Abstract
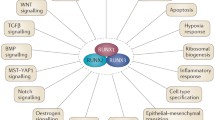
Over the past several years it has become increasingly evident that normal development and cancer share many properties. Both processes involve alterations in cell proliferation and differentiation, cell death, neovascularization, and cell motility and invasion. Thus, genes involved in normal development are frequently utilized in neoplasia. During development, numerous transcriptional regulatory mechanisms are used to ensure tight control over cellular proliferation. In this review we focus on a number of transcription factor families (homeobox, STAT, and Ets), and on inhibitors of transcription factors (Id), which have been implicated in controlling the cell cycle not only in normal mammary gland development but also in breast tumorigenesis.
Similar content being viewed by others
REFERENCES
M. T. Lewis (2000). Homeobox genes in mammary gland development and neoplasia. Breast Cancer Res. 2:158–169.
M. A. Dyer, F. J. Livesey, C. L. Cepko, and G. Oliver (2003). Prox1 function controls progenitor cell proliferation and horizontal cell genesis in the mammalian retina. Nat. Genet. 34:53–58.
C. Abate-Shen (2002). Deregulated homeobox gene expression in cancer: Cause or consequence? Nat. Rev. Cancer 2:777–785.
J. M. Bjornsson, N. Larsson, A. C. Brun, M. Magnusson, E. Andersson, P. Lundstrom, J. Larsson, E. Repetowska, M. Ehinger, R. K. Humphries, and S. Karlsson (2003). Reduced proliferative capacity of hematopoietic stem cells deficient in Hoxb3 and Hoxb4. Mol. Cell Biol. 23:3872–3883.
C. Kioussi, P. Briata, S. H. Baek, D. W. Rose, N. S. Hamblet, T. Herman, K. A. Ohgi, C. Lin, A. Gleiberman, J. Wang, V. Brault, P. Ruiz-Lozano, H. D. Nguyen, R. Kemler, C. K. Glass, A. Wynshaw-Boris, and M. G. Rosenfeld (2002). Identification of a Wnt/Dvl/beta-Catenin ≥ Pitx2 pathway mediating cell-type-specific proliferation during development. Cell 111:673–685.
V. Moucadel, M. S. Totaro, C. D. Dell, P. Soubeyran, J. C. Dagorn, J. N. Freund, and J. L. Iovanna (2002). The homeobox gene Cdx1 belongs to the p53-p21(WAF)-Bcl-2 network in intestinal epithelial cells. Biochem. Biophys. Res. Commun. 297:607–615.
A. W. Ledford, J. G. Brantley, G. Kemeny, T. L. Foreman, S. E. Quaggin, P. Igarashi, S. M. Oberhaus, M. Rodova, J. P. Calvet, and G. B. Vanden Heuvel (2002). Deregulated expression of the homeobox gene Cux-1 in transgenic mice results in downregulation of p27(kip1) expression during nephrogenesis, glomerular abnormalities, and multiorgan hyperplasia. Dev. Biol. 245:157–171.
H. L. Ford (1998). Homeobox genes: A link between development, cell cycle, and cancer? Cell Biol. Int. 22:397–400.
R. Tupler, G. Perini, and M. R. Green (2001). Expressing the human genome. Nature 409:832–833.
F. Apiou, D. Flagiello, C. Cillo, B. Malfoy, M. F. Poupon, and B. Dutrillaux (1996). Fine mapping of human HOX gene clusters. Cytogenet. Cell Genet. 73:114–115.
K. Kawakami, S. Sato, H. Ozaki, and K. Ikeda (2000). Six family genes—Structure and function as transcription factors and their roles in development. Bioessays 22:616–626.
V. C. Bromleigh and L. P. Freedman (2000). p21 is a transcriptional target of HOXA10 in differentiating myelomonocytic cells. Genes Dev. 14:2581–2586.
K. Sawada, Y. Konishi, M. Tominaga, Y. Watanabe, J. Hirano, S. Inoue, R. Kageyama, M. Blum, and A. Tominaga (2000). Goosecoid suppresses cell growth and enhances neuronal differentiation of PC12 cells. J. Cell Sci. 113:2705–2713.
