Abstract
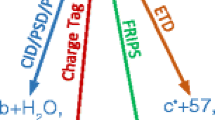
Nonenzymatic deamidation of asparagine residues in proteins generates aspartyl (Asp) and isoaspartyl (isoAsp) residues via a succinimide intermediate in a neutral or basic environment. Electron capture dissociation (ECD) can differentiate and quantify the relative abundance of these isomeric products in the deamidated proteins. This method requires the proteins to be digested, usually by trypsin, into peptides that are amenable to ECD. ECD of these peptides can produce diagnostic ions for each isomer; the c · + 58 and z − 57 fragment ions for the isoAsp residue and the fragment ion ((M + nH)(n−1)+· − 60) corresponding to the side-chain loss from the Asp residue. However, deamidation can also occur as an artifact during sample preparation, particularly when using typical tryptic digestion protocols. With 18O labeling, it is possible to differentiate deamidation occurring during trypsin digestion which causes a +3 Da (18O1 + 1D) mass shift from the pre-existing deamidation, which leads to a +1-Da mass shift. This paper demonstrates the use of 18O labeling to monitor three rapidly deamidating peptides released from proteins (calmodulin, ribonuclease A, and lysozyme) during the time course of trypsin digestion processes, and shows that the fast (∼4 h) trypsin digestion process generates no additional detectable peptide deamidations.
Article PDF
Similar content being viewed by others
References
Robinson, N. E.; Robinson, A. B. Molecular Clocks: Deamidation of Asparaginyl and Glutaminyl Residues in Peptides and Proteins. Althouse Press: Cave Junction, OR, 2004.
Robinson, N. E. Protein Deamidation. Proc. Natl. Acad. Sci. U. S. A.. 2002, 99, 5283–5288.
Robinson, N. E.; Robinson, A. B. Molecular Clocks. Proc. Natl. Acad. Sci. U. S. A.. 2001, 98, 944–949.
Lindner, H.; Sarg, B.; Hoertnagl, B.; Helliger, W. The Microheterogeneity of the Mammalian H1(0) Histone—Evidence for an Age-Dependent Deamidation. J. Biol. Chem. 1998, 273, 13324–13330.
Bischoff, R.; Kolbe, H. V. Deamidation of Asparagine and Glutamine Residues in Proteins and Peptides—Structural Determinants and Analytical Methodology. J. Chromatogr. B Biomed. Appl. 1994, 662, 261–278.
Lindner, H.; Helliger, W. Age-Dependent Deamidation of Asparagine Residues in Proteins. Exp. Gerontol. 2001, 36, 1551–1563.
Capasso, S. Estimation of the Deamidation Rate of Asparagine Side Chains. J. Pept. Res. 2000, 55, 224–229.
Geiger, T.; Clarke, S. Deamidation, Isomerization, and Racemization at Asparaginyl and Aspartyl Residues in Peptides—Succinimide-Linked Reactions That Contribute to Protein-Degradation. J. Biol. Chem. 1987, 262, 785–794.
Xie, M. L.; Schowen, R. L. Secondary Structure and Protein Deamidation. J. Pharm. Sci. 1999, 88, 8–13.
Robinson, N. E.; Robinson, A. B. Prediction of Protein Deamidation Rates from Primary and Three-Dimensional Structure. Proc. Natl. Acad. Sci. U. S. A. 2001, 98, 4367–4372.
Tylercross, R.; Schirch, V. Effects of Amino-Acid-Sequence, Buffers, and Ionic-Strength on the Rate and Mechanism of Deamidation of Asparagine Residues in Small Peptides. J. Biol. Chem. 1991, 266, 22549–22556.
Wright, H. T. Nonenzymatic Deamidation of Asparaginyl and Glutaminyl Residues in Proteins. Crit. Rev. Biochem. Mol. Biol. 1991, 26, 1–52.
Kossiakoff, A. A. Tertiary Structure Is a Principal Determinant to Protein Deamidation. Science. 1988, 240, 191–194.
Peters, B.; Trout, B. L. Asparagine Deamidation: pH-Dependent Mechanism from Density Functional Theory. Biochemistry. 2006, 45, 5384–5392.
Joshi, A. B.; Sawai, M.; Kearney, W. R.; Kirsch, L. E. Studies on the Mechanism of Aspartic Acid Cleavage and Glutamine Deamidation in the Acidic Degradation of Glucagon. J. Pharm. Sci. 2005, 94, 1912–1927.
