COMT and 4CL genes in sugarcane. In 2010 In Vitro Biology Meeting and IAPB 12th World Congress Abstract Issue, P-084. Springer, St. Louis, MO
Jung JH, Fouad WM, Vermerris W, Gallo M, Altpeter F (2012) RNAi suppression of lignin biosynthesis in sugarcane reduces recalcitrance for biofuel production from lignocellulosic biomass. Plant Biotech J 10:1067–1076
Jung JH, Vermerris W, Gallo M, Fedenko JR, Erickson JE, Altpeter F (2013) RNA interference suppression of lignin biosynthesis increases fermentable sugar yields for biofuel production from field-grown sugarcane. Plant Biotech J 11:709–716
Kim J, Gallo M, Altpeter F (2012) Analysis of transgene integration and expression following biolistic transfer of different quantities of minimal expression cassette into sugarcane (Saccharum spp. hybrids). Plant Cell Tissue Org 108:297–302
Lakshmanan P, Geijskes RJ, Aitken K, Grof CP, Bonnett G, Smith G (2005) Sugarcane biotechnology: The challenges and opportunities. In Vitro Cell Dev Biol Plant 41:345–363
Lee D, Meyer K, Chapple C, Douglas CJ (1997) Antisense suppression of 4-coumarate:coenzyme A ligase activity in Arabidopsis leads to altered lignin subunit composition. Plant Cell 9:1985–1998
Mudge SR, Basnayake SW, Moyle RL, Osabe K, Graham MW, Morgan TE, Birch RG (2013) Mature-stem expression of a silencing-resistant sucrose isomerase gene drives isomaltulose accumulation to high levels in sugarcane. Plant Biotechnol J 11:502–509
Nagaya S, Kawamura K, Shinmyo A, Kato K (2010) The HSP terminator of Arabidopsis thaliana increases gene expression in plant cells. Plant Cell Physiol 51:328–332
Nielsen KM (2003) Transgenic organisms-time for conceptual diversification? Nat. Biotech 21:227–228
Pichersky E, Gang DR (2000) Genetics and biochemistry of secondary metabolites in plants: an evolutionary perspective. Trends Plant Sci 5:439–445
Poovaiah CR, Nageswara-Rao M, Soneji JR, Baxter HL, Stewart CN (2014) Altered lignin biosynthesis using biotechnology to improve lignocellulosic biofuel feedstocks. Plant Biotech J 12:1163–1173
Rommens CM (2004) All-native DNA transformation: a new approach to plant genetic engineering. Trends Plant Sci 9:457–464
Rommens CM, Haring MA, Swords K, Davies HV, Belknap WR (2007) The intragenic approach as a new extension to traditional plant breeding. Trends Plant Sci 12:397–403
Saballos A, Sattler SE, Sanchez E, Foster TP, Xin Z, Kang C, Pedersen JF, Vermerris W (2012) Brown midrib2 (Bmr2) encodes the major 4-coumarate:coenzyme A ligase involved in lignin biosynthesis in sorghum (Sorghum bicolor (L.) Moench). Plant J 70:818–830
Schouten HJ, Jacobsen E (2008) Cisgenesis and intragenesis, sisters in innovative plant breeding. Trends Plant Sci 13:260–261
Schouten HJ, Krens FA, Jacobsen E (2006) Cisgenic plants are similar to traditionally bred plants. EMBO Rep 7:750–753
Sluiter A, Hames B, Ruiz R, Scarlata C, Sluiter J, Templeton D, Crocker D (2008) Determination of structural carbohydrates and lignin in biomass In Laboratory Analytical Procedure (LAP), Technical Report NREL⁄TP-510-42618). National Renewable Energy Laboratory, Golden
Somerville C, Youngs H, Taylor C, Davis SC, Long SP (2010) Feedstocks for Lignocellulosic Biofuels. Science 329:790–792
Stout AT, Davis AA, Domec J-C, Yang C, Shi R, King JS (2014) Growth under field conditions affects lignin content and productivity in transgenic Populus trichocarpa with altered lignin biosynthesis. Biomass Bioenergy 68:228–239
Tamura K, Stecher G, Peterson D, Filipski A, Kumar S (2013) MEGA6: Molecular evolutionary genetics analysis version 6.0. Mol Biol Evol 30:2725–2729
Taparia Y, Gallo M, Altpeter F (2012) Comparison of direct and indirect embryogenesis protocols, biolistic gene transfer and selection parameters for efficient genetic transformation of sugarcane. Plant Cell Tiss Org 111:131–141