M. J. B.-G. R. Kim, W. A. Banach-Petrosky, N. Desai, Y. Wang, S. W. Hayward, G. R. Cunha, R. D. Cardiff, M. M. Shen, and C. Abate-Shen (2002). Nkx3.1 mutant mice recapitulate early stages of prostate carcinogenesis. Cancer Res. 62:2999–3004.
R. C. Smith, D. Branellec, D. H. Gorski, K. Guo, H. Perlman, J. F. Dedieu, C. Pastore, A. Mahfoudi, P. Denefle, J. M. Isner, and K. Walsh (1997). p21CIP1-mediated inhibition of cell proliferation by overexpression of the gax homeodomain gene. Genes. Dev. 11:1674–1689.
T. Kawabe, A. J. Muslin, and S. J. Korsmeyer (1997). HOX11 interacts with protein phosphatases PP2A and PP1 and disrupts a G2/M cell-cycle checkpoint. Nature 385:454–458.
D. L. Rossi, A. Acebron, and P. Santisteban (1995). Function of the homeo and paired domain proteins TTF-1 and Pax-8 in thyroid cell proliferation. J. Biol. Chem. 270:23139–23142.
M. Burmeister, J. Novak, M. Y. Liang, S. Basu, L. Ploder, N. L. Hawes, D. Vidgen, F. Hoover, D. Goldman, V. I. Kalnins, T. H. Roderick, B. A. Taylor, M. H. Hankin, and R. R. McInnes (1996). Ocular retardation mouse caused by Chx10 homeobox null allele: Impaired retinal progenitor proliferation and bipolar cell differentiation. Nat. Genet. 12:376–384.
J. Krosl, S. Baban, G. Krosl, S. Rozenfeld, C, and G. Sauvageau (1998). Cellular proliferation and transformation induced by HOXB4 and HOXB3 proteins involves cooperation with PBX1. Oncogene 16:3403–3412.
Y. H. Liu, Z. Tang, R. K. Kundu, L. Wu, W. Luo, D. Zhu, F. Sangiorgi, M. L. Snead, and R. E. Maxson (1999). Msx2 gene dosage influences the number of proliferative osteogenic cells in growth centers of the developing murine skull: A possible mechanism for MSX2-mediated craniosynostosis in humans. Dev. Biol. 205:260–274.
M. Kobayashi, R. Toyama, H. Takeda, I. B. Dawid, and K. Kawakami (1998). Overexpression of the forebrain-specific homeobox gene six3 induces rostral forebrain enlargement in zebrafish. Development 125:2973–2982.
G. Goudreau, P. Petrou, L. W. Reneker, J. Graw, J. Loster, and P. Gruss (2002). Mutually regulated expression of Pax6 and Six3 and its implications for the Pax6 haploinsufficient lens phenotype. Proc. Natl. Acad. Sci. U.S.A. 99:8719–8724.
H. L. Ford, E. N. Kabingu, E. A. Bump, G. L. Mutter, and A. B. Pardee (1998). Abrogation of the G2 cell cycle checkpoint associated with overexpression of HSIX1: A possible mechanism of breast carcinogenesis. Proc. Natl. Acad. Sci. U.S.A. 95:12608–12613.
H. L. Ford, and A. B. Pardee (1999). Cancer and the cell cycle. J. Cell Biochem. (Suppl.) 32/33:166–172.
W. Zheng, L. Huang, Z. B. Wei, D. Silvius, B. Tang, and P. X. Xu (2003). The role of Six1 in mammalian auditory system development. Development 130:3989–4000.
H. Ozaki, K. Nakamura, J. I. Funahashi, K. Ikeda, G. Yamada, H. Tokano, H. O. Okamura, K. Kitamura, S. Muto, H. Kotaki, K. Sudo, R. Horai, Y. Iwakura, and K. Kawakami (2004). Six1 controls patterning of the mouse otic vesicle. Development 131:551–562.
