Abstract
Background
Pancreatic cancer is an exceptionally lethal disease with an annual mortality nearly equivalent to its annual incidence. This dismal rate of survival is due to several factors including late presentation with locally advanced, unresectable tumors, early metastatic disease, and rapidly arising chemoresistance. To study the mechanisms of chemoresistance in pancreatic cancer we developed two gemcitabine-resistant pancreatic cancer cell lines.
Methods
Resistant cells were obtained by culturing L3.6pl and AsPC-1 cells in serially increasing concentrations of gemcitabine. Stable cultures were obtained that were 40- to 50-fold increased in resistance relative to parental cells. Immunofluorescent staining was performed to examine changes in β-catenin and E-cadherin localization. Protein expression was determined by immunoblotting. Migration and invasion were determined by modified Boyden chamber assays. Fluorescence-activated cell sorting (FACS) analyses were performed to examine stem cell markers.
Results
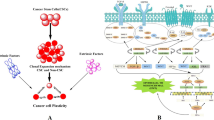
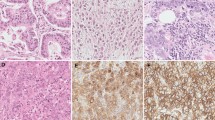
Gemcitabine-resistant cells underwent distinct morphological changes, including spindle-shaped morphology, appearance of pseudopodia, and reduced adhesion characteristic of transformed fibroblasts. Gemcitabine-resistant cells were more invasive and migratory. Gemcitabine-resistant cells were increased in vimentin and decreased in E-cadherin expression. Immunofluorescence and immunoblotting revealed increased nuclear localization of total β-catenin. These alterations are hallmarks of epithelial-to-mesenchymal transition (EMT). Resistant cells were activated in the receptor protein tyrosine kinase, c-Met and increased in expression of the stem cell markers CD (cluster of differentiation)24, CD44, and epithelial-specific antigen (ESA).
Conclusions
Gemcitabine-resistant pancreatic tumor cells are associated with EMT, a more-aggressive and invasive phenotype in numerous solid tumors. The increased phosphorylation of c-Met may also be related to chemoresistance and EMT and presents as an attractive adjunctive chemotherapeutic target in pancreatic cancer.




Similar content being viewed by others
References
Burris HA 3rd, Moore MJ, Andersen J, et al. Improvements in survival and clinical benefit with gemcitabine as first-line therapy for patients with advanced pancreas cancer: a randomized trial. J Clin Oncol 1997;15:2403–13
Moore MJ, Hamm J, Dancey J, et al. Comparison of gemcitabine versus the matrix metalloproteinase inhibitor BAY 12-9566 in patients with advanced or metastatic adenocarcinoma of the pancreas: A phase III trial of the national cancer institute of Canada clinical trials group. J Clin Oncol 2003;17:3296–302
Yang AD, Fan F, Camp ER, et al. Chronic oxaliplatin resistance induces epithelial-to-mesenchymal transition in colorectal cancer cell lines. Clin Cancer Res 2006;12:4147–53
Hiscox S, Jiang WG, Obermeier K, et al. Tamoxifen resistance in MCF7 cells promotes EMT-like behaviour and involves modulation of beta-catenin phosphorylation. Int J Cancer 2006;118:290–301
Hiscox S, Morgan L, Barrow D, et al. Tamoxifen resistance in breast cancer cells is accompanied by an enhanced motile and invasive phenotype: inhibition by gefitinib (‘Iressa’, ZD1839). Clin Exp Metastasis 2004;21:201–12
Thiery JP. Epithelial-mesenchymal transitions in tumour progression. Nat Rev Cancer 2002;2:442–54
Thiery JP, Sleeman JP. Complex networks orchestrate epithelial mesenchymal transitions. Nat Rev Mol Cell Biol 2006;7:131–42
Boyer B, Valles AM, Edme N. Induction and regulation of epithelial to mesenchymal transitions. Biochem Pharm 2000;60:1091–9
Savagner P. Leaving the neighborhood: molecular mechanisms involved during epithelial to mesenchymal transition. BioEssays 2001;23:912–23
Grotegut S, von Schweinitz D, Christofori G, Lehembre F. Hepatocyte growth factor induces cell scattering through MAPK/Egr-1-mediated upregulation of Snail. EMBO J 2006;25:3534–45
Jeffers M, Rong S, Vande Woude GF. Enhanced tumorigenicity and invasion-metastasis by hepatocyte growth factor/scatter factor-met signalling in human cells concomitant with induction of the urokinase proteolysis network. Mol Cell Biol 1996;16:1115–25
Bladt F, Riethmacher D, Isenmann S, Aguzzi A, Birchmeier C. Essential role for the c-met receptor in the migration of myogenic precursor cells into the limb bud. Nature 1995;376:768–71