Vanblaere T, Szankowski I, Schaart J, Schouten H, Flachowsky H, Broggini GAL, Gessler C (2011) The development of a cisgenic apple plant. J Biotech 154:304–311
Vermerris W (2008) Composition and biosynthesis of lignocellulosic biomass. In: Vermerris W (ed) Genetic Improvement of Bioenergy Crops. Springer, New York, pp 89–142
Vermerris W, Abril A (2015) Enhancing cellulose utilization for fuels and chemicals by genetic modification of plant cell wall architecture. Curr Opin Biotechnol 32:104–112
Voelker SL, Lachenbruch B, Meinzer FC, Jourdes M, Ki C, Patten AM, Davin LB, Lewis NG, Tuskan GE, Gunter L, Decker SR, Selig MG, Sykes R, Himmel ME, Kitin P, Shevchenko O, Strauss SH (2010) Antisense down-regulation of 4 CL expression alters lignification, tree growth, and saccharification potential of field-grown poplar. Plant Physiol 154:874–886
Voelker SL, Lachenbruch B, Meinzer FC, Kitin P, Strauss SH (2011) Transgenic poplars with reduced lignin show impaired xylem conductivity, growth efficiency and survival. Plant Cell Environ 34:655–668
Vogt T (2010) Phenylpropanoid biosynthesis. Mol Plant 3:2–20
Wagner A, Donaldson L, Kim H, Phillips L, Flint H, Steward D, Torr K, Koch G, Schmitt U, Ralph J (2009) Suppression of 4-coumarate-CoA ligase in the coniferous gymnosperm Pinus radiata. Plant Physiol 149:370–383
Wang H, Xue Y, Chen Y, Li R, Wei J (2012) Lignin modification improves biofuel production potential in transgenic Populus tomentosa. Ind Crops Prod 37:170–177
Weeks J, Ye J, Rommens C (2008) Development of an in planta method for transformation of alfalfa (Medicago sativa). Transgenic Res 17:587–597
Xu Z, Zhang D, Hu J, Zhou X, Ye X, Reichel KL, Stewart NR, Syrenne RD, Yang X, Gao P, Shi W, Doeppke C, Sykes RW, Burris JN, Bozell JJ, Cheng MZ, Hayes DG, Labbe N, Davis M, Stewart CN, Yuan JS (2009) Comparative genome analysis of lignin biosynthesis gene families across the plant kingdom. BMC Bioinform 10:S3
Xu B, Escamilla-Trevino LL, Sathitsuksanoh N, Shen Z, Shen H, Zhang YH, Dixon RA, Zhao B (2011) Silencing of 4-coumarate:coenzyme A ligase in switchgrass leads to reduced lignin content and improved fermentable sugar yields for biofuel production. New Phytol 192:611–625
Zale J, Jung JH, Kim JY, Pathak B, Karan R, Liu H, Chen X, Wu H, Candreva J, Zhai Z, Shanklin J, Altpeter F (2016) Metabolic engineering of sugarcane to accumulate energy-dense triacylglycerols in vegetative biomass. Plant Biotech J 14:661–669
Acknowledgments
The information, data or work presented herein were funded in part by Syngenta. The authors would like to thank Sun Gro Horticulture, Apopka, FL, for donation of Fafard #2 potting mix.
Author Contributions
FA conceived the experiments. FA, JHJ, BK and GMM designed the experiments. JHJ isolated candidate genes, evaluated their expression profile and constructed the plasmids. JHJ and HD generated transgenic plants and completed molecular analysis of transgenic plants, BK carried out the field trial including phenotypic, sugar and biomass yield evaluation, monitoring of plots and preparing reports for the regulatory agency. JHJ analyzed cell wall composition. GMM carried out the cell wall saccharification analysis, JHJ and FA wrote the manuscript. All authors read and approved the final manuscript.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors have no conflict of interest to declare.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Jung, J.H., Kannan, B., Dermawan, H. et al. Precision breeding for RNAi suppression of a major 4-coumarate:coenzyme A ligase gene improves cell wall saccharification from field grown sugarcane. Plant Mol Biol 92, 505–517 (2016). https://doi.org/10.1007/s11103-016-0527-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11103-016-0527-y