Abstract
Purpose
Connexin 43 (Cx43) is a widely expressed gap junction protein. It can also regulate various gap-junction independent processes, including cellular proliferation. The latter regulatory functions have been attributed to its carboxy-terminal domain, CT-Cx43. CT-Cx43 has been found to be expressed independent of full-length Cx43 in various cell types. Its nuclear localization has additionally raised the possibility that it may regulate the expression of particular genes, including miRNAs, known play a role in the regulation of cellular proliferation. Here, we set out to uncover the molecular mechanism(s) underlying CT-Cx43 mediated gene (de-)regulation in human breast cancer.
Methods
Western blotting and quantitative real time PCR were carried to assess the expression of CT-Cx43 and miR-125b in a panel of 60 primary human breast cancer tissues and its paired normal adjacent tissues. In addition, CT-Cx43 was exogenously expressed in the breast cancer-derived cell line MCF-7 and its effect on the expression of miR-125b and its downstream target p53 were evaluated, as well as its effect on cellular proliferation and death using MTT and LDH assays, respectively.
Results
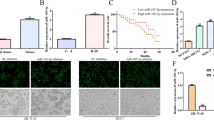
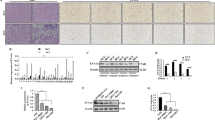
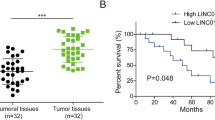
We found that CT-Cx43, but not full-length Cx43, was down-regulated in low grade human breast cancers. In addition, we found that the tumor suppressor protein p53 exhibited a decreased expression in the CT-Cx43 down-regulated samples. Interestingly, we found that miR-125b, a negative regulator of p53, exhibited an inverse expression relationship with CT-Cx43 in the breast cancer samples tested. This inverse relationship was confirmed by exogenous expression of CT-Cx43 in MCF-7 cells. In addition, we found that CT-Cx43 up-regulation and subsequent miR-125b down-regulation resulted in a decreased proliferation of MCF-7 cells.
Conclusions
Our data suggest a mechanism by which CT-Cx43 may regulate cell proliferation. Targeting of CT-Cx43 and/or miR-125b may be instrumental for therapeutic intervention in human breast cancer.




Similar content being viewed by others
References
G. Mese, G. Richard, T.W. White, Gap junctions: basic structure and function. J. Investig. Dermatol. 127, 2516–2524 (2007)
H.A. Dbouk, R.M. Mroue, M.E. El-Sabban, R.S. Talhouk, Connexins: a myriad of functions extending beyond assembly of gap junction channels, Cell. Commun. Signals 7, 4 (2009)
K. Willecke, J. Eiberger, J. Degen, D. Eckardt, A. Romualdi, M. Guldenagel, U. Deutsch, G. Sohl, Structural and functional diversity of connexin genes in the mouse and human genome. Biol. Chem. 383, 725–737 (2002)
G. Pointis, J. Gilleron, D. Carette, D. Segretain, Physiological and physiopathological aspects of connexins and communicating gap junctions in spermatogenesis, philosophical transactions of the royal society of london. Series B, Biol. Sci. 365, 1607–1620 (2010)
S. Sirnes, J. Bruun, M. Kolberg, A. Kjenseth, G.E. Lind, A. Svindland, A. Brech, A. Nesbakken, R.A. Lothe, E. Leithe, E. Rivedal, Connexin43 acts as a colorectal cancer tumor suppressor and predicts disease outcome. Int. J. Cancer 131, 570–581 (2012)
