Abstract
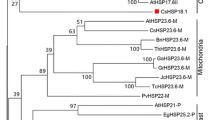
Small heat shock proteins (sHSPs) are crucial components of the plant response to heat shock. We identified and analyzed eight sHSP genes of Capsella bursa-pastoris to better understand the ability of this species to adapt. Eight genes were initially cloned and sequenced from the mature embryo cDNA pool. They belong to the cytosolic I (CI), cytosolic II (CII), and cytosolic III (CIII) subfamilies. One CI sHSP gene was homologous to that of C. rubella. Sequence analysis using 3′ RACE revealed that there are two or more variable 3′-untranslated regions (UTRs) in these sHSP transcripts. The transcriptional levels of the eight sHSP genes were analyzed in different organs and developmental stages via qRT-PCR. Eight genes were significantly up-regulated in young leaves exposed to heat stress at 42 °C, and also showed differential responses to ABA treatment. We also compared expression of these genes with corresponding Arabidopsis sHSP genes and found some differences between the two species.
Similar content being viewed by others
Abbreviations
- sHSP:
-
small heat shock protein
- qRT-PCR:
-
quantitatuve reverse transcriptase — polymerase chain reaction
- RACE:
-
rapid amplification of cDNA ends
- 3′ UTR:
-
3′ untranslated region
References
Arnold, K., Bordoli, L., Kopp, J., Schwede, T.: The SWISSMODEL workspace: a web-based environment for protein structure homology modelling. — Bioinformatics 22: 195–201, 2006.
Blanc, G., Wolfe, K.H.: Functional divergence of duplicated genes formed by polyploidy during Arabidopsis evolution. — Plant Cell 16: 1679–1691, 2004.
Bi, V.I., McMullen, M.D., Sanchez-Villeda, H., Schroeder, S., Gardiner, J., Polacco, M., Soderlund, C., Wing, R., Fang, Z., Coe E.H., Jr.: Single nucleotide polymorphisms and insertion-deletions for genetic markers and anchoring the maize fingerprint contig physical map. — Crop Sci. 46: 12–21, 2006.
Campbell, J.L., Klueva, N.Y., Zheng, H.G., Nieto-Sotelo, J., Ho, T.D., Nguyen, H.T.: Cloning of new members of heat shock protein HSP101 gene family in wheat (Triticum aestivum (L.) Moench) inducible by heat, dehydration, and ABA. — Biochim. biophys. Acta 1517: 270–277, 2001.
Casneuf, T., De Bodt, S., Raes, J., Maere, S., Van de Peer, Y.: Nonrandom divergence of gene expression following gene and genome duplications in the flowering plant Arabidopsis thaliana. — Genome Biol. 7: R13, 2006.
Dafny-Yelin, M., Tzfira, T., Vainstein, A., Adam Z.: Non-redundant functions of sHSP-CIs in acquired thermotolerance and their role in early seed development in Arabidopsis. — Plant mol. Biol. 67: 363–373, 2008.
Eveland, A.L., McCarty, D.R., Koch, K.E.: Transcript profiling by 3′-untranslated region sequencing resolves expression of gene families. — Plant Physiol. 146: 32–44, 2008.
Grigorova, B., Vaseva, I., Demirevska, K., Feller, U.: Combined drought and heat stress in wheat: changes in some heat shock proteins. — Biol. Plant. 55: 105–111, 2011.
Guex, N., Peitsch, M.C.: SWISS-MODEL and the Swiss-Pdb Viewer: an environment for comparative protein modelling. — Electrophoresis 18: 2714–2723, 1997.
Hurka, H., Bleeker, W., Neuffer, B.: Evolutionary process associated with biological invasions in the Brassicaceae. — Biol. Invasions 5: 281–292, 2003.
Hurka, H., Neuffer, B.: Evolutionary processes in the genus Capsella (Brassicaceae). — Plant Syst. Evol. 206: 295–316, 1997.
Hussain, S.S., Iqbal, M.T., Arif M.A., Amjad, M.: Beyond osmolytes and transcription factors: drought tolerance in plants via protective proteins and aquaporins. — Biol. Plant. 55: 401–413, 2011.
