Abstract
Purpose
This study aimed to identify novel molecular markers for the early detection of colorectal cancer liver metastasis.
Methods
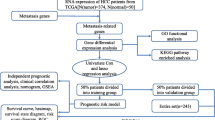
Genes related to hepatic metastasis were screened from the Oncomine database. The candidate markers were tested by immunohistochemistry, and their predictive accuracy was assessed by the cross-validation method and an independent test set.
Results
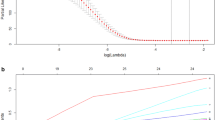
We got three datasets containing 2,973 genes that were highly expressed in primary colon cancer tissues compared with non-metastatic colon cancer tissues and identified 7 candidate molecules for immunohistochemical validation. A total of 213 colorectal cancer samples were randomly divided into a training set (113 cases) and a blind testing set (100 cases). In the training set, immunohistochemical analysis showed that HP, OPN, and PTGIS expression were significantly higher in the hepatic metastasis group than in the non-metastasis group. Logistic regression analysis showed that HP and OPN levels in primary tumors and lymph node metastasis status were the only significant (P < 0.05) parameters for detecting liver metastasis. The predictive accuracy of markers was assessed by the cross-validation method and an independent test set. In leave-one-out cross-validation, the two markers combined with clinicopathologic features resulted in 91.2% sensitivity and 96.4% specificity for hepatic metastasis detection. In an independent test set, the combination achieved 94.5% sensitivity and 88.9% specificity for predicting the hepatic metastasis of colorectal cancer.
Conclusions
Our results suggest that combined HP and OPN expression levels are significantly related to liver metastasis and prognosis, and, if this is validated, they could be used as accurate predictors of liver metastasis in colorectal cancer.


Similar content being viewed by others
References
Parkin DM, Bray F, Ferlay F, Pisani P. Global cancer statistics, 2002. CA Cancer J Clin. 2005;55:74–108.
Manfredi S, Lepage C, Hatem C, Coatmeur O, Faivre J, Bouvier AM. Epidemiology and management of liver metastases from colorectal cancer. Ann Surg. 2006;244:254–9.
McMillan DC, McArdle CS. Epidemiology of colorectal liver metastases. Surg Oncol. 2007;16:3–5.
Angelopoulos S, Kanellos I, Christophoridis E, Tsachalis T, Kanellou A, Betsis D. Five-year survival after curative resection for adenocarcinoma of the colon. Tech Coloproctol. 2004;8(Suppl. 1):S152–4.
Bakalakos EA, Kim JA, Young DC, Martin EW Jr. Determinants of survival following hepatic resection for metastatic colorectal cancer. World J Surg. 1998;22:399–404; discussion 404–5.
Di Martino E, Wild CP, Rotimi O, Darnton JS, Olliver RJ, Hardie LJ. IGFBP-3 and IGFBP-10 (CYR61) up-regulation during the development of Barrett’s oesophagus and associated oesophageal adenocarcinoma: potential biomarkers of disease risk. Biomarkers. 2006;11:547–61.
Yamazoe Y, Maetani S, Onodera H, Nishikawa T, Tobe T. Histopathological prediction of liver metastasis after curative resection of colorectal cancer. Surg Oncol. 1992;1:237–44.
Bird NC, Mangnall D, Majeed AW. Biology of colorectal liver metastases: a review. J Surg Oncol. 2006;94:68–80.
Ohji Y, Yao T, Eguchi T, et al. Evaluation of risk of liver metastasis in colorectal adenocarcinoma based on the combination of risk factors including CD10 expression: multivariate analysis of clinicopathological and immunohistochemical factors. Oncol Rep. 2007;17:525–30.
Barozzi C, Ravaioli M, D’Errico A, et al. Relevance of biologic markers in colorectal carcinoma: a comparative study of a broad panel. Cancer. 2002;94:647–57.
Takata O, Kawamura YJ, Konishi F, et al. cDNA array analysis for prediction of hepatic metastasis of colorectal carcinoma. Surg Today. 2006;36:608–14.
Jemal A, Bray F, Center MM, Ferlay J, Ward E, Forman D. Global cancer statistics. CA Cancer J Clin. 2011;61:69–90.
Li M, Lin YM, Hasegawa S, et al. Genes associated with liver metastasis of colon cancer, identified by genome-wide cDNA microarray. Int J Oncol. 2004;24:305–12.
Rhodes DR, Yu J, Shanker K, et al. ONCOMINE: a cancer microarray database and integrated data-mining platform. Neoplasia. 2004;6:1–6.
Marquardt JU, Quasdorff M, Varnholt H, et al. Neighbor of punc E11, a novel oncofetal marker for hepatocellular carcinoma. Int J Cancer. 2010.