Robinson, N. E.; Zabrouskov, V.; Zhang, J.; Lampi, K. J.; Robinson, A. B. Measurement of Deamidation of Intact Proteins by Isotopic Envelope and Mass Defect with Ion Cyclotron Resonance Fourier Transform Mass Spectrometry. Rapid Commun. Mass Spectrom. 2006, 20, 3535–3541.
Robinson, N. E.; Lampi, K. J.; Speir, J. P.; Kruppa, G.; Easterling, M.; Robinson, A. B. Quantitative Measurement of Young Human Eye Lens Crystallins by Direct Injection Fourier Transform Ion Cyclotron Resonance Mass Spectrometry. Mol. Vis. 2006, 12, 704–711.
Robinson, N. E.; Lampi, K. J.; McIver, R. T.; Williams, R. H.; Muster, W. C.; Kruppa, G.; Robinson, A. B. Quantitative Measurement of Deamidation in Lens Beta B2-Crystallin and Peptides by Direct Electrospray Injection and Fragmentation in a Fourier Transform Mass Spectrometer. Mol. Vis. 2005, 11, 1211–1219.
Schmid, D. G.; von der Mulbe, F.; Fleckenstein, B.; Weinschenk, T.; Jung, G. Broadband Detection Electrospray Ionization Fourier Transform Ion Cyclotron Resonance Mass Spectrometry to Reveal Enzymatically and Chemically Induced Deamidation Reactions within Peptides. Anal. Chem. 2001, 73, 6008–6013.
Chazin, W. J.; Kordel, J.; Thulin, E.; Hofmann, T.; Drakenberg, T.; Forsen, S. Identification of an Isoaspartyl Linkage Formed upon Deamidation of Bovine Calbindin-D9k and Structural Characterization by 2D 1H NMR. Biochemistry. 1989, 28, 8646–8653.
Carlson, A. D.; Riggin, R. M. Development of Improved High-Performance Liquid Chromatography Conditions for Nonisotopic Detection of Isoaspartic Acid to Determine the Extent of Protein Deamidation. Anal. Biochem. 2000, 278, 150–155.
Carr, S. A.; Hemling, M. E.; Bean, M. F.; Roberts, G. D. Integration of Mass-Spectrometry in Analytical Biotechnology. Anal. Chem. 1991, 63, 2802–2824.
Zhang, W.; Czupryn, M. J. Analysis of Isoaspartate in a Recombinant Monoclonal Antibody and Its Charge Isoforms. J. Pharm. Biomed. Anal. 2003, 30, 1479–1490.
Gonzalez, L. J.; Shimizu, T.; Satomi, Y.; Betancourt, L.; Besada, V.; Padron, G.; Orlando, R.; Shirasawa, T.; Shimonishi, Y.; Takao, T. Differentiating Alpha- and Beta-Aspartic Acids by Electrospray Ionization and Low-Energy Tandem Mass Spectrometry. Rapid Commun. Mass Spectrom. 2000, 14, 2092–2102.
Castet, S.; Enjalbal, C.; Fulcrand, P.; Guichou, J. F.; Martinez, J.; Aubagnac, J. L. Characterization of Aspartic Acid and Beta-Aspartic Acid in Peptides by Fast-Atom Bombardment Mass Spectrometry and Tandem Mass Spectrometry. Rapid Commun. Mass Spectrom. 1996, 10, 1934–1938.
Zubarev, R. A.; Kelleher, N. L.; McLafferty, F. W. Electron Capture Dissociation of Multiply Charged Protein Cations: A Nonergodic Process. J. Am. Chem. Soc. 1998, 120, 3265–3266.
Zubarev, R. A.; Haselmann, K. F.; Budnik, B.; Kjeldsen, F.; Jensen, F. Towards an Understanding of the Mechanism of Electron-Capture Dissociation: A Historical Perspective and Modern Ideas. Eur. J. Mass Spectrom. 2002, 8, 337–349.
Syrstad, E. A.; Turecek, F. Toward a General Mechanism of Electron Capture Dissociation. J. Am. Soc. Mass Spectrom. 2005, 16, 208–224.
Leymarie, N.; Costello, C. E.; O’Connor, P. B. Electron Capture Dissociation Initiates a Free Radical Reaction Cascade. J. Am. Chem. Soc. 2003, 125, 8949–8958.
