Abstract
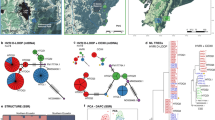
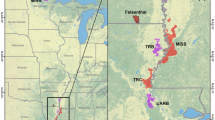
Estimating the genetic structure of a population is important for the conservation and management of wildlife. In the present study, our aim was to estimate the genetic structure of the brown bear (Ursus arctos) population in eastern Hokkaido by performing a Bayesian clustering analysis. To accomplish this goal, we used 15 microsatellites to generate genotypic data from tissue samples collected from 646 bears between 1996 and 2008. Using this genotypic data and the geographic locations where the bears were captured, GENELAND analysis detected six subpopulations. Based on the genotypic data, the STRUCTURE analysis revealed three subpopulations. As inferred from the GENELAND analysis, the core zones of the subpopulations (G-a through G-f) were located in the Shiranuka Hills (G-a), the northern area of the Shiranuka Hills (G-b), the eastern slope of the Daisetsuzan Mountains (G-c), the northern slope of the Akan Mountain Range (G-d), the Shiretoko Peninsula (G-e), and Akkeshi District (G-f). The STRUCTURE analysis indicated that G-b and G-d were influenced by gene flow from other subpopulations. National routes, towns, and farm fields were considered to have formed the distribution boundaries among the subpopulations. A high level of genetic differentiation was not observed among the six subpopulations, with the exception of G-f (F st = 1.35–0.176, D s = 0.246–0.349), which showed a geographically discontinuous distribution. We suggest that the loss of forest areas through future regional development and road building should be avoided to facilitate gene flow in brown bears in Hokkaido.






Similar content being viewed by others
References
Abe H (1980) Distribution of brown bear. In: Doubutsu Bunpu Chousa Houkoku Sho. (Honyuurui) Zenkoko Ban (Sono 2). Japan Wildlife Research Center, Tokyo, pp 87–98, in Japanese
Aoi T (2011) Home range and land use. In: Tsubota T, Yamazaki K (eds) Bears in Japan—biology of Hokkaido brown bears and Japanese black bears. University of Tokyo Press, Tokyo, pp 59–84 (in Japanese)
Bellemain E, Taberlet P (2004) Improved noninvasive genotyping method: application to brown bear (Ursus arctos) faeces. Mol Ecol Notes 4:519–522
Crooks KR (2002) Relative sensitivities of mammalian carnivores to habitat fragmentation. Conserv Biol 16:488–502
Dixon JD, Oli MK, Wooten MC, Eason TH, McCown JW, Paetkau D (2006) Effectiveness of a regional corridor in connecting two Florida black bear populations. Conserv Biol 20:155–162
Evanno G, Regnaut S, Goudet J (2005) Detecting the number of clusters of individuals using the software structure: a simulation study. Mol Ecol 14:2611–2620
Falush D, Stephens M, Pritchard JK (2003) Inference of population structure using multilocus genotype data: linked loci and correlated allele frequencies. Genetics 164:1567–1587
Glenn LP, Miller LH (1980) Seasonal movements of an Alaska Peninsula brown bear population. Int Conf Bear Res Manag 4:307–312
Goudet J (1995) FSTAT (version 1.2): a computer program to calculate F-statistics. J Hered 86:485–486
Guillot G, Estoup A, Mortier F, Cosson JF (2005a) A spatial statistical model for landscape genetics. Genetics 170:1261–1280
Guillot G, Mortier F, Estoup A (2005b) GENELAND: a computer package for landscape genetics. Mol Ecol Notes 5:712–715
Hokkaido Institute of Environmental Sciences (1994) Results of a survey related to sika deer and brown bear sightings in Hokkaido. Hokkaido Institute of Environmental Sciences, Sapporo, in Japanese
Hokkaido Institute of Environmental Sciences (1996) Report of brown bear and sika deer ecological survey II. Wildlife distribution and abundance research report: Brown bears (1991–1995). Hokkaido Institute of Environmental Sciences, Sapporo, in Japanese
Hokkaido Institute of Environmental Sciences (2000) Results of a survey related to sika deer and brown bear sighting on Hokkaido: IV. Hokkaido Institute of Environmental Sciences, Sapporo, in Japanese