X. Li, V. Perissi, F. Liu, D. W. Rose, and M. G. Rosenfeld (2002). Tissue-specific regulation of retinal and pituitary precursor cell proliferation. Science 297:1180–1183.
X. Li, K. A. Oghi, J. Zhang, A. Krones, K. T. Bush, C. K. Glass, S. K. Nigam, A. K. Aggarwal, R. Maas, D. W. Rose, and M. G. Rosenfeld (2003). Eya protein phosphatase activity regulates Six1-Dach-Eya transcriptional effects in mammalian organogenesis. Nature 426:247–254.
J. Lynch, E. R. Suh, D. G. Silberg, S. Rulyak, N. Blanchard, and P. G. Traber (2000). The caudal-related homeodomain protein Cdx1 inhibits proliferation of intestinal epithelial cells by down-regulation of D-type cyclins. J. Biol. Chem. 275:4499–4506.
T. V. Petrova, T. Makinen, T. P. Makela, J. Saarela, I. Virtanen, R. E. Ferrell, D. N. Finegold, D. Kerjaschki, S. Yla-Herttuala, and K. Alitalo (2002). Lymphatic endothelial reprogramming of vascular endothelial cells by the Prox-1 homeobox transcription factor. EMBO. J. 21:4593–4599.
J. M. Bjornsson, E. Andersson, P. Lundstrom, N. Larsson, X. Xu, E. Repetowska, R. K. Humphries, and S. Karlsson (2001). Proliferation of primitive myeloid progenitors can be reversibly induced by HOXA10. Blood 98:3301–3308.
D. M. Wellik and M. R. Capecchi (2003). Hox10 and Hox11 genes are required to globally pattern the mammalian skeleton. Science 301:363–367.
H. Chen and S. Sukumar (2003). Role of homeobox genes in normal mammary gland development and breast Tumorigenesis. J. Mammary Gland Biol. Neoplasia 21:157–174.
F. Chen and M. R. Capecchi (1999). Paralogous mouse Hox genes, Hoxa9, Hoxb9, and Hoxd9, function together to control development of the mammary gland in response to pregnancy. Proc. Natl. Acad. Sci. U.S.A 96:541–546.
T. R. Golub, D. K. Slonim, P. Tamayo, C. Huard, M. Gaasenbeek, J. P. Mesirov, H. Coller, M. L. Loh, J. R. Downing, M. A. Caligiuri, C. D. Bloomfield, and E. S. Lander (1999). Molecular classification of cancer: Class discovery and class prediction by gene expression monitoring. Science 286:531–537.
Y. Friedmann, C. A. Daniel, P. Strickland, and C. W. Daniel (1994). Hox genes in normal and neoplastic mouse mammary gland. Cancer Res. 54:5981–5985.
A. Srebrow, Y. Friedmann, A. Ravanpay, C. W. Daniel, and M. J. Bissell (1998). Expression of Hoxa-1 and Hoxb-7 is regulated by extracellular matrix-dependent signals in mammary epithelial cells. J. Cell Biochem. 69:377–391.
A. Care, A. Silvani, E. Meccia, G. Mattia, A. Stoppacciaro, G. Parmiani, C. Peschle, and M. P. Colombo (1996). HOXB7 constitutively activates basic fibroblast growth factor in melanomas. Mol. Cell Biol. 16:4842–4851.
A. Care, A. Silvani, E. Meccia, G. Mattia, C. Peschle, and M. P. Colombo (1998). Transduction of the SkBr3 breast carcinoma cell line with the HOXB7 gene induces bFGF expression, increases cell proliferation and reduces growth factor dependence. Oncogene 16:3285–3289.
A. Care, M. Valtieri, G. Mattia, E. Meccia, B. Masella, L. Luchetti, F. Felicetti, M. P. Colombo, and C. Peschle (1999). Enforced expression of HOXB7 promotes hematopoietic stem cell proliferation and myeloid-restricted progenitor differentiation. Oncogene 18:1993–2001.
A. Care, F. Felicetti, E. Meccia, L. Bottero, M. Parenza, A. Stoppacciaro, C. Peschle, and M. P. Colombo (2001). HOXB7: A key factor for tumor-associated angiogenic switch. Cancer Res. 61:6532–6539.