Monami G, Emiliozzi V, Bitto A, et al. Proepithelin regulates prostate cancer cell biology by promoting cell growth, migration, and anchorage-independent growth. Am J Pathol. 2009;174:1037–47.
Smith DD, Saetrom P, Snove OJ, et al. Meta-analysis of breast cancer microarray studies in conjunction with conserved cis-elements suggest patterns for coordinate regulation. BMC Bioinformatics. 2008;9:63.
Yang Y, Pospisil P, Iyer LK, Adelstein SJ, Kassis AI. Integrative genomic data mining for discovery of potential blood-borne biomarkers for early diagnosis of cancer. PLoS One. 2008;3:e3661.
Tanami H, Tsuda H, Okabe S, et al. Involvement of cyclin D3 in liver metastasis of colorectal cancer, revealed by genome-wide copy-number analysis. Lab Invest. 2005;85:1118–29.
Wanebo HJ, Chu QD, Vezeridis MP, Soderberg C. Patient selection for hepatic resection of colorectal metastases. Arch Surg. 1996;131:322–9.
Ramaswamy S, Ross KN, Lander ES, Golub TR. A molecular signature of metastasis in primary solid tumors. Nat Genet. 2003;33:49–54.
Hu H, Sun L, Guo C, et al. Tumor cell-microenvironment interaction models coupled with clinical validation reveal CCL2 and SNCG as two predictors of colorectal cancer hepatic metastasis. Clin Cancer Res. 2009;15:5485–93.
Macri A, Versaci A, Lupo G, et al. Role of osteopontin in breast cancer patients. Tumori. 2009;95:48–52.
Sun Y, Wang L, Jiang M, Huang J, Liu A, Wolfl S. Secreted phosphoprotein 1 upstream invasive network construction and analysis of lung adenocarcinoma compared with human normal adjacent tissues by integrative biocomputation. Cell Biochem Biophys. 2010;56:59–71.
Yu HB, Han XB, Liang YQ, Liu JG, Wang H. [Expression of osteopontin and survivin in prostate cancer and the clinical significance]. Nan Fang Yi Ke Da Xue Xue Bao. 2010;30:1141–3.
Coppola D, Szabo M, Boulware D, et al. Correlation of osteopontin protein expression and pathological stage across a wide variety of tumor histologies. Clin Cancer Res. 2004;10(1 Pt. 1):184–90.
Lee JL, Wang MJ, Sudhir PR, Chen GD, Chi CW, Chen JY. Osteopontin promotes integrin activation through outside-in and inside-out mechanisms: OPN-CD44 V interaction enhances survival in gastrointestinal cancer cells. Cancer Res. 2007;67:2089–97.
Allan AL, George R, Vantyghem SA, et al. Role of the integrin-binding protein osteopontin in lymphatic metastasis of breast cancer. Am J Pathol. 2006;169(1):233–46.
Ramankulov A, Lein M, Kristiansen G, Meyer HA, Loening SA, Jung K. Elevated plasma osteopontin as marker for distant metastases and poor survival in patients with renal cell carcinoma. J Cancer Res Clin Oncol. 2007;133(9):643–52.
Huntoon KM, Wang Y, Eppolito CA, et al. The acute phase protein haptoglobin regulates host immunity. J Leukoc Biol. 2008;84:170–81.
Abdullah M, Schultz H, Kahler D, et al. Expression of the acute phase protein haptoglobin in human lung cancer and tumor-free lung tissues. Pathol Res Pract. 2009;205:639–47.
Smeets MB, Fontijn J, Kavelaars A, Pasterkamp J, De Kleijn DPV. The acute phase protein haptoglobin is locally expressed in arthritic and oncological tissues. Int J Exp Pathol. 2003;84:69–74.
Harvey S, Kohga S, Sait SN, et al. Co-expression of urokinase with haptoglobin in human carcinomas. J Surg Res. 2009;152:189–97.
Lee CC, Ho HC, Lee MS, Hung SK, Yu CC, Su YC. Expression of haptoglobin predicts recurrence in head and neck squamous cell carcinoma. Clin Chim Acta. 2010;411:1116–21.
Acknowledgment
This study was supported by The National Key Basic Research Program of China (2009CB521804 and 2010CB933902) and by PUMC&CAMS Funds for Young Scholar.
Author information
Authors and Affiliations
Corresponding authors
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Sun, L., Pan, J., Peng, L. et al. Combination of Haptoglobin and Osteopontin Could Predict Colorectal Cancer Hepatic Metastasis. Ann Surg Oncol 19, 2411–2419 (2012). https://doi.org/10.1245/s10434-011-2177-2
Received:
Published:
Issue Date:
DOI: https://doi.org/10.1245/s10434-011-2177-2