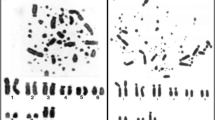
Abstract
Comparative chromosome painting was applied to the Indian spiny mouse (Mus platythrix) with mouse (M. musculus) chromosome-specific probes for understanding the process of chromosome rearrangements between the two species. The chromosome locations of the 5S and 18S-28S ribosomal RNA genes and the order of the 119 and Tcp-1 genes in the In(17)2 region of the t-complex were also compared. All the painting probes were successfully hybridized to the Indian spiny mouse chromosomes, and a total of 27 segments homologous to mouse chromosomes were identified. The comparative FISH analysis revealed that tandem fusions were major events in the chromosome evolution of the Indian spiny mouse. In addition, other types of chromosome rearrangements, i.e. reciprocal translocations and insertions, were also included.
Similar content being viewed by others
References
Bonhomme F, Guénet J-L (1996) The laboratory mouse and its wild relatives. In: Lyon MF, Rastan S, Brown SDM, eds. Genetic Variants and Strains of the Laboratory Mouse. Oxford, UK: Oxford University Press, pp 1577-1596.
Dev VG, Tantravahi R, Miller DA, Miller OJ (1977) Nucleolus organizers in Mus musculus subspecies and in the RAG mouse cell line. Genetics 86: 389-398.
Ellerman JR (1961) Mammalia Vol. 3 Rodentia. In: Roonwal ML, ed. The Fauna of India Including Pakistan, Burma and Ceylon. Calcutta,India: Baptist Mission Press, pp 771-782.
Hammer MF, Schimenti J, Silver LM (1989) Evolution of mouse chromosome 17 and the origin of inversions associated with t haplotypes. Proc Natl Acad Sci USA 86: 3261-3265.
Herrmann B, Bucan M, Mains PE, Frischauf A-M, Silver LM, Lehrach H (1986) Genetic analysis of the proximal portion of the mouse t complex: evidence for a second inversion within t haplotypes. Cell 44: 469-476.
Kominami R, Mishima Y, Urano Y, Sakai M, Muramatsu M (1982) Cloning and determination of the transcription termination site of ribosomal RNA gene of the mouse. Nucl Acids Res 10: 1963-1979.
Kuroiwa A, Tsuchiya K, Watanabe T et al. (2001) Conservation of the rat X chromosome gene order in rodent species. Chromosome Res 9: 61-67.
Matsuda Y, Chapman VM (1995) Application of fluorescence in situ hybridization in genome analysis of the mouse. Electrophoresis 16: 261-272.
Matsuda Y, Harada Y-N, Natsuume-Sakai S, Lee K, Shiomi T, Chapman VM (1992) Location of the mouse complement factor H gene (cfh) by FISH analysis and replication R-banding. Cytogenet Cell Genet 61: 282-285.
Matsuda Y, Moriwaki K, Chapman VM, Hoi-Sen Y, Akbarzadeh J, Suzuki H (1994) Chromosomal mapping of mouse 5S rRNA genes by direct R-banding fluorescence in situ hybridization. Cytogenet Cell Genet 66: 246-249.
Scherthan H, Cremer T, Arnason U, Weier H-U, Lima-de-Faria A, Frönicke L (1994) Comparative chromosome painting discloses homologous segments in distantly related mammals. Nature Genet 6: 342-347.
Schubert S, Dechend F, Skawran B, Krawczak M, Schmidtke J (2000) Molecular evolution of the murine tspy genes. Cytogenet Cell Genet 91: 239-242.
Seabright M (1971) A rapid banding technique for human chromosomes. Lancet 7731: 971-972.
Stanyon R, Yang F, Cavagna P et al. (1999) Reciprocal chromosome painting shows that genomic rearrangement between rat and mouse proceeds ten times faster than between humans and cats. Cytogenet Cell Genet 84: 150-155.
Suzuki H, Kurihara Y, Kanehisa T, Moriwaki K (1990) Variation in the distribution of silver-staining nucleolar organizer regions on the chromosomes of the wild mouse, Mus musculus. Mol Biol Evol 7: 271-282.
Tsuchiya K, Yosida TH (1972) Indian brown spiny mouse, Mus platythrix, having the lowest fundamental chromosome number. Ann Rep Natl Inst Genet Jpn 22: 51-52.
Wienberg J, Jauch A, Stanyon R, Cremer T (1990) Molecular cytotaxonomy of primates by chromosomal in situ suppression hybridization. Genomics 8: 347-350.
Wienberg J, Stanyon R (1995) Chromosome painting in mammals as an approach to comparative genomics. Curr Opin Genet Dev 5: 792-797.
Willison KR, Dudley K, Potter J (1986) Molecular cloning and sequence analysis of a haploid expressed gene encoding t complex polypeptide 1. Cell 44: 727-738.
Yang F, O'Brien PCM, Ferguson-Smith MA (2000) Comparative chromosome map of the laboratory mouse and Chinese hamster defined by reciprocal chromosome painting. Chromosome Res 8: 219-227.
Yosida TH (1979) Possible evidence for the karyotype evolution of the Indian spiny mouse due to tandem fusion of the house mouse chromosomes. Proc Jpn Acad 55: 270-274.
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Matsubara, K., Nishida-Umehara, C., Kuroiwa, A. et al. Identification of chromosome rearrangements between the laboratory mouse (Mus musculus) and the Indian spiny mouse (Mus platythrix) by comparative FISH analysis. Chromosome Res 11, 57–64 (2003). https://doi.org/10.1023/A:1022010116287
Issue Date:
DOI: https://doi.org/10.1023/A:1022010116287