Abstract
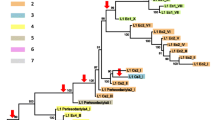
A large-scale study of short retroposon (SINE) B1 has been conducted in the genome of rodents from most of the known families of this mammalian order. The B1 nucleotide sequences of rodents from different families exhibited a number of characteristic features including substitutions, deletions, and tandem duplications. Comparing the distribution of these features among the rodent families, the currently discussed phylogenetic relationships were tested. The results of analysis indicated (1) an early divergence of Sciuridae and related families (Aplodontidae and Gliridae) from the other rodents; (2) a possible subsequent divergence of beavers (Castoridae); (3) a monophyletic origin of the group Hystricognathi, which includes several families, such as porcupines (Hystricidae) and guinea pigs (Caviidae); (4) a possible monophyletic origin of the group formed by the remaining families, including six families of mouselike rodents (Myodonta). Various approaches to the use of short retroposons for phylogenetic studies are discussed.
Similar content being viewed by others
References
Deininger, P.L. and Batzer, M.A., Mammalian Retroelements, Genome Res., 2002, vol. 12, no. 10, pp. 1455–1465.
Kramerov, D. and Vassetzky, N., Short Retroposons in Eukaryotic Genomes, Int. Rev. Cytol., 2005, vol. 247, pp. 165–221.
Okada, N., SINEs, Curr. Opin. Genet. Dev., 1991, vol. 1, no. 4, pp. 498–504.
Kapitonov, V.V. and Jurka, J., A Novel Class of SINE Elements Derived from 5S rRNA, Mol. Biol. Evol., 2003, vol. 20, no. 5, pp. 694–702.
Nishihara, H., Smit, A.F., and Okada, N., Functional Noncoding Sequences Derived from SINEs in the Mammalian Genome, Genome Res., 2006, vol. 16, no. 7, pp. 864–874.
Kramerov, D.A., Grigoryan, A.A., Ryskov, A.P., et al., Long Double-Stranded Sequences (dsRNA-B) of Nuclear Pre-mRNA Consist of a Few Highly Abundant Classes of Sequences: Evidence from DNA Cloning Experiments, Nucleic Acids Res, 1979, vol. 6, no. 2, pp. 697–713.
Krayev, A.S., Kramerov, D.A., Skryabin, K.G., et al., The Nucleotide Sequence of the Ubiquitous Repetitive DNA Sequence B1 Complementary to the Most Abundant Class of Mouse Fold-Back RNA, Nucleic Acids Res., 1980, vol. 8, no. 6, pp. 1201–1215.
Deininger, P.L., Jolly, D.J., Rubin, C.M., et al., Base Sequence Studies of 300 Nucleotide Renatured Repeated Human DNA Clones, J. Mol. Biol., 1981, vol. 151, no. 1, pp. 17–33.
Haynes, S.R., Toomey, T.P., Leinwand, L., et al., The Chinese Hamster Alu-Equivalent Sequence: A Conserved Highly Repetitious, Interspersed Deoxyribonucleic Acid Sequence in Mammals Has a Structure Suggestive of a Transposable Element, Mol. Cell. Biol., 1981, vol. 1, no. 7, pp. 573–583.
Ullu, E. and Tschudi, C., Alu Sequences Are Processed 7SL RNA Genes, Nature, 1984, vol. 312, no. 5990, pp. 171–172.
Daniels, G.R. and Deininger, P.L., A Second Major Class of Alu Family Repeated DNA Sequences in a Primate Genome, Nucleic Acids Res., 1983, vol. 11, no. 21, pp. 7595–7610.
Zietkiewicz, E., Richer, C., Sinnett, D., et al., Monophyletic Origin of Alu Elements in Primates, J. Mol. Evol., 1998, vol. 47, no. 2, pp. 172–182.
Roos, C., Schmitz, J., and Zischler, H., Primate Jumping Genes Elucidate Strepsirrhine Phylogeny, Proc. Natl. Acad. Sci. USA, 2004, vol. 101, no. 29, pp. 10 650–10 654.
Labuda, D., Sinnett, D., Richer, C., et al., Evolution of Mouse B1 Repeats: 7SL RNA Folding Pattern Conserved, J. Mol. Evol., 1991, vol. 32, no. 5, pp. 405–414.