Hokkaido Institute of Environmental Sciences (2004) Census of a wild animal (Ursus arctos, 1999–2003): a research report. Hokkaido Institute of Environmental Sciences, Sapporo, in Japanese
Inukai T, Kadosaki M, Tomikawa T, Mikami T, Iizuka J, Hatakeyama T, Owari K (1985) Status of the capture and the inhabitation of brown bears in Hokkaido, Japan (II). Annu rep Hist Mus Hokkaido 13:55–84 (in Japanese with English abstracts)
Itoh T, Sato Y, Mano T, Iwata R (2009a) Estimating a suitable microsatellite marker set for individual identification and parentage test of brown bear (Ursus arctos) in the Akan-Shiranuka Region, eastern Hokkaido, Japan. J For Res 14:117–122
Itoh T, Nakamura H, Koizumi S, Sato Y (2009b) Ecology and conservation of brown bear in the Urahoro area (the way for ten years and future of Urahoro Brown Bear Research Group). Bull Urahoro Hist Mus 9:17–26 (in Japanese)
Itoh T, Sato Y, Kobayashi K, Mano T, Iwata R (2012) Effective dispersal of brown bears (Ursus arctos) in eastern Hokkaido, inferred from analyses of mitochondrial DNA and microsatellites. Mamm Study 37:29–41
Latch EK, Scognamillo DG, Fike JA, Chamberlain MJ, Rhodes OE Jr (2008) Deciphering ecological barriers to North American river otter (Lontra canadensis) gene flow in the Louisiana landscape. J Hered 99:265–274
Laurence S, Coltman DW, Gorrell JC, Schulte-Hostedde AI (2011) Genetic structure of muskrat (Ondatra zibethicus) and its concordance with taxonomy in North America. J Hered 102:688–696
Lecis R, Ferrando A, Ruiz-Olmo J, Mañas S, Domingo-Roura X (2008) Population genetic structure and distribution of introduced American mink (Mustela vison) in Spain, based on microsatellite variation. Conserv Genet 9:1149–1161
Maehr DS, Smith JS, Cunningham MW, Barnwell ME, Larkin JL, Orlando MA (2003) Spatial characteristics of an isolated Florida black bear population. Southeast Nat 2:433–446
Manel S, Schwartz MK, Luikart G, Taberlet P (2003) Landscape genetics: combining landscape ecology and population genetics. Trends Ecol Evol 18:189–197
Mano T (2007) The status of brown bears in Japan. In: Japan Bear Network (ed) Understanding Asian bears to secure their future. Japan Bear Network, Ibaraki, pp 111–121
Matsuhashi T, Masuda R, Mano T, Yoshida MC (1999) Microevolution of the mitochondrial DNA control region in the Japanese brown bear (Ursus arctos) population. Mol Biol Ecol 16:676–684
McLellan BN, Hovey FW (2001) Natal dispersal of grizzly bears. Can J Zool 79:838–844
McLellan BN, Shackleton DM (1988) Grizzly bears and resource-extraction industries: effects of roads on behavior, habitat use and demography. J Appl Ecol 25:451–460
Nei M (1978) Estimation of average heterozygosity and genetic distance from a small number of individuals. Genetics 89:583–590
Nellemann C, Støen OG, Kindberg J, Swenson JE, Vistnes I, Ericsson G, Katajisto J, Kaltenborn BP, Martin J, Ordiz A (2007) Terrain use by an expanding brown bear population in relation to age, recreational resorts and human settlements. Biol Conserv 138:157–165
Norén K, Carmichael L, Dalén L, Hersteinsson P, Samelius G, Fuglei E, Kapel CMO, Menyushina I, Strobeck C, Angerbjörn A (2011) Arctic fox Vulpes lagopus population structure: circumpolar patterns and processes. Oikos 120:873–885
Paetkau D, Calvert W, Stirling I, Strobeck C (1995) Microsatellite analysis of population structure in Canadian polar bears. Mol Ecol 4:347–354
Paetkau D, Shields GF, Strobeck C (1998a) Gene flow between insular, coastal and interior populations of brown bears in Alaska. Mol Ecol 7:1283–1292
Paetkau D, Strobeck C (1994) Microsatellite analysis of genetic variation in black bear populations. Mol Ecol 3:489–495
Paetkau D, Waits LP, Clarkson PL, Craighead L, Vyse E, Ward R, Strobeck C (1998b) Variation in genetic diversity across the range of North American brown bears. Conserv Biol 12:418–429
Peakall R, Smouse PE (2006) GenAlEx 6: genetic analysis in Excel. Population genetic software for teaching and research. Mol Ecol Notes 6:288–295
Pritchard JK, Stephens M, Donelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155:945–959