Schmidt C, Bladt F, Goedecke S, et al. Scatter factor/hepatocyte growth factor is essential for liver development. Nature 1995;373:699–702
Matsumoto K, Nakamura T. HGF: its organotrophic role and therapeutic potential. Ciba Found Symp 1997;212:198–211
Miyazawa K, Shimomura T, Naka D, Kitamura N. Proteolytic activation of hepatocyte growth factor in response to tissue injury. J Biol Chem 1994;269:8966–70
Jiang W, Hiscox S, Matsumoto K, Nakamura T. Hepatocyte growth factor/scatter factor, its molecular, cellular and clinical implications in cancer. Crit Rev Oncol Hematol 1999;29:209–48
Herynk MH, Tsan R, Radinsky R, Gallick GE. Activation of c-Met in colorectal carcinoma cells leads to constitutive association of tyrosine-phosphorylated beta-catenin. Clin Exp Metastasis 2003;20:291–300
Gray MJ, Zhang J, Ellis LM, et al. HIF-1alpha, STAT3, CBP/p300 and Ref-1/APE are components of a transcriptional complex that regulates Src-dependant hypoxia-induced expression of VEGF in pancreatic and prostate carcinomas. Oncogene 2005;24:3110–20
Summy JM, Trevino JG, Baker CH, et al. c-Src regulates constitutive and EGF-mediated VEGF expression in pancreatic tumor cells through activation of phosphatidyl inosityol 3-kinase and p38 MAPK. Pancreas 2005;31:263–74
Rasola A, Fassetta M, De Bacco F, D’Alessandro L, Gramaglia D, Di Renzo MF, Comoglio PM. A positive feedback loop between hepatocyte growth factor receptor and beta-catenin sustains colorectal cancer cell invasive growth. Oncogene 2007;26:1078–87
Li C, Heidt DG, Dalerba P, et al. Identification of pancreatic cancer stem cells. Cancer Res 2007;67:1030–7
La Porta CA. Drug resistance in melanoma: new perspectives. Curr Med Chem 2007;14:387–91
Donnenberg VS, Donnenberg AD. Multiple drug resistance in cancer revisited: the cancer stem cell hypothesis. J Clin Pharmacol 2005;45:872–7
Jemal A, Tiwari RC, Murray T, et al. Cancer statistics, 2004. CA Cancer J Clin 2004;54:8–29
Nakano Y, Tanno S, Koizumi K, et al. Gemcitabine chemoresistance and molecular markers associated with gemcitabine transport and metabolism in human pancreatic cancer cells. Br J Cancer 2007;96:457–63
Shah AN, Gallick GE. Src, chemoresistance and epithelial to mesenchymal transition: are they related? Anticancer Drugs 2007;18:371–5
Shibamoto S, Hayakawa M, Takeuchi K, et al. Tyrosine phosphorylation of beta-catenin and plakoglobin enhanced by hepatocyte growth factor and epidermal growth factor in human carcinoma cells. Cell Adhes Commun 1994;1:295–305
Hiscox S, Jiang WG. Association of the HGF/SF receptor, c-met, with the cell-surface adhesion molecule, E-cadherin, and catenins in human tumor cells. Biochem Biophys Res Commun 1999;261:406–11
Monga SP, Mars WM, Pediaditakis P, et al. Hepatocyte growth factor induces Wnt-independent nuclear translocation of beta-catenin after Met-beta-catenin dissociation in hepatocytes. Cancer Res 2002;62:2064–71
Yang J, Mani SA, Donaher JL, et al. Twist, a master regulator of morphogenesis, plays an essential role in tumor metastasis. Cell 2004;117:927–39
Kang Y, Massague J. Epithelial-mesenchymal transitions: twist in development and metastasis. Cell 2004;118:277–9
Howe LR, Watanabe O, Leonard J, Brown AM. Twist is up-regulated in response to Wnt1 and inhibits mouse mammary cell differentiation. Cancer Res 2003;63:1906–13
Hoek K, Rimm DL, Williams KR, et al. Expression profiling reveals novel pathways in the transformation of melanocytes to melanomas. Cancer Res 2004;64:5270–82
Cheng GZ, Chan J, Wang Q, Zhang W, Sun CD, Wang LH. Twist transcriptionally up-regulates AKT2 in breast cancer cells leading to increased migration, invasion, and resistance to paclitaxel. Cancer Res 2007;67:1979–87
Minard MM, Herynk MH, Collard JG, Gallick GE. The guanine nucleotide exchange factor Tiam1 increases colon carcinoma growth at metastatic sites in an orthotopic nude mouse model. Oncogene 2005; 24:2568–2573
Acknowledgments
This research was supported in part by NIH U54 CA 090810 (G.E.G), NIH T32 CA 09599 (A.N.S.), the Eleanor B. Pillsbury Fellowship-University of Illinois Hospital (A.N.S.), the Lockton Fund (G.E.G., J.M.S.), and G.E.G is the Sowell-Huggins Professor and J.Z. the Sowell-Huggins Fellow in Cancer Biology.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Shah, A.N., Summy, J.M., Zhang, J. et al. Development and Characterization of Gemcitabine-Resistant Pancreatic Tumor Cells. Ann Surg Oncol 14, 3629–3637 (2007). https://doi.org/10.1245/s10434-007-9583-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1245/s10434-007-9583-5