E. Hyman, P. Kauraniemi, S. Hautaniemi, M. Wolf, S. Mousses, E. Rozenblum, M. Ringner, G. Sauter, O. Monni, A. Elkahloun, O. P. Kallioniemi, and A. Kallioniemi (2002). Impact of DNA amplification on gene expression patterns in breast cancer. Cancer Res. 62:6240–6245.
V. Raman, S. A. Martensen, D. Reisman, E. Evron, W. F. Odenwald, E. Jaffee, J. Marks, and S. Sukumar (2000). Compromised HOXA5 function can limit p53 expression in human breast tumours. Nature 405:974–978.
G. Oliver, R. Wehr, N. A. Jenkins, N. G. Copeland, B. N. Cheyette, V. Hartenstein, S. L. Zipursky, and P. Gruss (1995). Homeobox genes and connective tissue patterning. Development 121:693–705.
H. Ohto, S. Kamada, K. Tago, S. I. Tominaga, H. Ozaki, S. Sato, and K. Kawakami (1999). Cooperation of six and eya in activation of their target genes through nuclear translocation of Eya. Mol. Cell Biol. 19:6815–6824.
F. Relaix and M. Buckingham (1999). From insect eye to vertebrate muscle: Redeployment of a regulatory network. Genes Dev. 13:3171–3178.
C. Laclef, E. Souil, J. Demignon, and P. Maire (2003). Thymus, kidney and craniofacial abnormalities in Six1 deficient mice. Mech. Dev. 120:669–679.
A. P. Young, R. Nagarajan, and G. D. Longmore (2003). Mechanisms of transcriptional regulation by Rb-E2F segregate by biological pathway. Oncogene 22:7209–7217.
C. Laclef, G. Hamard, J. Demignon, E. Souil, C. Houbron, and P. Maire (2003). Altered myogenesis in Six1-deficient mice. Development 130:2239–2252.
K. J. Martin, E. Graner, Y. Li, L. M. Price, B. M. Kritzman, M. V. Fournier, E. Rhei, and A. B. Pardee (2001). High-sensitivity array analysis of gene expression for the early detection of disseminated breast tumor cells in peripheral blood. Proc. Natl. Acad. Sci. U.S.A. 98:2646–2651.
H. L. Ford, E. Landesman-Bollag, C. S. Dacwag, P. T. Stukenberg, A. B. Pardee, and D. C. Seldin (2000). Cell cycleregulated phosphorylation of the human SIX1 homeodomain protein. J. Biol. Chem. 275:22245–22254.
D. Davidson (1995). The function and evolution of Msx genes: Pointers and paradoxes. Trends Genet 11:405–411.
G. Hu, H. Lee, S. M. Price, M. M. Shen, and C. Abate-Shen (2001). Msx homeobox genes inhibit differentiation through upregulation of cyclin D1. Development 128:2373–2384.
I. Satokata, L. Ma, H. Ohshima, M. Bei, I. Woo, K. Nishizawa, T. Maeda, Y. Takano, M. Uchiyama, S. Heaney, H. Peters, Z. Tang, R. Maxson, and R. Maas (2000). Msx2 deficiency in mice causes pleiotropic defects in bone growth and ectodermal organ formation. Nat. Genet. 24:391–395.
T. C. Wang, R. D. Cardiff, L. Zukerberg, E. Lees, A. Arnold, and E. V. Schmidt (1994). Mammary hyperplasia and carcinoma in MMTV-cyclin D1 transgenic mice. Nature 369:669–671.
G. B. Silberstein, G. R. Dressler, and K. Van Horn (2002). Expression of the PAX2 oncogene in human breast cancer and its role in progesterone-dependent mammary growth. Oncogene 21:1009–1016.
A. Nepveu (2001). Role of the multifunctional CDP/Cut/Cux homeodomain transcription factor in regulating differentiation, cell growth and development. Gene 270:1–15.
J. Bromberg (2000). Signal transducers and activators of transcription as regulators of growth, apoptosis and breast development. Breast. Cancer Res. 2:86–90.