Proctor MF, McLellan BN, Strobeck C, Barclay RMR (2004) Gender-specific dispersal distances of grizzly bears estimated by genetic analysis. Can J Zool 82:1108–1118
Proctor MF, McLellan BN, Strobeck C, Barclay RMR (2005) Genetic analysis reveals demographic fragmentation of grizzly bears yielding vulnerably small populations. Proc R Soc B 272:2409–2416
Raymond M, Rousset F (1995) GENEPOP (version 1.2): population genetics software for exact tests and ecumenicism. J Hered 86:248–249
Rice WR (1989) Analyzing tables of statistical tests. Evolution 43:223–225
Rousset F (1997) Genetic differentiation and estimation of gene flow from F-statistics under isolation by distance. Genetics 145:1219–1228
Rousset F (1999) Genetic differentiation in populations with different classes of individuals. Theor Popul Biol 55:297–308
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425
Sato Y (2003) Changes in the number of killed brown bears in Urahoro, Hokkaido, Japan. Bull Urahoro Hist Mus 3:27–35
Sato Y, Aoi T, Kaji K, Takatsuki S (2004) Temporal changes in the population density and diet of brown bears in eastern Hokkaido, Japan. Mamm Study 29:47–53
Sato Y, Kobayashi Y, Urata T, Takatsuki S (2008) Home range and habitat use of female brown bear (Ursus arctos) in Urahoro, eastern Hokkaido, Japan. Mamm Study 33:99–109
Sato Y, Itoh T, Mori Y, Satoh Y, Mano T (2011) Dispersal of male bears into periphery inferred from mtDNA haplotypes. Ursus 22:120–132
Servheen C, Herrero S, Peyton B (1999) Bears. Status Survey and Conservation Action Plan. IUCN, Cambridge
Sumner J, Rousset F, Estoup A, Moritz C (2001) “Neighbourhood” size, dispersal and density estimates in the prickly forest skink (Gnypetoscincus queenslandiae) using individual genetic and demographic methods. Mol Ecol 10:1917–1927
Taberlet P, Camarra JJ, Griffin S, Uhrès E, Hanotte O, Waits LP, Dubois-Paganon C, Burke T, Bouvet J (1997) Noninvasive genetic tracking of the endangered Pyrenean brown bear population. Mol Ecol 6:869–876
Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S (2011) MEGA5: Molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol Biol Evol 28:2731–2739
van Oosterhout C, Hutchinson WF, Wills DPM, Shipley P (2004) MICRO-CHECKER: software for identifying and correcting geno-typing errors in microsatellite data. Mol Ecol Notes 4:535–538
Waits LP, Taberlet P, Swenson JE, Sandegren F, Franzen R (2000) Nuclear DNA microsatellite analysis of genetic diversity and gene flow in the Scandinavian brown bear (Ursus arctos). Mol Ecol 9:421–431
Weir BS, Cockerham CC (1984) Estimating F-statistics for the analysis of population structure. Evolution 38:1358–1370
Zedrosser A, Støen OG, Sæbø S, Swenson JE (2007) Should I stay or should I go? Natal dispersal in the brown bear. Anim Behav 74:369–376
Acknowledgments
We acknowledge the farmers of Urahoro, the staff of the Urahoro Town Office, the members of the Hunters Association of Urahoro and Kami-Urahoro, the staff of the Urahoro Brown Bear Research Group, the staff of Shiretoko Nature Foundation, and the Hokkaido Institute of Environmental Sciences for their cooperation during sampling and for their support during the present study. We are grateful to T. Urata for his help with sampling and support during fieldwork in the Urahoro region. This research was supported in part by a Grant-in-Aid for Young Scientists (B), 19780122, from the Ministry of Education, Culture, Sports, Science and Technology, Japan, to Y.S., by research fellowship of the Japan Society for the Promotion of Science for Young Scientists to T.I., and the Kimunkamui project.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by: Allan McDevitt
Rights and permissions
About this article
Cite this article
Itoh, T., Sato, Y., Tsuruga, H. et al. Estimating the population structure of brown bears in eastern Hokkaido based on microsatellite analysis. Acta Theriol 58, 127–138 (2013). https://doi.org/10.1007/s13364-012-0095-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13364-012-0095-8