Quentin, Y., A Master Sequence Related to a Free Left Alu Monomer (FLAM) at the Origin of the B1 Family in Rodent Genomes, Nucleic Acids Res., 1994, vol. 22, no. 12, pp. 2222–2227.
Hartenberger, J.-L., The Order Rodentia: Major Questions on Their Evolutionary Origin, Relationships and Suprafamilial Systematics, in Evolutionary Relationship among Rodents, Luckett, W. and Hartenberger, J.-L., Eds., New York: Plenum, 1985, pp. 1–33.
Pavlinov, I., Sistematika sovremennykh mlekopitayushchikh (Systematics of Contemporary Mammals), Moscow: Mosk. Gos. Univ., 2003.
Krayev, A.S., Markusheva, T.V., Kramerov, D.A., et al., Ubiquitous Transposon-Like Repeats B1 and B2 of the Mouse Genome: B2 Sequencing, Nucleic Acids Res., 1982, vol. 10, no. 23, pp. 7461–7475.
Den Dunnen, J.T. and Schoenmakers, J.G., Consensus Sequences of the Rattus norvegicus B1 and B2 Repeats, Nucleic Acids Res., 1987, vol. 15, no. 6, p. 2772.
Bains, W. and Temple-Smith, K., Similarity and Divergence among Rodent Repetitive DNA Sequences, J. Mol. Evol., 1989, vol. 28, no. 3, pp. 191–199.
Waterston, R.H., Lindblad-Toh, K., Birney, E., et al., Initial Sequencing and Comparative Analysis of the Mouse Genome, Nature, 2002, vol. 420, no. 6915, pp. 520–562.
Gibbs, R.A., Weinstock, G.M., Metzker, M.L., et al., Genome Sequence of the Brown Norway Rat Yields Insights into Mammalian Evolution, Nature, 2004, vol. 428, no. 6982, pp. 493–521.
Kim, J., Martignetti, J.A., Shen, M.R., et al., Rodent BC1 RNA Gene as a Master Gene for ID Element Amplification, Proc. Natl. Acad. Sci. USA, 1994, vol. 91, no. 9, pp. 3607–3611.
Serdobova, I.M. and Kramerov, D.A., Short Retroposons of the B2 Superfamily: Evolution and Application for the Study of Rodent Phylogeny, J. Mol. Evol., 1998, vol. 46, pp. 202–214.
Kramerov, D.A. and Vassetzky, N.S., Structure and Origin of a Novel Dimeric Retroposon B1-dID, J. Mol. Evol., 2001, vol. 52, no. 2, pp. 137–143.
Vassetzky, N.S., Ten, O.A., and Kramerov, D.A., B1 and Related SINEs in Mammalian Genomes, Gene, 2003, vol. 319, pp. 149–160.
Nishihara, H., Terai, Y., and Okada, N., Characterization of Novel Alu-and tRNA-Related SINEs from the Tree Shrew and Evolutionary Implications of Their Origins, Mol. Biol. Evol., 2002, vol. 19, no. 11, pp. 1964–1972.
Serdobova, I.M. and Kramerov, D.A., Usage of Short Retroposons as Phylogenetic Markers, Dokl. Akad. Nauk SSSR, 1994, vol. 335, no. 5, pp. 664–667.
Kramerov, D., Vassetzky, N., and Serdobova, I., The Evolutionary Position of Dormice (Gliridae) in Rodentia Determined by a Novel Short Retroposon, Mol. Biol. Evol., 1999, pp. 715–716.
Felsenstein, J., Phylogenies from Gene Frequencies: A Statistical Problem, Systematic Zool., 1985, vol. 34, no. 3, pp. 300–311.
Felsenstein, J., PHYLIP—Phylogeny Inference Package (Version 3.2), Cladistics, 1989, vol. 5, pp. 164–166.
Huchon, D. and Douzery, E.J., From the Old World to the New World: A Molecular Chronicle of the Phylogeny and Biogeography of Hystricognath Rodents, Mol. Phylogenet. Evol., 2001, vol. 20, no. 2, pp. 238–251.
Huchon, D., Madsen, O., Sibbald, M.J., et al., Rodent Phylogeny and a Timescale for the Evolution of Glires: Evidence from an Extensive Taxon Sampling Using Three Nuclear Genes, Mol. Biol. Evol., 2002, vol. 19, no. 7, pp. 1053–1065.
