Abstract
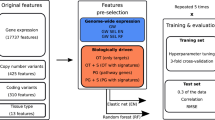
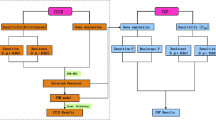
Prior work has shown that the sensitivity of a tumour to a specific drug can be predicted from a molecular signature of gene expressions. This is an important finding for improving drug efficacy and personalizing drug use. In this paper, we present an analysis strategy that, compared to prior work, maintains more information and leads to improved chemosensitivity prediction. Specifically we show (a) that prediction is improved when the GI50 value of a drug is estimated by all available measurements and fitting a sigmoid curve and (b) application of regression techniques often results in more accurate models compared to classification techniques. In addition, we show that (c) modern variable selection techniques, such as MMPC result in better predictive performance than simple univariate filtering. We demonstrate the strategy on 59 tumor cell lines after treatment with 118 fully characterized drugs obtained by the National Cancer Institute (NCI 60 screening) and biologically comment on the identified molecular signatures of the best predicted drugs.
Chapter PDF
Similar content being viewed by others
References
Potti, A., Dressman, H.K., Bild, A., Riedel, R.F., et al.: Genomic signatures to guide the use of chemotherapeutics (2006)
Augustin, C.K., Yoo, J.S., Potti, A., Yoshimoto, Y., et al.: Genomic and molecular profiling predicts response to temozolomide in melanoma. Clinical Cancer Res. 15(2) (2009)
Staunton, J.E., Slonim, D.K., Coller, H.A., et al.: Chemosensitivity prediction by transcriptional profiling. Proc. Natl. Acad. Sci. 98(19), 10787–10792 (2001)
Ma, Y., Ding, Z., Qian, Y., et al.: An integrative genomic and proteomic approach to chemosensitivity prediction. Int. J. Oncol. 34(1), 107–115 (2009)
Tsamardinos, I., Brown, L.E., Aliferis, C.F.: The max-min hill-climbing bayesian network structure learning algorithm. Journal of Machine Learning 65, 31–78 (2006)
Shankavaram, U.T., Varma, S., Kane, D., et al.: Cellminer: a relational database and query tool for the nci-60 cancer cell lines. BMC Genomics 10(277) (2009)
http://dtp.nci.nih.gov/docs/compare/compare_methodology.html
http://www.itl.nist.gov/div898/handbook/eda/section3/eda35h.htm
Aliferis, C.F., Statnikov, A., Tsamardinos, I., Mani, S., Koutsoukos, X.D.: Local causal and markov blanket induction for causal discovery and feature selection for classification part i: Algorithms and empirical evaluation. Journal of Machine Learning Research, Special Topic on Causality 11, 171–234 (2010)
Statnikov, A., Tsamardinos, T., Brown, L.E., Aliferis, C.F.: Causal explorer: A matlab library of algorithms for causal discovery and variable selection for classification. Challenges in Causality 1 (2009)
Chang, C.C., Lin, C.J.: LIBSVM: a library for support vector machines (2001), software http://www.csie.ntu.edu.tw/~cjlin/libsvm
Steel, R.G.D., Torrie, J.H.: Principles and Procedures of Statistics. McGraw-Hill, New York (1960)
Boser, B., Guyon, I., Vapnik, V.: An training algorithm for optimal margin classifiers. In: Fifth Annual Workshop on Computational Learning Theory, pp. 144–152 (1992)
Statnikov, A., Aliferis, C.F., Tsamardinos, I., Hardin, D., Levy, S.: A comprehensive evaluation of multicategory classification methods for microarray gene expression cancer diagnosis. Bioinformatics 21(5), 631–643 (2005)
Egyhazi, S., Bergh, J., Hansson, J., Karran, P., Ringborg, U.: Carmustine-induced toxicity, dna crosslinking and o6-methylguanine-dna methyltransferase activity in two human lung cancer cell lines. Eur. J. Cancer. 27, 1658–1662 (1991)
Waanders, E., van der Velden, V.H., van der Schoot, C.E., van Leeuwen, F.N., et al.: Integrated use of minimal residual disease classification and ikzf1 alteration status accurately predicts 79% of relapses in pediatric acute lymphoblastic leukemia. Leukemia 25, 254–258 (2011)
Zuo, Z., Jones, D., Yao, H., et al.: A pathway-based gene signature correlates with therapeutic response in adult patients with philadelphia chromosome-positive acute lymphoblastic leukemia. Mod. Pathol. 23, 1524–1534 (2010)
Laity, J.H., Lee, B.M., Wright, P.E.: Zinc finger proteins: new insights into structural and functional diversity. Curr. Opin. Struct. Biol 11(1), 39–46 (2001)
Meili, D., Kralovicova, J., Zagalak, J., Bonafe, L., et al.: Disease-causing mutations improving the branch site and polypyrimidine tract: pseudoexon activation of line-2 and antisense alu lacking the poly(t)-tail. Hum. Mutat. 30, 823–831 (2009)
Babst, M., Katzmann, D.J., Estepa-Sabal, E.J., Meerloo, T., Emr, S.D.: Escrt-iii: an endosome-associated heterooligomeric protein complex required for mvb sorting. Dev. Cell. 3, 271–282 (2002)
Rusten, T.E., Simonsen, A.: Escrt functions in autophagy and associated disease. Cell Cycle 7, 1166–1172 (2008)
Tso, P.H., Wang, Y., Wong, S.Y., Poon, L.S., Chan, A.S., et al.: Rgs19 enhances cell proliferation through its c-terminal pdz motif. Cell Signal 22, 1700–1707 (2010)
Maris, J.M., Mosse, Y.P., Bradfield, J.P., et al.: Chromosome 6p22 locus associated with clinically aggressive neuroblastoma. N. Engl. J. Med. 358 (2008)
Dunwell, T., Hesson, L., Rauch, T.A., Wang, L., et al.: A genome-wide screen identifies frequently methylated genes in haematological and epithelial cancers. Mol Cancer 9(44) (2010)
Mariani, L., McDonough, W.S., Hoelzinger, D.B., Beaudry, C., Kaczmarek, E., et al.: Identification and validation of p311 as a glioblastoma invasion gene using laser capture microdissection. Cancer Res. 61, 4190–4196 (2001)
Saletta, F., Rahmanto, Y.S., Richardson, D.R.: The translational regulator eif3a: the tricky eif3 subunit! Biochim Biophys Acta 1806(2), 275–286 (2010)
Author information
Authors and Affiliations
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2011 International Federation for Information Processing
About this paper
Cite this paper
Christodoulou, E.G., Røe, O.D., Folarin, A., Tsamardinos, I. (2011). Information-Preserving Techniques Improve Chemosensitivity Prediction of Tumours Based on Expression Profiles. In: Iliadis, L., Jayne, C. (eds) Engineering Applications of Neural Networks. EANN AIAI 2011 2011. IFIP Advances in Information and Communication Technology, vol 363. Springer, Berlin, Heidelberg. https://doi.org/10.1007/978-3-642-23957-1_50
Download citation
DOI: https://doi.org/10.1007/978-3-642-23957-1_50
Publisher Name: Springer, Berlin, Heidelberg
Print ISBN: 978-3-642-23956-4
Online ISBN: 978-3-642-23957-1
eBook Packages: Computer ScienceComputer Science (R0)