J. Bromberg (2002). Stat proteins and oncogenesis. J. Clin. Invest. 109:1139–1142.
C. J. Watson (2001). Stat transcription factors in mammary gland development and tumorigenesis. J. Mammary Gland Biol. Neoplasia 6:115–127.
X. Liu, G. W. Robinson, and L. Hennighausen (1996). Activation of Stat5a and Stat5b by tyrosine phosphorylation is tightly linked to mammary gland differentiation. Mol. Endocrinol 10:1496–1506.
E. Iavnilovitch, B. Groner, and I. Barash (2002). Overex-pression and forced activation of stat5 in mammary gland of transgenic mice promotes cellular proliferation, enhances differentiation, and delays postlactational apoptosis. Mol. Cancer Res. 1:32–47.
T. Bowman, R. Garcia, J. Turkson, and R. Jove (2000). STATs in oncogenesis. Oncogene 19:2474–2488.
P. L. Welcsh, M. K. Lee, R. M. Gonzalez-Hernandez, D. J. Black, M. Mahadevappa, E. M. Swisher, J. A. Warrington, and M. C. King (2002). BRCA1 transcriptionally regulates genes involved in breast tumorigenesis. Proc. Natl. Acad. Sci. U.S.A. 99:7560–7565.
A. Widschwendter, S. Tonko-Geymayer, T. Welte, G. Daxenbichler, C. Marth, and W. Doppler (2002). Prognostic significance of signal transducer and activator of transcription 1 activation in breast cancer. Clin. Cancer Res. 8:3065–3074.
P. J. Real, A. Sierra, A. De Juan, J. C. Segovia, J. M. Lopez-Vega, and J. L. Fernandez-Luna (2002). Resistance to chemotherapy via Stat3-dependent overexpression of Bcl-2 in metastatic breast cancer cells. Oncogene 21:7611–7618.
L. Li and P. E. Shaw (2002). Autocrine-mediated activation of STAT3 correlates with cell proliferation in breast carcinoma lines. J. Biol. Chem. 277:17397–17405.
R. Garcia, T. L. Bowman, G. Niu, H. Yu, S. Minton, C. A. Muro-Cacho, C. E. Cox, R. Falcone, R. Fairclough, S. Parsons, A. Laudano, A. Gazit, A. Levitzki, A. Kraker, and R. Jove (2001). Constitutive activation of Stat3 by the Src and JAK tyrosine kinases participates in growth regulation of human breast carcinoma cells. Oncogene 20:2499–2513.
F. Zhang, C. Li, H. Halfter, and J. Liu (2003). Delineating an oncostatin M-activated STAT3 signaling pathway that coordinates the expression of genes involved in cell cycle regulation and extracellular matrix deposition of MCF-7 cells. Oncogene 22:894–905.
S. Ren, H. R. Cai, M. Li, and P. A. Furth (2002). Loss of Stat5a delays mammary cancer progression in a mouse model. Oncogene 21:4335–4339.
H. Yamashita, H. Iwase, T. Toyama, and Y. Fujii (2003). Naturally occurring dominant-negative Stat5 suppresses transcriptional activity of estrogen receptors and induces apoptosis in T47D breast cancer cells. Oncogene 22:1638–1652.
T. Oikawa and T. Yamada (2003). Molecular biology of the Ets family of transcription factors. Gene 303:11–34.
A. D. Sharrocks (2001). The ETS-domain transcription factor family. Nat. Rev. Mol. Cell Biol. 2:827–237.
V. I. Sementchenko and D. K. Watson (2000). Ets target genes: past, present and future. Oncogene 19:6533–6548.
I. Matushansky, F. Radparvar, and A. I. Skoultchi (2003). CDK6 blocks differentiation: coupling cell proliferation to the block to differentiation in leukemic cells. Oncogene 22:4143–4149.
N. Ohtani, Z. Zebedee, T. J. Huot, J. A. Stinson, M. Sugimoto, Y. Ohashi, A. D. Sharrocks, G. Peters, and E. Hara (2001). Opposing effects of Ets and Id proteins on p16INK4a expression during cellular senescence. Nature 409:1067–1070.
