Abstract
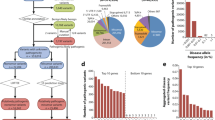
A survey of a select panel of 14 genetic diseases with mixed inheritance confirms that, while autosomal recessive (AR) disease genes are more numerous than autosomal dominant (AD) or X-linked (XL) ones, they make a smaller average contribution to disease. Data collected from N-ethyl-N-nitrosourea (ENU) mutagenesis studies show a similar excess of AR mutations. The smaller AR contribution may partially reflect disease severity, but only in the comparison of AR with AD mutations. On the contrary, XL mutations for the 14 diseases are generally more severe. Genome-wide associations studies (GWAS) data provide fresh insight into the shortage, with a limited negative selection effect mediated by the pleiotropic expression of recessive disease genes in other deleterious phenotypes. Genomic data provide further evidence of purging selection in a past European population bottleneck followed by a dramatic population explosion, now more clearly associated with past climate change. We consider these likely to be the main factors responsible for the low AR to AD/XL inheritance ratio.


Similar content being viewed by others
References
Acevedo-Arozena A, Wells S, Potter P, Kelly M, Cox RD, Brown SD (2008) ENU mutagenesis, a way forward to understand gene function. Annu Rev Genomics Hum Genet 9:49–69
Allan W (1939) Relation of hereditary pattern to clinical severity as illustrated by peroneal atrophy. Arch Intern Med 63:1123–1131
Bittles AH, Black ML (2010) Evolution in health and medicine Sackler colloquium: Consanguinity, human evolution, and complex diseases. Proc Natl Acad Sci U S A 107(Suppl 1):1779–1786
Bittles AH, Neel JV (1994) The costs of human inbreeding and their implications for variations at the DNA level. Nat Genet 8:117–121
Blekhman R, Man O, Herrmann L, et al (2008) Natural selection on genes that underlie human disease susceptibility. Curr Biol 18:883–889
Buckley RH (2004) The multiple causes of human SCID. J Clin Invest 114:1409–1411
Chang YF, Imam JS, Wilkinson MF (2007) The nonsense-mediated decay RNA surveillance pathway. Annu Rev Biochem 76:51–74
Charlesworth D, Willis JH (2009) The genetics of inbreeding depression. Nat Rev Genet 10:783–796
Coventry A, Bull-Otterson LM, Liu X et al (2010) Deep resequencing reveals excess rare recent variants consistent with explosive population growth. Nat Commun 1:131
Dean M, Carrington M, O’Brien SJ (2002) Balanced polymorphism selected by genetic versus infectious human disease. Annu Rev Genomics Hum Genet 3:263–292
Erickson RP (2009) Autosomal recessive diseases among the Athabaskans of the southwestern United States: recent advances and implications for the future. Am J Genet A 149A:2602–2611
Eriksson A, Betti L, Friend AD, et al (2012) Late Pleistocene climate change and the global expansion of anatomically modern humans. Proc Natl Acad Sci U S A 109:16089–16094
Fu W, O’Connor TD, Jun G et al (2013) Analysis of 6,515 exomes reveals the recent origin of most human protein-coding variants. Nature 493:216–220
Furney SJ, Albà MM, López-Bigas N (2006) Differences in the evolutionary history of disease genes affected by dominant or recessive mutations. BMC Genomics 7:165
Gabriková D, Bernasovská J, Mačeková S et al (2012) Unique frequencies of HFE gene variants in Roma/Gypsies. J Appl Genet 53:183–187
Gabrikova D, Mistrik M, Bernasovska J et al (2013) Founder mutations in NDRG1 and HK1 genes are common causes of inherited neuropathies among Roma/Gypsies in Slovakia. J Appl Genet 54:455–460
Hartong DT, Berson EL, Dryja TP (2006) Retinitis pigmentosa. Lancet 368:1795–1809
Hrabé de Angelis MH, Flaswinkel H, Fuchs H et al (2000) Genome-wide, large-scale production of mutant mice by ENU mutagenesis. Nat Genet 25:444–447
Inoue K, Ohyama T, Sakuragi Y et al (2004) Translation of SOX10 3’ untranslated region causes a complex severe neurocristopathy by generation of a deleterious functional domain. Hum Mol Genet 16:3037–3046
Kääriäinen H (1987) Polycystic kidney disease in children: a genetic and epidemiological study of 82 Finnish patients. J Med Genet 24:474–481
Keinan A, Clark AG (2012) Recent explosive human population growth has resulted in an excess of rare genetic variants. Science 336:740–743
