Abstract
Genomic prediction of the extreme forms of adult body height or stature is of practical relevance in several areas such as pediatric endocrinology and forensic investigations. Here, we examine 770 extremely tall cases and 9,591 normal height controls in a population-based Dutch European sample to evaluate the capability of known height-associated DNA variants in predicting tall stature. Among the 180 normal height-associated single nucleotide polymorphisms (SNPs) previously reported by the Genetic Investigation of ANthropocentric Traits (GIANT) genome-wide association study on normal stature, in our data 166 (92.2 %) showed directionally consistent effects and 75 (41.7 %) showed nominally significant association with tall stature, indicating that the 180 GIANT SNPs are informative for tall stature in our Dutch sample. A prediction analysis based on the weighted allele sums method demonstrated a substantially improved potential for predicting tall stature (AUC = 0.75; 95 % CI 0.72–0.79) compared to a previous attempt using 54 height-associated SNPs (AUC = 0.65). The achieved accuracy is approaching practical relevance such as in pediatrics and forensics. Furthermore, a reanalysis of all SNPs at the 180 GIANT loci in our data identified novel secondary association signals for extreme tall stature at TGFB2 (P = 1.8 × 10−13) and PCSK5 (P = 7.8 × 10−11) suggesting the existence of allelic heterogeneity and underlining the importance of fine analysis of already discovered loci. Extrapolating from our results suggests that the genomic prediction of at least the extreme forms of common complex traits in humans including common diseases are likely to be informative if large numbers of trait-associated common DNA variants are available.



Similar content being viewed by others
References
Aulchenko YS, Struchalin MV, Belonogova NM, Axenovich TI, Weedon MN, Hofman A, Uitterlinden AG, Kayser M, Oostra BA, van Duijn CM, Janssens AC, Borodin PM (2009) Predicting human height by Victorian and genomic methods. Eur J Hum Genet 17:1070–1075. doi:10.1038/ejhg.2009.5
Branicki W, Liu F, van Duijn K, Draus-Barini J, Pospiech E, Walsh S, Kupiec T, Wojas-Pelc A, Kayser M (2011) Model-based prediction of human hair color using DNA variants. Hum Genet 129:443–454. doi:10.1007/s00439-010-0939-8
Campbell CD, Ogburn EL, Lunetta KL, Lyon HN, Freedman ML, Groop LC, Altshuler D, Ardlie KG, Hirschhorn JN (2005) Demonstrating stratification in a European American population. Nat Genet 37:868–872
Carmichael CM, McGue M (1995) A cross-sectional examination of height, weight, and body mass index in adult twins. J Gerontol A Biol Sci Med Sci 50:B237–B244
Chan Y, Holmen OL, Dauber A, Vatten L, Havulinna AS, Skorpen F, Kvaloy K, Silander K, Nguyen TT, Willer C, Boehnke M, Perola M, Palotie A, Salomaa V, Hveem K, Frayling TM, Hirschhorn JN, Weedon MN (2011) Common variants show predicted polygenic effects on height in the tails of the distribution, except in extremely short individuals. PLoS Genet 7:e1002439. doi:10.1371/journal.pgen.1002439
Chitramuthu BP, Baranowski DC, Cadieux B, Rousselet E, Seidah NG, Bennett HP (2010) Molecular cloning and embryonic expression of zebrafish PCSK5 co-orthologues: functional assessment during lateral line development. Dev Dyn 239:2933–2946. doi:10.1002/dvdy.22426
Dagoneau N, Benoist-Lasselin C, Huber C, Faivre L, Megarbane A, Alswaid A, Dollfus H, Alembik Y, Munnich A, Legeai-Mallet L, Cormier-Daire V (2004) ADAMTS10 mutations in autosomal recessive Weill-Marchesani syndrome. Am J Hum Genet 75:801–806. doi:10.1086/425231
Fredriks AM, van Buuren S, Burgmeijer RJ, Meulmeester JF, Beuker RJ, Brugman E, Roede MJ, Verloove-Vanhorick SP, Wit JM (2000) Continuing positive secular growth change in The Netherlands 1955–1997. Pediatr Res 47:316–323