C. Zhang, M. M. Kavurma, A. Lai, and L. M. Khachigian (2003). Ets-1 protects vascular smooth muscle cells from undergoing apoptosis by activating p21WAF1/Cip1: ETs-1 regulates basal and inducible p21WAF1/Cip1 transcription via distinct cis-acting elements in the p21WAF1/Cip1 promoter. J. Biol. Chem. 278:27903–27909.
Y. Miyazaki, P. Boccuni, S. Mao, J. Zhang, H. Erdjument-Bromage, P. Tempst, H. Kiyokawa, and S. D. Nimer (2001). Cyclin A-dependent phosphorylation of the ETS-related protein, MEF, restricts its activity to the G1 phase of the cell cycle. J. Biol. Chem. 276:40528–40536.
D. N. Sgouras, M. A. Athanasiou, G. J. Beal, Jr., R. J. Fisher, D. G. Blair, and G. J. Mavrothalassitis (1995). ERF: An ETS domain protein with strong transcriptional repressor activity, can suppress ets-associated tumorigenesis and is regulated by phosphorylation during cell cycle and mitogenic stimulation. EMBO J. 14:4781–4793.
L. Le Gallic, D. Sgouras, G. Beal, Jr., and G. Mavrothalassitis (1999). Transcriptional repressor ERF is a Ras/mitogen-activated protein kinase target that regulates cellular proliferation. Mol. Cell Biol. 19:4121–4133.
A. Chotteau-Lelievre, R. Montesano, J. Soriano, P. Soulie, X. Desbiens, and Y. de Launoit (2003). PEA3 transcription factors are expressed in tissues undergoing branching morphogenesis and promote formation of duct-like structures by mammary epithelial cells in vitro. Dev. Biol. 259:241–257.
I. G. Maroulakou and D. B. Bowe (2000). Expression and function of Ets transcription factors in mammalian development: A regulatory network. Oncogene 19:6432–6442.
K. Kas, E. Finger, F. Grall, X. Gu, Y. Akbarali, J. Boltax, A. Weiss, P. Oettgen, R. Kapeller, and T. A. Libermann (2000). ESE-3, a novel member of an epithelium-specific ets transcription factor subfamily, demonstrates different target gene specificity from ESE-1. J. Biol. Chem. 275:2986–2998.
R. Neve, C. H. Chang, G. K. Scott, A. Wong, R. R. Friis, N. E. Hynes, and C. C. Benz (1998). The epithelium-specific ets transcription factor ESX is associated with mammary gland development and involution. FASEB. J. 12:1541–1550.
J. Zhou, A. Y. Ng, M. J. Tymms, L. S. Jermiin, A. K. Seth, R. S. Thomas, and I. Kola (1998). A novel transcription factor, ELF5, belongs to the ELF subfamily of ETS genes and maps to human chromosome 11p13-15, a region subject to LOH and rearrangement in human carcinoma cell lines. Oncogene 17:2719–2732.
N. A. Kurpios, N. A. Sabolic, T. G. Shepherd, G. M. Fidalgo, and J. A. Hassell (2003). Function of PEA3 Ets transcription factors in mammary gland development and oncogenesis. J. Mammary Gland Biol. Neoplasia 8:177–190.
T. Shepherd and J. A. Hassell (2001). Role of Ets transcription factors in mammary gland development and oncogenesis. J. Mammary Gland Biol. Neoplasia 6:129–140.
P.N. Span, P. Manders, J. J. Heuvel, C. M. Thomas, R. R. Bosch, L. V. Beex, and C. G. Sweep (2002). Expression of the transcription factor Ets-1 is an independent prognostic marker for relapse-free survival in breast cancer. Oncogene 21:8506–8509.
A. Ghadersohi, and A. K. Sood (2001). Prostate epithelium-derived Ets transcription factor mRNA is overexpressed in human breast tumors and is a candidate breast tumor marker and a breast tumor antigen. Clin. Cancer Res. 7:2731–2738.