Coon, J. J.; Syka, J. E. P.; Schwartz, J. C.; Shabanowitz, J.; Hunt, D. F. Anion Dependence in the Partitioning between Proton and Electron Transfer in Ion/Ion Reactions. Int. J. Mass Spectrom. 2004, 236, 33–42.
Syka, J. E. P.; Coon, J. J.; Schroeder, M. J.; Shabanowitz, J.; Hunt, D. F. Peptide and Protein Sequence Analysis by Electron Transfer Dissociation Mass Spectrometry. Proc. Natl. Acad. Sci. U. S. A. 2004, 101, 9528–9533.
Cournoyer, J. J.; Lin, C.; Bowman, M. J.; O’Connor, P. B. Quantitating the Relative Abundance of Isoaspartyl Residues in Deamidated Proteins by Electron Capture Dissociation. J. Am. Soc. Mass Spectrom. 2007, 18, 48–56.
Cournoyer, J. J.; Lin, C.; O’Connor, P. B. Detecting Deamidation Products in Proteins by Electron Capture Dissociation. Anal. Chem. 2006, 78, 1264–1271.
Cournoyer, J. J.; Pittman, J. L.; Ivleva, V. B.; Fallows, E.; Waskell, L.; Costello, C. E.; O’Connor, P. B. Deamidation: Differentiation of Aspartyl from Isoaspartyl Products in Peptides by Electron Capture Dissociation. Protein Sci. 2005, 14, 452–463.
O’Connor, P. B.; Cournoyer, J. J.; Pitteri, S. J.; Chrisman, P. A.; McLuckey, S. A. Differentiation of Aspartic and Isoaspartic Acids Using Electron Transfer Dissociation. J. Am. Soc. Mass Spectrom. 2006, 17, 15–19.
Horn, D. M.; Ge, Y.; McLafferty, F. W. Activated Ion Electron Capture Dissociation for Mass Spectral Sequencing of Larger (42 kDa) Proteins. Anal. Chem. 2000, 72, 4778–4784.
Yao, X. D.; Freas, A.; Ramirez, J.; Demirev, P. A.; Fenselau, C. Proteolytic O-18 Labeling for Comparative Proteomics: Model Studies with Two Serotypes of Adenovirus. Anal. Chem. 2004, 76, 2675–2675.
Stewart, I. I.; Thomson, T.; Figeys, D. O-18 Labeling: A Tool for Proteomics. Rapid Commun. Mass Spectrom. 2001, 15, 2456–2465.
Heller, M.; Mattou, H.; Menzel, C.; Yao, X. D. Trypsin Catalyzed O-16-to-O-18 Exchange for Comparative Proteomics: Tandem Mass Spectrometry Comparison Using MALDI-TOF, ESI-QTOF, and ESI-Ion Trap Mass Spectrometers. J. Am. Soc. Mass Spectrom. 2003, 14, 704–718.
Yao, X. D.; Freas, A.; Ramirez, J.; Demirev, P. A.; Fenselau, C. Proteolytic O-18 Labeling for Comparative Proteomics: Model Studies with Two Serotypes of Adenovirus. Anal. Chem. 2001, 73, 2836–2842.
Miyagi, M.; Rao, K. C. S. Proteolytic O-18-Labeling Strategies for Quantitative Proteomics. Mass Spectrom. Rev. 2007, 26, 121–136.
Katta, V.; Chait, B. T. Hydrogen/Deuterium Exchange Electrospray Ionization Mass Spectrometry: A Method for Probing Protein Conformational Changes in Solution. J. Am. Chem. Soc. 1993, 115, 6317–6321.
Wales, T. E.; Engen, J. R. Hydrogen Exchange Mass Spectrometry for the Analysis of Protein Dynamics. Mass Spectrom. Rev. 2006, 25, 158–170.
Garcia, R. A.; Pantazatos, D.; Villarreal, F. J. Hydrogen/Deuterium Exchange Mass Spectrometry for Investigating Protein-Ligand Interactions. Assay Drug. Dev. Technol. 2004, 2, 81–91.
Kheterpal, I.; Cook, K. D.; Wetzel, R. Hydrogen/Deuterium Exchange Mass Spectrometry Analysis of Protein Aggregates. Methods Enzymol. 2006, 413, 140–166.
Kheterpal, I.; Wetzel, R. Hydrogen/Deuterium Exchange Mass Spectrometries—A Window into Amyloid Structure. Acc. Chem. Res. 2006, 39, 584–593.