Gudbjartsson DF, Walters GB, Thorleifsson G, Stefansson H, Halldorsson BV, Zusmanovich P, Sulem P, Thorlacius S, Gylfason A, Steinberg S, Helgadottir A, Ingason A, Steinthorsdottir V, Olafsdottir EJ, Olafsdottir GH, Jonsson T, Borch-Johnsen K, Hansen T, Andersen G, Jorgensen T, Pedersen O, Aben KK, Witjes JA, Swinkels DW, den Heijer M, Franke B, Verbeek AL, Becker DM, Yanek LR, Becker LC, Tryggvadottir L, Rafnar T, Gulcher J, Kiemeney LA, Kong A, Thorsteinsdottir U, Stefansson K (2008) Many sequence variants affecting diversity of adult human height. Nat Genet 40:609–615. doi:10.1038/ng.122
Haji-Seyed-Javadi R, Jelodari-Mamaghani S, Paylakhi SH, Yazdani S, Nilforushan N, Fan JB, Klotzle B, Mahmoudi MJ, Ebrahimian MJ, Chelich N, Taghiabadi E, Kamyab K, Boileau C, Paisan-Ruiz C, Ronaghi M, Elahi E (2012) LTBP2 mutations cause Weill-Marchesani and Weill-Marchesani-like syndrome and affect disruptions in the extracellular matrix. Hum Mutat 33:1182–1187. doi:10.1002/humu.22105
Hendriks AE, Boellaard WP, van Casteren NJ, Romijn JC, de Jong FH, Boot AM, Drop SL (2010) Fatherhood in tall men treated with high-dose sex steroids during adolescence. J Clin Endocrinol Metab 95:5233–5240. doi:10.1210/jc.2010-0435
Hendriks AE, Brown MR, Boot AM, Oostra BA, de Jong FH, Drop SL, Parks JS (2011a) Common polymorphisms in the GH/IGF-1 axis contribute to growth in extremely tall subjects. Growth Horm IGF Res 21:318–324. doi:10.1016/j.ghir.2011.08.001
Hendriks AE, Brown MR, Boot AM, Oostra BA, Drop SL, Parks JS (2011b) Genetic variation in candidate genes like the HMGA2 gene in the extremely tall. Horm Res Paediatr 76:307–313. doi:10.1159/000330764
Hendriks AE, Laven JS, Valkenburg O, Fong SL, Fauser BC, de Ridder MA, de Jong FH, Visser JA, van Ginneken AM, Boot AM, Drop SL (2011c) Fertility and ovarian function in high-dose estrogen-treated tall women. J Clin Endocrinol Metab 96:1098–1105. doi:10.1210/jc.2010-2244
Hendriks AE, Drop SL, Laven JS, Boot AM (2012) Fertility of tall girls treated with high-dose estrogen, a dose-response relationship. J Clin Endocrinol Metab 97:3107–3114. doi:10.1210/jc.2012-1078
Hirschhorn JN, Lindgren CM, Daly MJ, Kirby A, Schaffner SF, Burtt NP, Altshuler D, Parker A, Rioux JD, Platko J, Gaudet D, Hudson TJ, Groop LC, Lander ES (2001) Genomewide linkage analysis of stature in multiple populations reveals several regions with evidence of linkage to adult height. Am J Hum Genet 69:106–116. doi:10.1086/321287
Hofman A, Grobbee DE, de Jong PT, van den Ouweland FA (1991) Determinants of disease and disability in the elderly: the Rotterdam Elderly Study. Eur J Epidemiol 7:403–422
Hofman A, Breteler MM, van Duijn CM, Krestin GP, Pols HA, Stricker BH, Tiemeier H, Uitterlinden AG, Vingerling JR, Witteman JC (2007) The Rotterdam Study: objectives and design update. Eur J Epidemiol 22:819–829. doi:10.1007/s10654-007-9199-x
Hofman A, Breteler MM, van Duijn CM, Janssen HL, Krestin GP, Kuipers EJ, Stricker BH, Tiemeier H, Uitterlinden AG, Vingerling JR, Witteman JC (2009) The Rotterdam Study: 2010 objectives and design update. Eur J Epidemiol 24:553–572. doi:10.1007/s10654-009-9386-z
Janssens AC, Pardo MC, Steyerberg EW, van Duijn CM (2004) Revisiting the clinical validity of multiplex genetic testing in complex diseases. Am J Hum Genet 74:585–588. doi:10.1086/382052 (author reply 588–589)
Kayser M, de Knijff P (2011) Improving human forensics through advances in genetics, genomics and molecular biology. Nat Rev Genet 12:179–192. doi:10.1038/nrg2952
Kayser M, Schneider PM (2009) DNA-based prediction of human externally visible characteristics in forensics: motivations, scientific challenges, and ethical considerations. Forensic Sci Int Genet 3:154–161. doi:10.1016/j.fsigen.2009.01.012