Jansen, R.P.: mRNA localization: message on the move. — Nat. Rev. mol. cell. Biol. 2: 247–256, 2001.
Jiang, C., Xu, J., Zhang, H., Zhang, X., Shi, J., Li, M., Ming, F.: A cytosolic class I small heat shock protein, RcHSP17.8, of Rosa chinensis confers resistance to a variety of stresses to Escherichia coli, yeast and Arabidopsis thaliana. — Plant Cell Environ. 32: 1046–1059, 2009.
Kalemba, E.M., Pukacka, S.: Changes in late embryogenesis abundant proteins and a small heat shock protein during storage of beech (Fagus sylvatica L.) seeds. — Environ. exp. Bot. 63: 274–280, 2008.
Kim, R., Kim, K.K., Yokota, H., Kim, S.H.: Small heat shock protein of Methanococcus jannaschii, a hyperthermophile. — Proc. nat. Acad. Sci. USA 95: 9129–9133, 1998.
Kotak, S., Vierling, E., Bäumlein, H., Von Koskull-Döring, P.: A novel transcriptional cascade regulating expression of heat stress proteins during seed development of Arabidopsis. — Plant Cell 19: 182–195, 2007.
Kumar, S., Tamura, K., Nei, M.: MEGA3: integrated software for molecular evolutionary genetics analysis and sequence alignment. — Brief Bioinform. 5: 150–163, 2004.
Kurkela, S., Franck, M.: Cloning and characterization of a coldand ABA-inducible Arabidopsis gene. — Plant mol. Biol. 15: 137–144, 1990.
Mitchell, P., Tollervey, D.: mRNA turnover. — Curr. Opin. Cell Biol. 13: 320–325, 2001.
Nautiyal, P.C., Shono, M.: Analysis of the role of mitochondrial and endoplasmic reticulum localized small heat shock proteins in tomato. — Biol. Plant. 54: 715–719, 2010.
Nishizawa, A., Yabuta, Y., Yoshida, E., Maruta, T., Yoshimura, K., Shigeoka S.: Arabidopsis heat shock transcription factor A2 as a key regulator in response to several types of environmental stress. — Plant J. 48:535-547, 2006.
Rampino, P., Mita, G., Assab, E., De Pascali, M., Giangrande, E., Treglia, A.S., Perrotta, C.: Two sunflower 17.6HSP genes, arranged in tandem and highly homologous, are induced differently by various elicitors. — Plant Biol. 12: 13–22, 2010.
Sato, Y., Yokoya, S.: Enhanced tolerance to drought in transgenic rice plants overexpressing a small heat-shock protein, sHSP17.7. — Plant Cell Rep. 27: 329–334, 2008.
Scharf, K.D., Siddique, M., Vierling, E.: The expanding family of Arabidopsis thaliana small heat stress proteins and a new family of proteins containing alpha-crystallin domains (Acd proteins). — Cell Stress Chaperon. 6: 225–237, 2001.
Schefe, J.H., Lehmann, K.E., Buschmann, I.R., Unger, T., Funke-Kaiser, H.: Quantitative real-time RT-PCR data analysis: current concepts and the novel “gene expression’s CT difference” formula. — J. Mol. Med. 84: 901–910, 2006.
Siddique, M., Gernhard, S., Von Koskull-Döring, P., Vierling, E., Scharf, K.D.: The plant sHSP superfamily: five new members in Arabidopsis thaliana with unexpected properties. — Cell Stress Chaperon. 13: 183–197, 2008.
Simon, B., Sengupta-Gopalan, C.: The 3′ untranslated region of the two cytosolic glutamine synthetase (GS1) genes in alfalfa (Medicago sativa) regulates transcript stability in response to glutamine. — Planta 232: 1151–1162, 2010.
Slotte, T., Ceplitis, A., Neuffer, B., Hurka, H., Lascoux, M.: Intrageneric phylogeny of Capsella (Brassicaceae) and the origin of the tetraploid C. bursa-pastoris based on chloroplast and nuclear DNA sequences. — Amer. J. Bot. 93: 1714–1724, 2006.