Xiao, G.; Bondarenko, P. V.; Jacob, J.; Chu, G. C.; Chelius, D. O18 Labeling Method for Identification and Quantification of Succinimide in Proteins. Anal. Chem. 2007, 79, 2714–2721.
Kubota, K.; Yoneyama-Takazawa, T.; Ichikawa, K. Determination of Sites Citrullinated by Peptidylarginine Deiminase Using O-18 Stable Isotope Labeling and Mass Spectrometry. Rapid Commun. Mass Spectrom. 2005, 19, 683–688.
Schnolzer, M.; Jedrzejewski, P.; Lehmann, W. D. Protease-Catalyzed Incorporation of O-18 into Peptide Fragments and Its Application for Protein Sequencing by Electrospray and Matrix-Assisted Laser Desorption/Ionization Mass Spectrometry. Electrophoresis. 1996, 17, 945–953.
Antonov, V. K.; Ginodman, L. M.; Rumsh, L. D.; Kapitannikov, Y. V.; Barshevskaya, T. N.; Yavashev, L. P.; Gurova, A. G.; Volkova, L. I. Studies on the Mechanisms of Action of Proteolytic Enzymes Using Heavy Oxygen Exchange. Eur. J. Biochem. 1981, 117, 195–200.
O’Connor, P. B.; Pittman, J. L.; Thomson, B. A.; Budnik, B. A.; Cournoyer, J. C.; Jebanathirajah, J.; Lin, C.; Moyer, S.; Zhao, C. A New Hybrid Electrospray Fourier Transform Mass Spectrometer: Design and Performance Characteristics. Rapid Commun. Mass Spectrom. 2006, 20, 259–266.
Jebanathirajah, J. A.; Pittman, J. L.; Thomson, B. A.; Budnik, B. A.; Kaur, P.; Rape, M.; Kirschner, M.; Costello, C. E.; O’Connor, P. B. Characterization of a New qQq-FTICR Mass Spectrometer for Post-translational Modification Analysis and Top-Down Tandem Mass Spectrometry of Whole Proteins. J. Am. Soc. Mass Spectrom. 2005, 16, 1985–1999.
Tsybin, Y. O.; Witt, M.; Baykut, G.; Hakansson, P. Electron Capture Dissociation Fourier Transform Ion Cyclotron Resonance Mass Spectrometry in the Electron Energy Range 0–50 eV. Rapid Commun. Mass Spectrom. 2004, 18, 1607–1613.
Tsybin, Y. O.; Hakansson, P.; Budnik, B. A.; Haselmann, K. F.; Kjeldsen, F.; Gorshkov, M.; Zubarev, R. A. Improved Low-Energy Electron Injection Systems for High Rate Electron Capture Dissociation in Fourier Transform Ion Cyclotron Resonance Mass Spectrometry. Rapid Commun. Mass Spectrom. 2001, 15, 1849–1854.
Tsybin, Y. O.; Ramstrom, M.; Witt, M.; Baykut, G.; Hakansson, P. Peptide and Protein Characterization by High-Rate Electron Capture Dissociation Fourier Transform Ion Cyclotron Resonance Mass Spectrometry. J. Mass Spectrom. 2004, 39, 719–729.
Cheung, W. Y. Calmodulin Plays a Pivotal Role in Cellular Regulation. Science. 1980, 207, 19–27.
Potter, S. M.; Henzel, W. J.; Aswad, D. W. In-Vitro Aging of Calmodulin Generates Isoaspartate at Multiple Asn-Gly and Asp-Gly Sites in Calcium-Binding Domains II, III, and IV. Protein Sci. 1993, 2, 1648–1663.
Zubarev, R. A. Electron-Capture Dissociation Tandem Mass Spectrometry. Curr. Opin. Biotechnol. 2004, 15, 12–16.
Yergey, J. A. A general approach to calculating isotopic distributions for mass spectrometry. Int. J. Mass Spectrom. Ion Processes. 1983, 52, 337–349.
Author information
Authors and Affiliations
Corresponding author
Additional information
Published online March 5, 2008
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Li, X., Cournoyer, J.J., Lin, C. et al. Use of 18O labels to monitor deamidation during protein and peptide sample processing. J Am Soc Mass Spectrom 19, 855–864 (2008). https://doi.org/10.1016/j.jasms.2008.02.011
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1016/j.jasms.2008.02.011