Kim JJ, Park YM, Baik KH, Choi HY, Yang GS, Koh I, Hwang JA, Lee J, Lee YS, Rhee H, Kwon TS, Han BG, Heath KE, Inoue H, Yoo HW, Park K, Lee JK (2012) Exome sequencing and subsequent association studies identify five amino acid-altering variants influencing human height. Hum Genet 131:471–478. doi:10.1007/s00439-011-1096-4
Lango Allen H, Estrada K, Lettre G, Berndt SI, Weedon MN, Rivadeneira F, Willer CJ, Jackson AU, Vedantam S, Raychaudhuri S, Ferreira T, Wood AR, Weyant RJ, Segre AV, Speliotes EK, Wheeler E, Soranzo N, Park JH, Yang J, Gudbjartsson D, Heard-Costa NL, Randall JC, Qi L, Vernon Smith A, Magi R, Pastinen T, Liang L, Heid IM, Luan J, Thorleifsson G, Winkler TW, Goddard ME, Sin Lo K, Palmer C, Workalemahu T, Aulchenko YS, Johansson A, Carola Zillikens M, Feitosa MF, Esko T, Johnson T, Ketkar S, Kraft P, Mangino M, Prokopenko I, Absher D, Albrecht E, Ernst F, Glazer NL, Hayward C, Hottenga JJ, Jacobs KB, Knowles JW, Kutalik Z, Monda KL, Polasek O, Preuss M, Rayner NW, Robertson NR, Steinthorsdottir V, Tyrer JP, Voight BF, Wiklund F, Xu J, Hua Zhao J, Nyholt DR, Pellikka N, Perola M, Perry JR, Surakka I, Tammesoo ML, Altmaier EL, Amin N, Aspelund T, Bhangale T, Boucher G, Chasman DI, Chen C, Coin L, Cooper MN, Dixon AL, Gibson Q, Grundberg E, Hao K, Juhani Junttila M, Kaplan LM, Kettunen J, Konig IR, Kwan T, Lawrence RW, Levinson DF, Lorentzon M, McKnight B, Morris AP, Muller M, Suh Ngwa J, Purcell S, Rafelt S, Salem RM, Salvi E et al (2010) Hundreds of variants clustered in genomic loci and biological pathways affect human height. Nature 467:832–838. doi:10.1038/nature09410
Le Goff C, Cormier-Daire V (2012) From tall to short: the role of TGFbeta signaling in growth and its disorders. Am J Med Genet C Semin Med Genet 160C:145–153. doi:10.1002/ajmg.c.31337
Lettre G, Jackson AU, Gieger C, Schumacher FR, Berndt SI, Sanna S, Eyheramendy S, Voight BF, Butler JL, Guiducci C, Illig T, Hackett R, Heid IM, Jacobs KB, Lyssenko V, Uda M, Boehnke M, Chanock SJ, Groop LC, Hu FB, Isomaa B, Kraft P, Peltonen L, Salomaa V, Schlessinger D, Hunter DJ, Hayes RB, Abecasis GR, Wichmann HE, Mohlke KL, Hirschhorn JN (2008) Identification of ten loci associated with height highlights new biological pathways in human growth. Nat Genet 40:584–591. doi:10.1038/ng.125
Li Y, Willer C, Sanna S, Abecasis G (2009) Genotype imputation. Annu Rev Genomics Hum Genet 10:387–406
Liu F, van Duijn K, Vingerling JR, Hofman A, Uitterlinden AG, Janssens AC, Kayser M (2009a) Eye color and the prediction of complex phenotypes from genotypes. Curr Biol 19:R192–R193. doi:10.1016/j.cub.2009.01.027
Liu S, Tang W, Fang J, Ren J, Li H, Xiao Z, Quarles LD (2009b) Novel regulators of Fgf23 expression and mineralization in Hyp bone. Mol Endocrinol 23:1505–1518. doi:10.1210/me.2009-0085
Macgregor S, Cornes BK, Martin NG, Visscher PM (2006) Bias, precision and heritability of self-reported and clinically measured height in Australian twins. Hum Genet 120:571–580. doi:10.1007/s00439-006-0240-z
Perola M, Sammalisto S, Hiekkalinna T, Martin NG, Visscher PM, Montgomery GW, Benyamin B, Harris JR, Boomsma D, Willemsen G, Hottenga JJ, Christensen K, Kyvik KO, Sorensen TI, Pedersen NL, Magnusson PK, Spector TD, Widen E, Silventoinen K, Kaprio J, Palotie A, Peltonen L (2007) Combined genome scans for body stature in 6,602 European twins: evidence for common Caucasian loci. PLoS Genet 3:e97. doi:10.1371/journal.pgen.0030097
Phillips K, Matheny AP Jr (1990) Quantitative genetic analysis of longitudinal trends in height: preliminary results from the Louisville Twin Study. Acta Genet Med Gemellol (Roma) 39:143–163
Silventoinen K, Kaprio J, Lahelma E, Koskenvuo M (2000) Relative effect of genetic and environmental factors on body height: differences across birth cohorts among Finnish men and women. Am J Public Health 90:627–630
