Abstract
Purpose
Platinum-based chemotherapy is the most common treatment for patients with advanced non-small cell lung cancer (NSCLC). Genetic polymorphisms in the base excision repair (BER) pathway are suspected to influence the response of patients to this type of therapy. In this study, we investigated whether nonsynonymous single nucleotide polymorphisms (SNPs) in the BER pathway were associated with the response, progression-free survival (PFS) and overall survival (OS) of NSCLC patients following platinum-based chemotherapy.
Methods
We used TaqMan to genotype four SNPs (APE1 Asp148Glu, PARP1 Va1762Ala, XRCC1 Arg194Trp and XRCC1 Arg399Gln) in 147 patients with advanced NSCLC who had undergone routine platinum-based chemotherapy.
Results
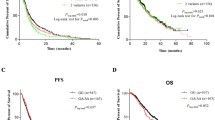
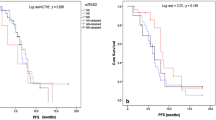
Logistic regression analysis showed that subjects with the XRCC1-399 A allele had a significantly better response to platinum-based chemotherapy than those with the XRCC1-399 GG genotype (AA/AG vs. GG: adjusted OR = 2.35, 95 % CI = 1.11–5.00). Furthermore, multivariate Cox proportional hazard regression analysis showed that the PARP1-762 CC genotype was a significantly unfavorable prognostic factor for PFS (CC vs. CT/TT: adjusted HR = 1.90, 95 % CI = 1.02–3.52). In contrast, the APE1-148 GG genotype was a significantly protective prognostic factor for OS (GG vs. TT: adjusted HR = 0.33, 95 % CI = 0.12–0.92).
Conclusions
We found that XRCC1 Arg399Gln, PARP1 Va1762Ala and APE1 Asp148Glu SNPs in the BER pathway may influence the prognosis of advanced NSCLC patients following platinum-based chemotherapy.


Similar content being viewed by others
References
Parkin DM, Bray F, Ferlay J, Pisani P (2005) Global cancer statistics, 2002. CA Cancer J Clin 55:74–108
Zhang H, Cai B (2003) The impact of tobacco on lung health in China. Respirology 8:17–21
Yang L, Parkin DM, Li L, Chen Y (2003) Time trends in cancer mortality in China: 1987–1999. Int J Cancer 106:771–783
Bahl A, Falk S (2001) Meta-analysis of single agents in the chemotherapy of NSCLC: what do we want to know? Br J Cancer 84:1143–1145
Ludwig JA, Weinstein JN (2005) Biomarkers in cancer staging, prognosis and treatment selection. Nat Rev Cancer 5:845–856
Sweeney C, Nazar-Stewart V, Stapleton PL, Eaton DL, Vaughan TL (2003) Glutathione S-transferase M1, T1, and P1 polymorphisms and survival among lung cancer patients. Cancer Epidemiol Biomarkers Prev 12:527–533
Gurubhagavatula S, Liu G, Park S, Zhou W, Su L, Wain JC, Lynch TJ, Neuberg DS, Christiani DC (2004) XPD and XRCC1 genetic polymorphisms are prognostic factors in advanced non-small-cell lung cancer patients treated with platinum chemotherapy. J Clin Oncol 22:2594–2601
Wu X, Zhao H, Amos CI, Shete S, Makan N, Hong WK, Kadlubar FF, Spitz MR (2002) p53 Genotypes and haplotypes associated with lung cancer susceptibility and ethnicity. J Natl Cancer Inst 94:681–690
Friedberg EC (2003) DNA damage and repair. Nature 421:436–440
Hoogervorst EM, van Steeg H, de Vries A (2005) Nucleotide excision repair- and p53-deficient mouse models in cancer research. Mutat Res 574:3–21
Sancar A, Lindsey-Boltz LA, Unsal-Kacmaz K, Linn S (2004) Molecular mechanisms of mammalian DNA repair and the DNA damage checkpoints. Annu Rev Biochem 73:39–85
Schiller JH, Harrington D, Belani CP, Langer C, Sandler A, Krook J, Zhu J, Johnson DH (2002) Comparison of four chemotherapy regimens for advanced non-small-cell lung cancer. N Engl J Med 346:92–98
Hoang T, Xu R, Schiller JH, Bonomi P, Johnson DH (2003) Clinical model to predict survival in chemonaive patients with advanced non-small-cell lung cancer treated with third-generation chemotherapy regimens based on eastern cooperative oncology group data. J Clin Oncol 23:175–183