M. Oyamada, Y. Oyamada, T. Takamatsu, Regulation of connexin expression. Biochim. Bophys. Acta 1719, 6–23 (2005)
R. Ismail, R. Rashid, K. Andrabi, F.Q. Parray, S. Besina, M.A. Shah, M. Ul Hussain, Pathological implications of Cx43 down-regulation in human colon cancer. Asian Pac. J. Cancer Prev. 15, 2987–2991 (2014)
C. Moorby, M. Patel, Dual functions for connexins: Cx43 regulates growth independently of gap junction formation. Exp. Cell Res. 271, 238–248 (2001)
J.E. Trosko, R.J. Ruch, Gap junctions as targets for cancer chemoprevention and chemotherapy. Curr. Drug Targets 3, 465–482 (2002)
X. Dang, B.W. Doble, E. Kardami, The carboxy-tail of connexin-43 localizes to the nucleus and inhibits cell growth. Mol. Cell. Biochem. 242, 35–38 (2003)
W.D. Foulkes, p53–master and commander. New Eng. J. Med. 357, 2539–2541 (2007)
E. Wawryk-Gawda, P. Chylinska-Wrzos, M. Lis-Sochocka, K. Chlapek, K. Bulak, M. Jedrych, B. Jodlowska-Jedrych, P53 protein in proliferation, repair and apoptosis of cells. Protoplasma 251, 525–533 (2014)
L.A. Voutsadakis, The chemosensitivity of testicular germ cell tumors. Cell. Oncol. 37, 79–94 (2014)
V.S. Bielicka, P. Domagala, D. Bielicki, K. Safranow, W. Domagala, Thymidylate synthase expression and P21WAF1/P53 phenotype of colon cancers identify patients who may benefit from 3-flurouracil based therapy. Cell. Oncol. 37, 17–28 (2014)
N. Almog, V. Rotter, Involvement of p53 in cell differentiation and development. Biochim. Biophys. Acta 1333, F1–27 (1997)
A.J. Giaccia, M.B. Kastan, The complexity of p53 modulation: emerging patterns from divergent signals. Genes Dev. 12, 2973–2983 (1998)
K. Sakaguchi, J.E. Herrera, S. Saito, T. Miki, M. Bustin, A. Vassilev, C.W. Anderson, E. Appella, DNA damage activates p53 through a phosphorylation-acetylation cascade. Genes Dev. 12, 2831–2841 (1998)
S.W. Lowe, Activation of p53 by oncogenes. Endocr. Relat. Cancer 6, 45–48 (1999)
L. Zheng, J.Q. Ren, H. Li, Z.L. Kong, H.G. Zhu, Downregulation of wild-type p53 protein by HER-2/neu mediated PI3K pathway activation in human breast cancer cells: its effect on cell proliferation and implication for therapy. Cell Res. 14, 497–506 (2004)
N. Rivlin, R. Brosh, M. Oren, V. Rotter, Mutations in the p53 tumor suppressor gene: important milestones at the various steps of tumorigenesis. Genes Cancer 2, 466–474 (2011)
P.A. Muller, K.H. Vousden, p53 mutations in cancer. Nat. Cell Biol. 15, 2–8 (2013)
M. Ul Hussain, Micro-RNAs (miRNAs): genomic organisation, biogenesis and mode of action. Cell Tissue Res. 349, 405–413 (2012)
R. Maqbool, M. Ul Hussain, MicroRNAs and human diseases: diagnostic and therapeutic potential. Cell Tissue Res. 358, 1–15 (2014)
R. Nagadia, P. Pandit, W.B. Coman, J.C. White, C. Punyadeera, MicroRNAs in head and neck cancer revisited. Cell. Oncol. 36, 1–7 (2013)
Y. Wang, M. Li, W. Zang, Y. Ma, N. Wang, P. Li, G. Zhao, MiR-429 upregulation induces apoptosis and suppresses invasion by targeting Bcl-2 and SP-1 in esophageal carcinoma. Cell. Oncol. 36, 385--394 (2013)
L. Rask, E. Balslev, R. Sokilde, E. Hogdall, H. Flyger, J. Eriksen, T. Litman, Differential expression of miR-139, miR-486 and miR-21 in breast cancer patients sub classified according to lymph node status. Cell. Oncol. 37, 215–227 (2014)
