Abstract
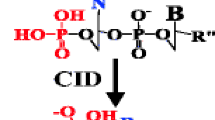
The dissociation of model RNA anions has been studied as a function of anion charge state and excitation amplitude using ion trap collisional activation. Similar to DNA anions, the precursor ion charge state of an RNA anion plays an important role in directing the preferred dissociation channels. Generally, the complementary c/y-ions from 5′ P-O bond cleavage dominate at low to intermediate charge states, while other backbone cleavages appear to a limited extent but increase in number and relative abundance at higher excitation energies. The competition between base loss, either as a neutral or as an anion, as well as the preference for the identity of the lost base are also observed to be charge-state dependent. To gain further insight into the partitioning of the dissociation products among the various possible channels, model dinucleotide anions have been subjected to a systematic study. In comparison to DNA, the 2′-OH group on RNA significantly facilitates the dissociation of the 5′ P-O bond. However, the degree of excitation required for a 5′ base loss and the subsequent 3′ C-O bond cleavage are similar for the analogous RNA and DNA dinucleotides. Data collected for protonated dinucleotides, however, suggest that the 2′-OH group in RNA can stabilize the glycosidic bond of a protonated base. Therefore, base loss from low charge state oligonucleotide anions, in which protonation of one or more bases via intramolecular proton transfer can occur, may also be stabilized in RNA anions relative to corresponding DNA anions.
Article PDF
Similar content being viewed by others
References
Mattick, J. S. Noncoding RNAs, the Architects of Eukaryotic Complexity. EMBO Rep. 2001, 2, 986–991.
Mattick, J. S. The Functional Genomics of Noncoding RNA. Science 2005, 309, 1527–1528.
Carninci, P., et al. The Transcriptional Landscape of the Mammalian Genome, P.. Science 2005, 309, 1559–1563.
Aravin, A. A.; Hannon, G. J.; Brennecke, J. The Piwi-piRNA Pathway Provides an Adaptive Defense in the Transposon Arms Race. Science 2007, 318, 761–764.
Hahn, C. S.; Strauss, E. G.; Strauss, J. H. Dideoxy Sequencing of RNA Using Reverse Transcriptase. Methods Enzymol. 1989, 180, 121–130.
Cerny, R. L.; Tomer, K. B.; Gross, M. L.; Grotjahn, L. Fast Atom Bombardment Combined with Tandem Mass Spectrometry for the Study of Dinucleotides. Anal. Biochem. 1986, 156, 424–435.
Phillips, D. R.; McCloskey, J. A. A Comprehensive Study of the Low Energy Collision-induced Dissociation of Dinucleoside Monophosphates. Int. J. Mass Spectrom. Ion Processes 1993, 128, 61–82.
Wu, J.; McLuckey, S. A. Gas-Phase Fragmentation of Oligonucleotide Ions. Int. J. Mass Spectrom. 2004, 237, 197–241.
Kirpekar, F.; Krogh, T. N. RNA Fragmentation Studied in a Matrix-Assisted Laser Desorption/Ionization Tandem Quadrupole/Orthogonal Time-of-Flight Mass Spectrometer: Rapid Commun. Mass Spectrom. 2001, 15, 8–14.
Schürch, S.; Bernal-Mendez, E.; Leumann, C. J. Electrospray Tandem Mass Spectrometry of Mixed-Sequence RNA/DNA Oligonucleotides. J. Am. Soc. Mass Spectrom. 2002, 13, 936–945.
Tromp, J. M.; Schürch, S. Gas-Phase Dissociation of Oligoribonucleotides and their Analogues Studied by Electrospray Ionization Tandem Mass Spectrometry. J. Am. Soc. Mass Spectrom. 2005, 16, 1262–1268.
Andersen, T. E.; Kirpekar, F.; Haselmann, K. F. R. N. A. Fragmentation in MALDI Mass Spectrometry Studied by H/D-Exchange: Mechanisms of General Applicability to Nucleic Acids. J. Am. Soc. Mass Spectrom. 2006, 17, 1353–1368.
Ni, J.; Pomerantz, C.; Rozenski, J.; Zhang, Y.; McCloskey, J. A. Interpretation of Oligonucleotide Mass Spectra for Determination of Sequence Using Electrospray Ionization and Tandem Mass Spectrometry. Anal. Chem. 1996, 68(13), 1989–1999.
McCloskey, J. A.; Whitehill, A. B.; Rozenski, J.; Qiu, F.; Crain, P. F. New Techniques for the Rapid Characterization of Oligonucleotides by Mass Spectrometry. Nucleosides Nucleotides 1999, 18(6/7), 1549–1553.
Mengel-Jørgensen, J.; Kirpekar, F. Detection of Pseudouridine and Other Modifications in tRNA by Cyanoethylation and MALDI Mass Spectrometry. Nucleic Acids Res. 2002, 30, e135.