Keinan A, Mullikin JC, Patterson N, Reich D (2007) Measurement of the human allele frequency spectrum demonstrates greater genetic drift in East Asians than in Europeans. Nat Genet 39:1251–1255
Kenneson A, Kolor K, Yang Q, Olney RS, Rasmussen SA, Friedman JM (2006) Trends and racial disparities in muscular dystrophy deaths in the United States, 1983–1998: an analysis of multiple cause mortality data. Am J Med Genet A 140:2289–2297
Krawczak M, Ball EV, Cooper DN (2008) Neighboring-nucleotide effects on the rates of germ-line single-base-pair substitution in human genes. Am J Hum Genet 63:474–488
Kryukov GV, Pennacchio LA, Sunyaev SR (2007) Most rare missense alleles are deleterious in humans: implications for complex disease and association studies. Am J Hum Genet 80:727–739
Lohmueller KE, Indap AR, Schmidt S et al (2008) Proportionally more deleterious genetic variation in European than in African populations. Nature 451:994–997
Lynch M (2010) Rate, molecular spectrum, and consequences of human mutation. Proc Natl Acad Sci U S A 107:961–968
MacArthur DG, Balasubramanian S, Frankish A et al (2012) A systematic survey of loss-of-function variants in human protein-coding genes. Science 335:823–828
Marth GT, Czabarka E, Murvai J, Sherry ST (2004) The allele frequency spectrum in genome-wide human variation data reveals signals of differential demographic history in three large world populations. Genetics 166:351–372
McQuillan R, Eklund N, Pirastu N, et al (2012) Evidence of inbreeding depression on human height. PLoS Genet 8(7):e1002655
Mitchison NA, Clarke B (2008) From ENU mutagenesis to population genetics. Mamm Genome 19:221–225
Mitchison NA, Bhattacharya S, Tuddenham EG (2011) Human congenital diseases with mixed modes of inheritance have a shortage of recessive disease. A demographic scenario? Ann Hum Genet 75:688–693
Morriss-Kay GM (2010) The evolution of human artistic creativity. J Anat 216:158–176
Mort M, Ivanov D, Cooper DN, Chuzhanova NA (2008) A meta-analysis of nonsense mutations causing human genetic disease. Hum Mutat 29:1037–1047
Morton NE, Crow JF, Muller HJ (1956) An estimate of the mutational damage in man from data on consanguineous marriages. Proc Natl Acad Sci U S A 42:855–863
Nelson J, Crowhurst J, Carey B, Greed L (2003) Incidence of the mucopolysaccharidoses in Western Australia. J Med Genet A 123A(3):310–313
Nguyen N, Judd LM, Kalantzis A, Whittle B, Giraud AS, van Driel IR (2011) Random mutagenesis of the mouse genome: a strategy for discovering gene function and the molecular basis of disease. Am J Physiol Gastrointest Liver Physiol 300:G1–G11
Pelak K, Shianna KV, Ge D et al (2010) The characterization of twenty sequenced human genomes. PLoS Genet 6(9):e1001111
Pettigrew HD, Teuber SS, Gershwin ME (2009) Clinical significance of complement deficiencies. Ann N Y Acad Sci 1173:108–123
Peyvandi F, Spreafico M (2008) National and international registries of rare bleeding disorders. Blood Transfus 6(Suppl 2):s45–s48
Saporta AS, Sottile SL, Miller LJ, Feely SM, Siskind CE, Shy ME (2011) Charcot–Marie–Tooth disease subtypes and genetic testing strategies. Ann Neurol 69:22–33
Sohocki MM, Daiger SP, Bowne SJ et al (2001) Prevalence of mutations causing retinitis pigmentosa and other inherited retinopathies. Hum Mutat 17:42–51
Sowińska-Seidler A, Socha M, Jamsheer A (2014) Split-hand/foot malformation - molecular cause and implications in genetic counseling. J Appl Genet 55(1):105–115
Vogel F (1984) Clinical consequences of heterozygosity for autosomal-recessive diseases. Clin Genet 25:381–415
Weiss J, Hurley LA, Harris RM et al (2012) ENU mutagenesis in mice identifies candidate genes for hypogonadism. Mamm Genome 23:346–355
Williamson SH, Hernandez R, Fledel-Alon A, Zhu L, Nielsen R, Bustamante CD (2005) Simultaneous inference of selection and population growth from patterns of variation in the human genome. Proc Natl Acad Sci U S A 102:7882–7887
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Erickson, R.P., Mitchison, N.A. The low frequency of recessive disease: insights from ENU mutagenesis, severity of disease phenotype, GWAS associations, and demography: an analytical review. J Appl Genetics 55, 319–327 (2014). https://doi.org/10.1007/s13353-014-0203-3
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13353-014-0203-3