C. Salazar, R. Nagadia, P. Pandit, J.C. White, N. Banerjee, N. Dimitrova, W.B. Coman, C. Pundyadeera, A novel saliva based microRNA biomarker panel to detect head and neck cancers. Cell. Oncol. 37, 331–338 (2014)
J. Banzhaf-Strathmann, D. Edbauer, Good guy or bad guy: the opposing roles of microRNA 125b in cancer. Cell Commun. Signal 12, 30 (2014)
C. Chen, D.A. Ridzon, A.J. Broomer, Z. Zhou, D.H. Lee, J.T. Nguyen, M. Barbisin, N.L. Xu, V.R. Mahuvakar, M.R. Andersen, K.Q. Lao, K.J. Livak, K.J. Guegler, Real-time quantification of microRNAs by stem-loop RT-PCR. Nucleic Acids Res. 33, e179 (2005)
M. Ul-Hussain, S. Olk, B. Schoenebeck, B. Wasielewski, C. Meier, N. Prochnow, C. May, S. Galozzi, K. Marcus, G. Zoidl, R. Dermietzel, Internal ribosomal entry site (IRES) activity generates endogenous carboxyl-terminal domains of Cx43 and is responsive to hypoxic conditions. J. Biol. Chem. 289, 20979–20990 (2014)
N. Wu, X. Lin, X. Zhao, L. Zheng, L. Xiao, J. Liu, L. Ge, S. Cao, MiR-125b acts as an oncogene in glioblastoma cells and inhibits cell apoptosis through p53 and p38MAPK-independent pathways. Br. J. Cancer 109, 2853–2863 (2013)
A. Feliciano, J. Castellvi, A. Artero-Castro, J.A. Leal, C. Romagosa, J. Hernandez-Losa, V. Peg, A. Fabra, F. Vidal, H. Kondoh, Y.C.S. Ramon, M.E. Lleonart, miR-125b acts as a tumor suppressor in breast tumorigenesis via its novel direct targets ENPEP, CK2-alpha, CCNJ, and MEGF9. PLoS One 8, e76247 (2013)
A. Geurts van Kessel, The cancer genome: from structure to function. Cell. Oncol. 37, 155–165 (2014)
H.J. Wang, Y.Q. Guo, G. Tan, L. Dong, L. Cheng, K.J. Li, Z.Y. Wang, H.F. Luo, miR-125b regulates side population in breast cancer and confers a chemoresistant phenotype, J.Cell. Biochem. 114, 2248–2257 (2013)
H. Wang, G. Tan, L. Dong, L. Cheng, K. Li, Z. Wang, H. Luo, Circulating MiR-125b as a marker predicting chemoresistance in breast cancer, PloS One.7, e34210 (2012)
D.A. Iacobas, E. Scemes, D.C. Spray, Gene expression alterations in connexin null mice extend beyond the gap junction. Neurochem. Int. 45, 243–250 (2004)
E. Kardami, X. Dang, D.A. Iacobas, B.E. Nickel, M. Jeyaraman, W. Srisakuldee, J. Makazan, S. Tanguy, D.C. Spray, The role of connexins in controlling cell growth and gene expression. Prog. Biophys. Mol. Biol. 94, 245–264 (2007)
M. Hollstein, D. Sidransky, B. Vogelstein, C.C. Harris, p53 mutations in human cancers. Science 253, 49–53 (1991)
U.M. Moll, G. Riou, A.J. Levine, Two distinct mechanisms alter p53 in breast cancer: mutation and nuclear exclusion. Proc. Natl. Acad. Sci. U. S. A. 89, 7262–7266 (1992)
Acknowledgments
The junior research fellowships to RM (UGC-JRF 19-06/2011 Ci EU-IV), SN (ICMR JRF-2012/HRD-29) and project fellowships to RI (BT/PR14336/MED/30/493/2010) and RR (SR/WOS-A/LS-661/2012) are highly acknowledged. The present work was supported by project grant (BT/PR11917/MED/30/181/2009) from the Department of Biotechnology (DBT), New Delhi, to MUH.
Conflict of interest
All authors declare no conflict of interest
Author information
Authors and Affiliations
Corresponding author
Additional information
Raihana Maqbool and Rabiya Rashid contributed equally to the work.