Silventoinen K, Sammalisto S, Perola M, Boomsma DI, Cornes BK, Davis C, Dunkel L, De Lange M, Harris JR, Hjelmborg JV, Luciano M, Martin NG, Mortensen J, Nistico L, Pedersen NL, Skytthe A, Spector TD, Stazi MA, Willemsen G, Kaprio J (2003) Heritability of adult body height: a comparative study of twin cohorts in eight countries. Twin Res 6:399–408
Su PH, Wang SL, Chen JY (2011) Gender differences of final height contributed by parents’ height among healthy individuals. Pediatr Neonatol 52:183–189. doi:10.1016/j.pedneo.2011.05.003
Szumska D, Pieles G, Essalmani R, Bilski M, Mesnard D, Kaur K, Franklyn A, El Omari K, Jefferis J, Bentham J, Taylor JM, Schneider JE, Arnold SJ, Johnson P, Tymowska-Lalanne Z, Stammers D, Clarke K, Neubauer S, Morris A, Brown SD, Shaw-Smith C, Cama A, Capra V, Ragoussis J, Constam D, Seidah NG, Prat A, Bhattacharya S (2008) VACTERL/caudal regression/Currarino syndrome-like malformations in mice with mutation in the proprotein convertase Pcsk5. Genes Dev 22:1465–1477. doi:10.1101/gad.479408
Thodberg HH, Jenni OG, Caflisch J, Ranke MB, Martin DD (2009) Prediction of adult height based on automated determination of bone age. J Clin Endocrinol Metab 94:4868–4874. doi:10.1210/jc.2009-1429
Unrath M, Thodberg HH, Schweizer R, Ranke MB, Binder G, Martin DD (2012) Automation of bone age reading and a new prediction model improve adult height prediction in children with short stature. Horm Res Paediatr 78:312–319. doi:10.1159/000345875
Visscher PM, Medland SE, Ferreira MA, Morley KI, Zhu G, Cornes BK, Montgomery GW, Martin NG (2006) Assumption-free estimation of heritability from genome-wide identity-by-descent sharing between full siblings. PLoS Genet 2:e41
Walsh S, Lindenbergh A, Zuniga SB, Sijen T, de Knijff P, Kayser M, Ballantyne KN (2011a) Developmental validation of the IrisPlex system: determination of blue and brown iris colour for forensic intelligence. Forensic Sci Int Genet 5:464–471. doi:10.1016/j.fsigen.2010.09.008
Walsh S, Liu F, Ballantyne KN, van Oven M, Lao O, Kayser M (2011b) IrisPlex: a sensitive DNA tool for accurate prediction of blue and brown eye colour in the absence of ancestry information. Forensic Sci Int Genet 5:170–180. doi:10.1016/j.fsigen.2010.02.004
Walsh S, Liu F, Wollstein A, Kovatsi L, Ralf A, Kosiniak-Kamysz A, Branicki W, Kayser M (2012a) The HIrisPlex system for simultaneous prediction of hair and eye colour from DNA. Forensic Sci Int Genet. doi:10.1016/j.fsigen.2012.07.005
Walsh S, Wollstein A, Liu F, Chakravarthy U, Rahu M, Seland JH, Soubrane G, Tomazzoli L, Topouzis F, Vingerling JR, Vioque J, Fletcher AE, Ballantyne KN, Kayser M (2012b) DNA-based eye colour prediction across Europe with the IrisPlex system. Forensic Sci Int Genet 6:330–340. doi:10.1016/j.fsigen.2011.07.009
Weedon MN, Lango H, Lindgren CM, Wallace C, Evans DM, Mangino M, Freathy RM, Perry JR, Stevens S, Hall AS, Samani NJ, Shields B, Prokopenko I, Farrall M, Dominiczak A, Johnson T, Bergmann S, Beckmann JS, Vollenweider P, Waterworth DM, Mooser V, Palmer CN, Morris AD, Ouwehand WH, Zhao JH, Li S, Loos RJ, Barroso I, Deloukas P, Sandhu MS, Wheeler E, Soranzo N, Inouye M, Wareham NJ, Caulfield M, Munroe PB, Hattersley AT, McCarthy MI, Frayling TM (2008) Genome-wide association analysis identifies 20 loci that influence adult height. Nat Genet 40:575–583. doi:10.1038/ng.121
Yang J, Benyamin B, McEvoy BP, Gordon S, Henders AK, Nyholt DR, Madden PA, Heath AC, Martin NG, Montgomery GW, Goddard ME, Visscher PM (2010) Common SNPs explain a large proportion of the heritability for human height. Nat Genet 42:565–569. doi:10.1038/ng.608