Adkins, R.M., Gelke, E.L., Rowe, D., et al., Molecular Phylogeny and Divergence Time Estimates for Major Rodent Groups: Evidence from Multiple Genes, Mol. Biol. Evol., 2001, vol. 18, no. 5, pp. 777–791.
Adkins, R.M., Walton, A.H., and Honeycutt, R.L., Higher-Level Systematics of Rodents and Divergence Time Estimates Based on Two Congruent Nuclear Genes, Mol. Phylogenet. Evol., 2003, vol. 26, no. 3, pp. 409–420.
Steppan, S., Adkins, R., and Anderson, J., Phylogeny and Divergence-Date Estimates of Rapid Radiations in Muroid Rodents Based on Multiple Nuclear Genes, Syst. Biol., 2004, vol. 53, no. 4, pp. 533–553.
Opazo, J.C., A Molecular Timescale for Caviomorph Rodents (Mammalia, Hystricognathi), Mol. Phylogenet. Evol., 2005, vol. 37, no. 3, pp. 932–937.
Quentin, Y., Emergence of Master Sequences in Families of Retroposons Derived from 7sl RNA, Genetics, 1994, vol. 93, nos. 1–3, pp. 203–215.
Carrol, R., Vertebrate Paleontology and Evolution, New York: Freeman, 1988.
Reyes, A., Pesole, G., and Saccone, C., Long-Branch Attraction Phenomenon and the Impact of Among-Site Variation on Rodent Phylogeny, Gene, 2000, vol. 259, nos. 1–2, pp. 177–187.
D’erchia, A.M., Gissi, C., Pesole, G., et al., The Guinea-Pig Is Not a Rodent, Nature, 1996, vol. 381, no. 6583, pp. 597–600.
Murphy, W.J., Eizirik, E., O’Brien, S.J., et al., Resolution of the Early Placental Mammal Radiation Using Bayesian Phylogenetics, Science, 2001, vol. 294, no. 5550, pp. 2348–2351.
Shimamura, M., Yasue, H., Ohshima, K., et al., Molecular Evidence from Retroposons that Whales Form a Clade within Even-Toed Ungulates, Nature, 1997, vol. 388, no. 6643, pp. 666–670.
Stoneking, M., Fontius, J.J., Clifford, S.L., et al., Alu Insertion Polymorphisms and Human Evolution: Evidence for a Larger Population Size in Africa, Genome Res., 1997, vol. 7, no. 11, pp. 1061–1071.
Watkins, W.S., Rogers, A.R., Ostler, C.T., et al., Genetic Variation Among World Populations: Inferences from 100 Alu Insertion Polymorphisms, Genome Res., 2003, vol. 13, no. 7, pp. 1607–1618.
Nikaido, M., Nishihara, H., Hukumoto, Y., et al., Ancient SINEs from African Endemic Mammals, Mol. Biol. Evol., 2003, vol. 20, no. 4, pp. 522–527.
Rothenburg, S., Eiben, M., Koch-Nolte, E., et al., Independent Integration of Rodent Identifier (ID) Elements into Orthologous Sites of Some RT6 Alleles of Rattus norvegicus and Rattus rattus, J. Mol. Evol., 2002, vol. 55, no. 3, pp. 251–259.
Van De Lagemaat, L.N., Gagnier, L., Medstrand, P., et al., Genomic Deletions and Precise Removal of Transposable Elements Mediated by Short Identical DNA Segments in Primates, Genome Res., 2005, vol. 15, no. 9, pp. 1243–1249.
Terai, Y., Takahashi, K., Nishida, M., et al., Using SINEs to Probe Ancient Explosive Speciation: “Hidden” Radiation of African Cichlids?, Mol. Biol. Evol., 2003, vol. 20, no. 6, pp. 924–930.
Author information
Authors and Affiliations
Additional information
Original Russian Text © N.A. Veniaminova, N.S. Vassetzky, L.A. Lavrenchenko, S.V. Popov, D.A. Kramerov, 2007, published in Genetika, 2007, Vol. 43, No. 7, pp. 916–929.
Rights and permissions
About this article
Cite this article
Veniaminova, N.A., Vassetzky, N.S., Lavrenchenko, L.A. et al. Phylogeny of the order rodentia inferred from structural analysis of short retroposon B1. Russ J Genet 43, 757–768 (2007). https://doi.org/10.1134/S1022795407070071
Received:
Issue Date:
DOI: https://doi.org/10.1134/S1022795407070071