Wood RD, Mitchell M, Sgouros J, Lindahl T (2001) Human DNA repair genes. Science 291:1284–1289
Stoehlmacher J, Ghaderi V, Iobal S, Groshen S, Tsao-Wei D, Park D, Lenz HJ (2001) A polymorphism of the XRCC1 gene predicts for response to platinum based treatment in advanced colorectal cancer. Anticancer Res 21:3075–3079
Lockett KL, Hall MC, Xu J, Zheng SL, Berwick M, Chuang SC, Clark PE, Cramer SD, Lohman K, Hu JJ (2004) The ADPRT V762A genetic variant contributes to prostate cancer susceptibility and deficient enzyme function. Cancer Res 64:6344–6348
Fishel ML, Vasko MR, Kelley MR (2007) DNA repair in neurons: so if they don’t divide what’s to repair? Mutat Res 614:24–36
Fan J, Otterlei M, Wong HK, Tomkinson AE, Wilson DM 3rd (2004) XRCC1 co-localizes and physically interacts with PCNA. Nucleic Acids Res 32:2193–2201
Whitehouse CJ, Taylor RM, Thistlethwaite A, Zhang H, Karimi-Busheri F, Lasko DD, Weinfeld M, Caldecott KW (2001) XRCC1 stimulates human polynucleotide kinase activity at damaged DNA termini and accelerates DNA single-strand break repair. Cell 104:107–117
Bernig T, Chanock SJ (2006) Challenges of SNP genotyping and genetic variation: its future role in diagnosis and treatment of cancer. Expert Rev Mol Diagn 6:319–331
Therasse P, Arbuck SG, Eisenhauer EA, Wanders J, Kaplan RS, Rubinstein L, Verweij J, Van Glabbeke M, van Oosterom AT, Christian MC, Gwyther SG (2000) New guidelines to evaluate the response to treatment in solid tumors. European Organization for Research and Treatment of Cancer, National Cancer Institute of the United States, National Cancer Institute of Canada. J Natl Cancer Inst 92:205–216
Lindahl T, Wood RD (1999) Quality control by DNA repair. Science 286:1897–1905
Masson M, Niedergang C, Schreiber V, Muller S, Menissier-de Murcia J, de Murcia G (1998) XRCC1 is specifically associated with poly(ADP-ribose) polymerase and negatively regulates its activity following DNA damage. Mol Cell Biol 18:3563–3571
Giachino DF, Ghio P, Regazzoni S, Mandrile G, Novello S, Selvaggi G, Gregori D, DeMarchi M, Scagliotti GV (2007) Prospective assessment of XPD Lys751Gln and XRCC1 Arg399Gln single nucleotide polymorphisms in lung cancer. Clin Cancer Res 13:2876–2881
Kalikaki A, Kanaki M, Vassalou H, Souglakos J, Voutsina A, Georgoulias V, Mavroudis D (2009) DNA repair gene polymorphisms predict favorable clinical outcome in advanced non-small-cell lung cancer. Clin Lung Cancer 10:118–123
de las Penas R, Sanchez-Ronco M, Alberola V, Taron M, Camps C, Garcia-Carbonero R, Massuti B, Queralt C, Botia M, Garcia-Gomez R, Isla D, Cobo M, Santarpia M, Cecere F, Mendez P, Sanchez JJ, Rosell R (2006) Polymorphisms in DNA repair genes modulate survival in cisplatin/gemcitabine-treated non-small-cell lung cancer patients. Ann Oncol 17:668–675
Shiraishi K, Kohno T, Tanai C, Goto Y, Kuchiba A, Yamamoto S, Tsuta K, Nokihara H, Yamamoto N, Sekine I, Ohe Y, Tamura T, Yokota J, Kunitoh H (2010) Association of DNA repair gene polymorphisms with response to platinum-based doublet chemotherapy in patients with non-small-cell lung cancer. J Clin Oncol 28:4945–4952
Yin Z, Zhou B, He Q, Li M, Guan P, Li X, Cui Z, Xue X, Su M, Ma R, Bai W, Xia S, Jiang Y, Xu S, Lv Y (2009) Association between polymorphisms in DNA repair genes and survival of non-smoking female patients with lung adenocarcinoma. BMC Cancer 9:439
Wang ZH, Miao XP, Tan W, Zhang XR, Xu BH, Lin DX (2004) Single nucleotide polymorphisms in XRCC1 and clinical response to platin-based chemotherapy in advanced non-small cell lung cancer. Ai Zheng 23:865–868