M. Mitas, K. Mikhitarian, L. Hoover, M. A. Lockett, L. Kelley, A. Hill, W. E. Gillanders, and D. J. Cole (2002). Prostate-Specific Ets (PSE) factor: A novel marker for detection of metastatic breast cancer in axillary lymph nodes. Br. J. Cancer 86:899–904.
D. Scheurle, M. P. DeYoung, D. M. Binninger, H. Page, M. Jahanzeb, and R. Narayanan (2000). Cancer gene discovery using digital differential display. Cancer Res. 60:4037–4043.
R. J. Feldman, V. I. Sementchenko, M. Gayed, M. M. Fraig, and D. K. Watson (2003). Pdef expression in human breast cancer is correlated with invasive potential and altered gene expression. Cancer Res. 63:4626–4631.
A. Seth and T. S. Papas (1990). The c-ets-1 proto-oncogene has oncogenic activity and is positively autoregulated. Oncogene 5:1761–1767.
A. Seth, D. K. Watson, D. G. Blair, and T. S. Papas (1989). c-ets-2 Protooncogene has mitogenic and oncogenic activity. Proc. Natl. Acad. Sci. U.S.A. 86:7833–7837.
R. Pereira, C. T. Quang, I. Lesault, H. Dolznig, H. Beug, and J. Ghysdael (1999). FLI-1 inhibits differentiation and induces proliferation of primary erythroblasts. Oncogene 18:1597–1608.
A. H. Hart, C. M. Corrick, M. J. Tymms, P. J. Hertzog, and I. Kola (1995). Human ERG is a proto-oncogene with mitogenic and transforming activity. Oncogene 10:1423–1430.
P. J. Schedin, K. L. Eckel, S. M. McDaniel, J. D. Prescott, K. S. Brodsky, J. J. Tentler, and A. Gutierrez-Hartmann (2004). ESX induces transformation and functional epithelial to mesenchymal transition in MCF-12A mammary epithelial cells. Oncogene 23:1776–1779.
E. Sapi, M. B. Flick, S. Rodov, and B. M. Kacinski (1998). Ets-2 transdominant mutant abolishes anchorage-independent growth and macrophage colony-stimulating factor-stimulated invasion by BT20 breast carcinoma cells. Cancer Res. 58:1027–1033.
A. Delannoy-Courdent, V. Mattot, V. Fafeur, W. Fauquette, I. Pollet, T. Calmels, C. Vercamer, B. Boilly, B. Vandenbunder, and X. Desbiens (1998). The expression of an Ets1 transcription factor lacking its activation domain decreases uPA proteolytic activity and cell motility, and impairs normal tubulogenesis and cancerous scattering in mammary epithelial cells. J. Cell Sci. 111(Pt. 11):1521–1534.
N. Neznanov, A. K. Man, H. Yamamoto, C. A. Hauser, R. D. Cardiff, and R. G. Oshima (1999). A single targeted Ets2 allele restricts development of mammary tumors in transgenic mice. Cancer Res. 59:4242–4246.
J. P. Coppe, A. P. Smith, and P. Y. Desprez (2003). Id proteins in epithelial cells. Exp. Cell Res. 285:131–145.
H. A. Sikder, M. K. Devlin, S. Dunlap, B. Ryu, and R. M. Alani (2003). Id proteins in cell growth and tumorigenesis. Cancer Cell. 3:525–530.
A. Lasorella, T. Uo, and A. Iavarone (2001). Id proteins at the cross-road of development and cancer. Oncogene 20:8326–8333.
E. Hara, M. Hall, and G. Peters (1997). Cdk2-dependent phosphorylation of Id2 modulates activity of E2A-related transcription factors. EMBO J. 16:332–342.
R. W. Deed, E. Hara, G. T. Atherton, G. Peters, and J. D. Norton (1997). Regulation of Id3 cell cycle function by Cdk-2-dependent phosphorylation. Mol. Cell Biol. 17:6815–6821.