Electronic supplementary material
Supplementary Fig. 1
Total RNA was isolated from the normal and breast cancer samples as described in material and methods. To check the integrity of the RNA, 5 μl of each sample was loaded on formaldehyde agarose gel. After electrophoresis, the agarose gel was incubated for 5–10 min in MOPS buffer containing ethidium bromide. The gel was visualized in gel-documentation apparatus. As shown in suppl. Fig. 1, intact and prominent bands, corresponding to 28S and18S rRNA can be detected. Moreover, RNA bands corresponding to small RNAs can also be detected. Thus, the presence of intact RNA bands established the integrity of the RNA obtained from the tissue samples. (GIF 181 kb)
Supplementary Fig. 2
Total protein was extracted from the tumor and adjacent normal breast samples as described in material and methods. 30 μg of protein was loaded on SDS-PAGE and electrophoresis was performed. After PAGE, Western blot was performed using anti-Cx43 antibody as described previously. As shown in suppl. Fig. 2, anti-Cx43 detected FL-Cx43 and CT-Cx43 in the normal samples (N). However, the tumor samples (T) showed decreased expression of FL-Cx43 as well as CT-Cx43. (GIF 169 kb)
Supplementary Fig. 3
To investigate whether CT-Cx43 has the potential to bind the miR-125b promoter, we expressed CT-Cx43 as GST fusion protein in bacterial system. The GST-CT-Cx43 was purified from the bacterial lysate using Gluthatione-agarose beads. Additionally, 200 bps of miR-125b promoter (upstream of transcriptional site containing the Tata box; Gene ID: 406,911) was PCR amplified using the forward primers 5`-TTTGAAAAGACACTAAATCCT-3` and the reverse primer 5’ACAATGTTTTCTTTCTTGAGA-3`. Prior to PCR, the primers were 32p labeled using T4 polynucleotide kinase and gamma phosphate (32P) labeled ATP. The 200 bp DNA probe and purified GST-CT-CX43 were incubated together and gel-shift assay was performed. As shown in suppl. Fig. 3, lane II, addition of CT-Cx43 did not show any binding to the miR-125 DNA. Moreover, addition of anti-CT-Cx43 did not show and supershift of miR-125b probe (lane III). Thus, our initial data indicate that CT-Cx43 does not bind the proximal promoter element of miR-125b. Further upstream of 200 bp needs to further screened for any binding of CT-Cx43. (GIF 62.9 kb)
Supplementary Fig. 4
In order study the effect of CT-Cx43 on the expression of various growth related miRNAs, we transiently transfected MCF-7 cells with either CT-Cx43 plasmid or with empty plasmid. After 36 h transfection, RNA was isolated and using stem-loop primers strategy (for details consult material methods of the main manuscript) many miRNAs were PCR amplified (here denoted as “a, b, c, d, e, f, g, h, I, j), including the miR-125b. As shown in suppl Fig. 4, various miRNAs showed differential regulation between empty plasmid and CT-Cx43 transfected MCF-7 cells (compare “a & a`; e & e`; f & f`). Interestingly, miR-125b showed highly significant up-regulation in CT-Cx43 transfected MCF-7 cells. (GIF 291 kb)
Supplementary Fig. 5
In order to establish the possibility of nuclear localization of CT-Cx43 in breast samples, we isolated the nuclear and cytosolic fraction from the normal and tumor breast cancer samples using nuclear extraction kit (Sigma adrich). 30 μgs of each fraction was run on 12 % SDS-PAG and Western blot was performed using anti-Cx43 antibody as described in material and methods. As shown in Suppl. Fig. 5, anti-Cx43 antibody detected FL-Cx43 in the cytosolic fraction (Cyt) fraction of normal (N) as well as in the tumor (T) samples. Moreover, CT-Cx43 was detected in the cytosolic (Cyt.) and nuclear fractions (Nuc.) of the normal samples (N). However, very less/or no signal of CT-Cx43 was detected from the nuclear fraction (Nuc.) of tumor samples (T). Hence, it may be concluded that CT-Cx43 has the potential to localize in cell nucleus when it is expressed in the cells. (GIF 122 kb)
Rights and permissions
About this article
Cite this article
Maqbool, R., Rashid, R., Ismail, R. et al. The carboxy-terminal domain of connexin 43 (CT-Cx43) modulates the expression of p53 by altering miR-125b expression in low-grade human breast cancers . Cell Oncol. 38, 443–451 (2015). https://doi.org/10.1007/s13402-015-0240-x
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13402-015-0240-x