Sun X, Li F, Sun N, Shukui Q, Baoan C, Jifeng F, Lu C, Zuhong L, Hongyan C, YuanDong C, Jiazhong J, Yingfeng Z (2009) Polymorphisms in XRCC1 and XPG and response to platinum-based chemotherapy in advanced non-small cell lung cancer patients. Lung Cancer 65:230–236
Lunn RM, Langlois RG, Hsieh LL, Thompson CL, Bell DA (1999) XRCC1 polymorphisms: effects on aflatoxin B1-DNA adducts and glycophorin A variant frequency. Cancer Res 59:2557–2561
Duell EJ, Wiencke JK, Cheng TJ, Varkonyi A, Zuo ZF, Ashok TD, Mark EJ, Wain JC, Christiani DC, Kelsey KT (2000) Polymorphisms in the DNA repair genes XRCC1 and ERCC2 and biomarkers of DNA damage in human blood mononuclear cells. Carcinogenesis 21:965–971
Matullo G, Palli D, Peluso M, Guarrera S, Carturan S, Celentano E, Krogh V, Munnia A, Tumino R, Polidoro S, Piazza A, Vineis P (2001) XRCC1, XRCC3, XPD gene polymorphisms, smoking and (32)P-DNA adducts in a sample of healthy subjects. Carcinogenesis 22:1437–1445
Hu JJ, Smith TR, Miller MS, Mohrenweiser HW, Golden A, Case LD (2001) Amino acid substitution variants of APE1 and XRCC1 genes associated with ionizing radiation sensitivity. Carcinogenesis 22:917–922
Oei SL, Ziegler M (2000) ATP for the DNA ligation step in base excision repair is generated from poly(ADP-ribose). J Biol Chem 275:23234–23239
Sanderson RJ, Lindahl T (2002) Down-regulation of DNA repair synthesis at DNA single-strand interruptions in poly(ADP-ribose) polymerase-1 deficient murine cell extracts. DNA Repair (Amst) 1:547–558
Kim M, Kang HG, Lee SY, Lee HC, Lee EB, Choi YY, Lee WK, Cho S, Jin G, Jheon HS, Son JW, Lee MH, Jung DK, Cha SI, Kim CH, Kang YM, Kam S, Jung TH, Jheon S, Park JY (2010) Comprehensive analysis of DNA repair gene polymorphisms and survival in patients with early stage non-small-cell lung cancer. Cancer Sci 101:2436–2442
Wang D, Xiang DB, Yang XQ, Chen LS, Li MX, Zhong ZY, Zhang YS (2009) APE1 overexpression is associated with cisplatin resistance in non-small cell lung cancer and targeted inhibition of APE1 enhances the activity of cisplatin in A549 cells. Lung Cancer 66:298–304
Matakidou A, el Galta R, Webb EL, Rudd MF, Bridle H, Eisen T, Houlston RS (2007) Genetic variation in the DNA repair genes is predictive of outcome in lung cancer. Hum Mol Genet 16:2333–2340
Walker LJ, Robson CN, Black E, Gillespie D, Hickson ID (1993) Identification of residues in the human DNA repair enzyme HAP1 (Ref-1) that are essential for redox regulation of Jun DNA binding. Mol Cell Biol 13:5370–5376
Su D, Ma S, Liu P, Jiang Z, Lv W, Zhang Y, Deng Q, Smith S, Yu H (2007) Genetic polymorphisms and treatment response in advanced non-small cell lung cancer. Lung Cancer 56:281–288
Acknowledgments
This work was supported in part by grants from the Education Department of Jiangsu Province Green Blue Project (2010), the Scientific Research Foundation for the Returned Overseas Chinese Scholars, State Education Ministry 2009 (IA09), the National Natural Science Foundation of China (81071643), the National Natural Science Foundation of China (30772549), the Surface Project of the Health Department of Jiangsu Province (H201001) and the Innovation Project of Jiangsu Province.
Author information
Authors and Affiliations
Corresponding authors
Additional information
Wan Zhao, Lingmin Hu and Jiali Xu contributed equally to this work.
Rights and permissions
About this article
Cite this article
Zhao, W., Hu, L., Xu, J. et al. Polymorphisms in the base excision repair pathway modulate prognosis of platinum-based chemotherapy in advanced non-small cell lung cancer. Cancer Chemother Pharmacol 71, 1287–1295 (2013). https://doi.org/10.1007/s00280-013-2127-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00280-013-2127-8