Abstract
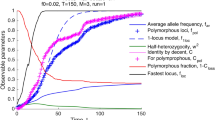
In a simple computer model of population evolution, we have shown that frequency of recombination between haplotypes during the gamete production influences the effectiveness of the reproduction strategy. High recombination rates keeps the fraction of defective alleles low while low recombination rate or uneven distributed recombination spots change the strategy of genomes' evolution and result in the accumulation of heterozygous loci in the genomes. Even short fragment of chromosome with restricted recombination influences the genetic structure of neighboring regions.
Similar content being viewed by others
References
Ayala, F.J., Kiger, J.A., 1980. Modern Genetics. The Benjamin/Cumings Pub. Comp. Inc., California.
Azbel, M., 1999. Phenomenological theory of mortality evolution: its singularities, universality, and superuniversality. Proc. Natl. Acad. Sci. USA 96, 3303–3307.
Cebrat, S., P(cekalski, A., Scharf, F., 2006. Monte Carlo simulations of the inside intron recombination. Int. J. Mod. Phys. C 17, 305–315.
Coe, J.B., Mao, Y., 2005. Gompertz mortality law and scaling behavior of the Penna model. Phys. Rev. E 72, 051925.
Dudkiewicz, M., Mackiewicz, P., Nowicka, A., Kowalczuk, M., Mackiewicz, D., Polak, N., Smolarczyk, K., Banaszak, J., Dudek, M.R., Cebrat, S., 2005. Correspondence between mutation and selection pressure and the genetic code degeneracy in the gene evolution. Future Generat. Comput. Systems 21 (7), 1033–1139.
Garver-Apgar, C.E., Gangestad, S.W., Thorhill, R., Miller, R.D., Olp, J.J., 2006. Major histocompatibility complex alleles, sexual responsivity, and unfaithfulness in romantic couples. Psych. Sci. 17, 830–835.
Gorlov, I.P., Gorlova, O.Y., 2001. Cost-benefit analysis of Recombination and its application for understanding of chiasma interference. J. Theor. Biol. 12, 12–45.
Hedrick, P.W., Black, F.L., 1997. Random mating selection within families against homozygotes for HLA in South Ameridians. Hereditas 127, 51–58.
Kliman, R.M.K., Rey, J., 1993. Reduced natural selection associated with low recombination in Drosophila melanogaster. Mol. Biol. Evol. 10, 1239–1258.
Laszkiewicz, A., Szymczak, Sz., Cebrat, S., 2003. Speciation effect in the Penna aging model. Int. J. Mod. Phys. C 14, 765–774.
Milinski, M., 1994. Hybridogenetic frogs on an evolutionary dead end raw. Trends Ecol. Evol. 9, 62.
de Oliveira, P.M.C., 2006. Chromosome length scaling in haploid asexual reproduction. J. Phys.: Condens. Matter 18, 1–9.
Penn, D.J., Potts, W.K., 1999. The evolution of mating preferences and major histocompatibility complex genes. Am. Naturalist 153, 145–164.
Stauffer, D., Cebrat, S., 2006. Extinction in genetic bit-string model with sexual recombination. Adv. Compl. Syst. 9, 147–156.
Yu, A., et al., 2001. Nature Comparison of human genetic and sequence-based physical maps. 409, 951–953.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Zawierta, M., Biecek, P., Waga, W. et al. The role of intragenomic recombination rate in the evolution of population's genetic pool. Theory Biosci. 125, 123–132 (2007). https://doi.org/10.1016/j.thbio.2007.02.002
Issue Date:
DOI: https://doi.org/10.1016/j.thbio.2007.02.002