Slotte, T., Holm, K., McIntyre, L.M., Lagercrantz, U., Lascoux, M.: Differential expression of genes important for adaptation in Capsella bursa-pastoris (Brassicaceae). — Plant Physiol. 145: 160–173, 2007.
Sun, W., Bernard, C., Van de Cotte, B., Montagu, M.V., Verbruggen, N.: At-HSP17.6A, encoding a small heat-shock protein in Arabidopsis, can enhance osmotolerance upon overexpression. — Plant J. 27: 407–415, 2001.
Sun, W., Montagu, M.V., Verbruggen, N.: Small heat shock proteins and stress tolerance in plants. — Biochim. biophys. Acta 1577: 1–9, 2002.
Sun, Y., MacRae, T.H.: Small heat shock proteins: molecular structure and chaperone function. — Cell. Mol. Life Sci. 62: 2460–2476, 2005.
Sundby, C., Härndahl, U., Gustavsson, N., Åhrman, E., Murphy, D.J.: Conserved methionines in chloroplasts. — Biochim. biophys. Acta 1703: 191–202, 2005.
Tejedor-Cano, J., Prieto-Dapena, P., Almoguera, C., Carranco, R., Hiratsu, K., Ohme-Takagi, M., Jordano, J.: Loss of function of the HSFA9 seed longevity program. — Plant Cell Environ. 33: 1408–1417, 2010.
Van Montfort, R.L., Basha, E., Friedrich, K.L., Slingsby, C., Vierling, E.: Crystal structure and assembly of a eukaryotic small heat shock protein. — Nat. Struct. Biol. 8: 1025–1030, 2001.
Van Montfort, R.L., Slingsby, C., Vierling, E.: Structure and function of the small heat shock protein/alpha-crystallin family of molecular chaperones. — Adv. Protein Chem. 59: 105–156, 2002.
Wachter, A., Tunc-Ozdemir, M., Grove, B.C., Green, P.J., Shintani, D.K., Breaker, R.R.: Riboswitch control of gene expression in plants by splicing and alternative 3′ end processing of mRNAs. — Plant Cell 19: 3437–3450, 2007.
Waters, E.R., Aevermann, B.D., Sanders-Reed, Z.: Comparative analysis of the small heat shock proteins in three angiosperm genomes identifies new subfamilies and reveals diverse evolutionary patterns. — Cell Stress Chaperon. 13: 127–142, 2008.
Waters, E.R., Rioflorido, I.: Evolutionary analysis of the small heat shock proteins in five complete algal genomes. — J. Mol. Evol. 65: 162–174, 2007.
Waters, E.R., Vierling, E.: Chloroplast small heat shock proteins: Evidence for atypical evolution of an organelle-localized protein. — Proc. nat. Acad. Sci. USA 96: 14394–14399, 1999a.
Waters, E.R., Vierling, E.: The diversification of plant cytosolic small heat shock proteins preceded the divergence of mosses. — Mol. Biol. Evol. 16: 127–139, 1999b.
Wang, W., Vinocur, B., Shoseyov, O., Altman, A.: Roles of plant heat-shock proteins and molecular chaperones in the abiotic stress response. — Trends Plant Sci. 9: 244–252, 2004.
Xue, Y., Peng, R., Xiong, A., Li, X., Zha, D., Yao, Q.: Over-expression of heat shock protein gene hsp26 in Arabidopsis thaliana enhances heat tolerance. — Biol. Plant. 54: 105–111, 2010.
Author information
Authors and Affiliations
Corresponding author
Additional information
Acknowledgments: This work was supported by the State Key Basic Research and Development Plan of China (2007CB108700), the National Natural Science Foundation of China (31070204, 30570112) and the Chinese 111 Project.
Rights and permissions
About this article
Cite this article
Tao, P., Liu, L. & Wang, J.B. Characterization of eight cytosolic sHSP genes and their expression in Capsella bursa-pastoris . Biol Plant 56, 648–656 (2012). https://doi.org/10.1007/s10535-012-0239-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10535-012-0239-3