Emmerechts, G.; Barbé, S.; Herdewijin, P.; Anné, J.; Rozenski, J. Post-Transcriptional Modification Mapping in the Clostridium acetobutylicum 16S rRNA by Mass Spectrometry and Reverse Transcriptase Assays. Nucleic Acids. Res. 2007, 35, 3494–3503.
McLuckey, S. A.; Goeringer, D. E. Slow Heating Methods in Tandem Mass Spectrometry. J. Mass Spectrom. 1997, 32, 461–474.
Xia, Y.; Chrisman, P. A.; Erickson, D. E.; Liu, J.; Liang, X.; Londry, F. A.; Yang, M. J.; McLuckey, S. A. Implementation of Ion/Ion Reactions in a Quadrupole/Time-of-Flight Tandem Mass Spectrometer. Anal. Chem. 2006, 78(12), 4146–4154.
Rozenski, J. Nucleic Acids Masspec Toolbox (Mongo Oligo Mass Calculator ver. 2.06). http://medlib.med.utah.edu/masspec.
McLuckey, S. A.; Vaidyanathan, G.; Habibi-Goudarzi, S. Charged vs. Neutral Nucleobase Loss from Multiply Charged Oligonucleotide Anions. J. Mass Spectrom. 1995, 30, 1222–1229.
Pan, S.; Verhoeven, K.; Lee, J. K. Investigation of the Initial Fragmentation of the Oligodeoxynucleotides in a Quadrupole Ion Trap: Charge Level-Related Base Loss. J. Am. Soc. Mass Spectrom. 2005, 16, 1853–1865.
Rodgers, M. T.; Campbell, S.; Marzluff, E. M.; Beauchamp, J. L. Low-Energy Collision-Induced Dissociation of Deprotonated Dinucleotides: Determination of the Energetically Favored Dissociation Pathways and the Relative Acidities of the Nucleic Acid Bases. Int. J. Mass Spectrom. 1994, 137, 121–149.
Habibi-Ghoudarzi, S.; McLuckey, S. A. Ion Trap Collisional Activation of the Deprotonated Deoxymononucleoside and Deoxydinucleoside Monophosphates. J. Am. Soc. Mass Spectrom. 1995, 6, 102–113.
McLuckey, S. A.; Habibi-Goudarzi, S. Decompositions of Multiply-Charged Oligonucleotide Anions. J. Am. Chem. Soc. 1993, 115, 12085–12095.
McLuckey, S. A.; Van Berkel, G. J.; Glish, G. L. Mass Spectrometry/Mass Spectrometry of Small Multiply Charged Oligonucleotides. J. Am. Soc. Mass Spectrom. 1992, 3, 60–70.
Nordhoff, E.; Cramer, R.; Karas, M.; Hillenkamp, F.; Kirpekar, F.; Kristiansen, K.; Roepstorff, P. Ion Stability of Nucleic Acids in Infrared Matrix-Assisted Laser Desorption/Ionization Mass Spectrometry. Nucleic Acids Res. 1993, 21, 3347–3357.
Tang, W.; Zhu, L.; Smith, L. M. Controlling DNA Fragmentation in MALDI-MS by Chemical Modification. Anal. Chem. 1997, 69, 302–312.
Wang, Z.; Wan, K. X.; Ramanathan, R.; Taylor, J. S.; Gross, M. L. Structure and Fragmentation Mechanisms of Isomeric T-Rich Oligodeoxynucleotides: A Comparison of Four Tandem Mass Spectrometric Methods. J. Am. Soc. Mass Spectrom. 1998, 9, 683–691.
Wan, K. X.; Gross, J.; Hillenkamp, F.; Gross, M. L. Fragmentation Mechanisms of Oligodeoxynucleotides Studied by H/D Exchange and Electrospray Ionization Tandem Mass Spectrometry. J. Am. Soc. Mass Spectrom. 2001, 12, 193–205.
Wan, K. X.; Gross, M. L. Fragmentation Mechanisms of Oligodeoxynucleotides: Effects of Replacing Phosphates with Methylphosphonates and Thymines with Other Bases in T-Rich Sequences. J. Am. Soc. Mass Spectrom. 2001, 12, 580–589.
Author information
Authors and Affiliations
Corresponding author
Additional information
Published online August 19, 2008
Rights and permissions
About this article
Cite this article
Huang, Ty., Kharlamova, A., Liu, J. et al. Ion trap collision-induced dissociation of multiply deprotonated RNA: c/y-ions versus (a-B)/w-ions. J Am Soc Mass Spectrom 19, 1832–1840 (2008). https://doi.org/10.1016/j.jasms.2008.08.009
Received:
Revised:
Accepted:
Issue Date:
DOI: https://doi.org/10.1016/j.jasms.2008.08.009