S. Parrinello, C. Q. Lin, K. Murata, Y. Itahana, J. Singh, A. Krtolica, J. Campisi, and P. Y. Desprez (2001). Id-1, ITF-2, and Id-2 comprise a network of helix-loop-helix proteins that regulate mammary epithelial cell proliferation, differentiation, and apoptosis. J. Biol. Chem. 276:39213–39219.
C. Q. Lin, S. Parrinello, J. Campisi, and P. Y. Desprez (1999). Regulation of mammary epithelial cell phenotypes by the helix-loop-helix protein, Id-1. Endocr. Relat. Cancer 6:49–50.
P. Y. Desprez, E. Hara, M. J. Bissell, J. Campisi (1995). Suppression of mammary epithelial cell differentiation by the helix-loop-helix protein Id-1. Mol. Cell Biol. 15:3398–3404.
S. Mori, S. I. Nishikawa, and Y. Yokota (2000). Lactation defect in mice lacking the helix-loop-helix inhibitor Id2. EMBO J. 19:5772–5781.
J. D. Norton, and G. T. Atherton (1998). Coupling of cell growth control and apoptosis functions of Id proteins. Mol. Cell Biol. 18:2371–2381.
R. M. Alani, J. Hasskarl, M. Grace, M. C. Hernandez, M. A. Israel, and K. Munger (1999). Immortalization of primary human keratinocytes by the helix-loop-helix protein, Id-1. Proc. Natl. Acad. Sci. U.S.A. 96:9637–9641.
A. Lasorella, R. Boldrini, C. Dominici, A. Donfrancesco, Y. Yokota, A. Inserra, and A. Iavarone (2002). Id2 is critical for cellular proliferation and is the oncogenic effector of N-myc in human neuroblastoma. Cancer Res. 62:301–306.
C. Beger, L. N. Pierce, M. Kruger, E. G. Marcusson, J. M. Robbins, P. Welcsh, P. J. Welch, K. Welte, M. C. King, J. R. Barber, and F. Wong-Staal (2001). Identification of Id4 as a regulator of BRCA1 expression by using a ribozyme-library-based inverse genomics approach. Proc. Natl. Acad. Sci. U.S.A. 98:130–135.
A. Swarbrick, J. Hunter, S. Lee, M. Sergio, R. L. Sutherlan, and E. A. Musgrove (2002). Id1 is a critical target of c-myc in breast cancer cells. In Molecular Biology of the Cell, 42nd American Society for Cell Biology Annual Meeting, Abstract 2438. 2002.
C. Q. Lin, J. Singh, K. Murata, Y. Itahana, S. Parrinello, S. H. Liang, C. E. Gillett, J. Campisi, and P. Y. Desprez (2000). A role for Id-1 in the aggressive phenotype and steroid hormone response of human breast cancer cells. Cancer Res. 60:1332–1340.
S. F. Schoppmann, M. Schindl, G. Bayer, K. Aumayr, J. Dienes, R. Horvat, M. Rudas, M. Gnant, R. Jakesz, and P. Birner (2003). Overexpression of Id-1 is associated with poor clinical outcome in node negative breast cancer. Int. J. Cancer 104:677–682.
P. Y. Desprez, T. Sumida, and J. P. Coppe (2003). helix-loop-helix proteins in mammary gland development and breast cancer. J. Mammary Gland Biol. Neoplasia 8:225–239.
R. D. Coletta, K. Christensen, K. J. Reichenberger, J. Lamb, D. Micomonaco, L. Huang, D. M. Wolf, C. Müller-Tidow, T. R. Golub, K. Kawakami, and H. L. Ford. The Six1 homeoprotein stimulates tumorigenesis via reactivation of cyclin A1. Proc. Natl. Acad. Sci. U. S. A., in press.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Coletta, R.D., Jedlicka, P., Gutierrez-Hartmann, A. et al. Transcriptional Control of the Cell Cycle in Mammary Gland Development and Tumorigenesis. J Mammary Gland Biol Neoplasia 9, 39–53 (2004). https://doi.org/10.1023/B:JOMG.0000023587.40966.f6
Issue Date:
DOI: https://doi.org/10.1023/B:JOMG.0000023587.40966.f6