Zerath E, Holy X, Mouillon JM, Farbos B, Machwate M, Andre C, Renault S, Marie PJ (1997) TGF-beta2 prevents the impaired chondrocyte proliferation induced by unloading in growth plates of young rats. Life Sci 61:2397–2406
Acknowledgments
The authors are grateful to the study participants, the staff from the Rotterdam Study and the participating general practitioners and pharmacists. We thank Pascal Arp, Mila Jhamai, Marijn Verkerk, Lizbeth Herrera and Marjolein Peters for their help in generating the GWAS database, as well as Karol Estrada and Maksim V. Struchalin for their support in generating and analyzing imputed genomic data. This study was supported in part by funding from the Netherlands Forensic Institute (NFI), by a grant from the Netherlands Genomics Initiative (NGI)/Netherlands Organization for Scientific Research (NWO) within the framework of the Forensic Genomics Consortium Netherlands (FGCN), the Swedish Research Council (K2010-54X-15073-07-3), and by Ferring Pharmaceuticals, Eli Lilly and Company, and Ace Pharmaceuticals. The generation and management of GWAS genotype data for the Rotterdam Study are supported by the Netherlands Organisation of Scientific Research NWO Investments (nr. 175.010.2005.011, 911-03-012). This study is funded by the Research Institute for Diseases in the Elderly (014-93-015; RIDE2), the Netherlands Genomics Initiative/Netherlands Organisation for Scientific Research (NWO) project nr. 050-060-810. The Rotterdam Study is funded by the Erasmus MC University Medical Center Rotterdam, the Erasmus University Rotterdam, the Netherlands Organization for the Health Research and Development (ZonMw), the Research Institute for Diseases in the Elderly (RIDE), the Ministry of Education, Culture and Science of the Netherlands, the Ministry for Health, Welfare and Sports of the Netherlands, the European Commission (DG XII), the Municipality of Rotterdam and the Netherlands Genomics Initiative (NGI)/Netherlands Organization for Scientific Research (NWO) within the framework of the Netherlands Consortium on Healthy Ageing (NCHA). None of the funding agencies had influenced the design, execution or results of this study.
Conflict of interest
SLS Drop has received research grants from Ace, Ferring and Eli Lilly. All other authors declare no conflicts of interest.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
439_2013_1394_MOESM3_ESM.tif
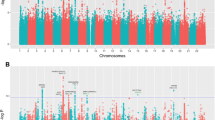
Supplementary Fig. 1a Association result of the case–control designed GWAS for tall stature in 770 tall and 9,591 non-tall Dutch Europeans. A, Manhattan plot. (TIFF 1321 kb)
439_2013_1394_MOESM5_ESM.tif
Supplementary Fig. 2. Performance of six classifiers in predicting tall stature from DNA variants. Box plots of AUC values were generated for weighted allele sums (WAS); least absolute shrinkage and selection operator (LASSO); support vector machine (SVM), neuron networks (NN); classification and regression trees (CART); and random forest (RF). For each method, 1,000 AUC values were derived from cross-validations using randomly split training–testing samples (80 % - 20 %). Summary statistics in the box plots include median (black line), 25-75 % quartiles (grey box), minimum and maximum (dashed interval), and outliers (dots). A. using randomly selected 180 SNPs over the genome. B. using the 180 SNPs from the GIANT paper (Lango Allen et al. 2010). Note that this analysis included 20 % of controls and all cases because the computational burden of SVM scales with sample size. (TIFF 714 kb)
Rights and permissions
About this article
Cite this article
Liu, F., Hendriks, A.E.J., Ralf, A. et al. Common DNA variants predict tall stature in Europeans. Hum Genet 133, 587–597 (2014). https://doi.org/10.1007/s00439-013-1394-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00439-013-1394-0