Abstract
The oomycete genus Phytophthora contains a large number of plant pathogens that cause significant damage to natural and agricultural systems. Until recently species have been distinguished using a limited set of morphological characters. The development of DNA-based technologies has revealed much broader and more complex diversity than previously recognised, and has led to the recent description of many new species. This review looks at the underlying mechanisms for the generation of diversity within the genus. The intercontinental movement and transplantation of infected plant material partially explains the appearance of new species in unexpected places. However, it is also likely that novel species arise as a result of the hybridisation and rapid evolution of introduced species under episodic selection pressures. Hybrid progeny may possess equal or greater virulence than parent species, thereby posing an increasing risk to our natural environment and agricultural production systems. These discoveries amplify the threats posed by the introduction of plant pathogens into new environments, and expose a crucial weakness in current evidence-based biosecurity regimes. Further work is required to identify hybrids, anticipate and understand the occurrence of hybridisation, and to implement appropriate quarantine and risk management measures.
Similar content being viewed by others
Introduction
The genus Phytophthora contains plant pathogens that cause significant damage to both agricultural and natural systems. DNA and RNA sequencing technologies have been used to describe new species responsible for recent ecologically and economically damaging disease outbreaks (Tyler et al. 2006; Haas et al. 2009; Runge et al. 2011; Grunwald et al. 2012; Yang et al. 2014). Consequently, about 70 of the 120 or so currently described species of Phytophthora have been added since 2000 (Fig. 1) (Érsek and Ribeiro 2010; Runge et al. 2011; Yang et al. 2014), and Brasier (2009) predicts there may be up to 600 extant species.
Number of Phytophthora species described within each of the currently recognised molecular phylogenetic clades discovered before (pale grey) and after (dark grey) the year 2000 (Scott et al. 2013)
While new technologies have uncovered previously unrecognised diversity in situ, trade in infected plants has been linked to new confrontations in exotic environments (Brasier 2008; Davison et al. 2006; Fry et al. 1993; Grunwald et al. 2012; Ristaino 2002; Scott et al. 2013). Otherwise benign species have been shown to evolve rapidly when exposed to selective pressures in new environments, a phenomenon termed “episodic selection” (Brasier 1995). This selection, together with the potential for hybridisation of introduced and indigenous species, can lead to the rapid emergence of new or modified species.
The emergence of new Phytophthora species has very serious implications for biosecurity and risk management (Scott et al. 2013). In Australia for example, Phytophthora dieback has always been attributed to P. cinnamomi, however recent surveys of waterways in Western Australia, and soils in the Gondwana Rainforests of Eastern Australia, have identified the presence of numerous known and unknown Phytophthora species (Burgess et al. 2009; Daniel et al. unpublished; Huberli et al. 2013). This review will argue that, as well as new technologies that distinguish existing but morphologically similar species, the emergence of new Phytophthora species can be linked to globalisation and international plant trade, resulting in episodic selection, hybridisation and the rapid evolution of novel species, some with extended host ranges.
Identifying Phytophthora species
Traditionally, fungal and oomycete species have been identified on a limited set of morphological characters that may be homoplastic, and may vary with environmental conditions (Cooke and Duncan 1997; Cooke et al. 2000; Förster et al. 2000; Burgess et al. 2009). Consequently, morphology alone is now considered to be an unreliable indicator of identity and phylogenetic relationships (Burgess et al. 2009; Jung and Burgess 2009; Jung et al. 2011; Villa et al. 2006). Cooke et al. (2000) used rRNA Internal Transcribed Spacer (ITS) sequences to construct a 10-clade phylogeny that has since been supported by further DNA analyses (Blair et al. 2008; Runge et al. 2011; Yang et al. 2014). These molecular approaches have been vital in revealing the evolution and natural phylogeny of the genus.
Episodic selection as a force driving rapid evolution
The increased volume of world trade in plants has led to an increase in the spread of plant pathogens (Brasier 2008; Scott et al. 2013). Poor hygiene in nurseries, the use of systemic fungicides that suppress disease symptoms without eradicating the pathogen (Linderman and Davis 2006), and the use of recycled irrigation water may have further contributed to an increase in the regional and global spread of plant pathogens (Brasier et al. 2003; Huberli et al. 2013; Scott et al. 2013). The impact of these exotic plant pathogens may also be intensified due to habitat disturbance, climate change and pollution (Scott et al. 2013).
Within their natural distributions the coevolution of plants and their associated microbial pathogens has resulted in an evolutionary arms race that tends to dampen the severity of epiphytotics (Kaschani and Van der Hoorn 2011). Because the “founder effect” initially limits the genetic diversity of an introduced species, the lonely intruder may fail to survive and establish (Facon et al. 2006; Mayr 1942). Alternatively, the naïve host species may be at a disadvantage, not having co-evolved with the pathogen, and the invasion of a new environment may release the pathogen from environmental and host constraints, precipitating disastrous environmental damage and crop losses (Facon et al. 2006).
Consequently, the most destructive genotypes of the pathogen establish dominant founder populations. This is a particular issue if the virulent new pathogen is introduced into monocultures of an agricultural crop, where plants lack genetic diversity and tend to be bred for yield and robustness against current, rather than exotic, pathogens. There are countless examples of exotic plant pathogens causing disease epidemics throughout history, including the Irish Potato Famine in the mid-19th century (Ristaino 2002), the coffee rust epidemics in Ceylon (1870s) and Brazil (1970s) (Arneson 2011), the two Dutch elm disease pandemics in the 20th century (Heybroek 1993), Phytophthora dieback in Australia in the last 50 years (Podger 1968), and stripe rust of wheat in Australasia in the 1980s (Wellings and McIntosh 1990).
Under certain circumstances episodic selection of founder populations can lead to the rapid selection and evolution of introduced organisms (Brasier 1995; Man in’t Veld et al. 2007). Alternatively, episodic selection pressures such as geographic transposition, exposure to a new host, a new climate or different competition can lead to recombination and the generation of better adapted genotypes (Fig. 2) (Brasier 1995; Facon et al. 2006; Ahmed et al. 2012). In the case of exotic Phytophthora pathogens, episodic selection has already resulted in the emergence of more virulent strains and species that appear better equipped to target indigenous hosts (Bertier et al. 2013; Bonants et al. 2000; Goss et al. 2011).
Some of the potential outcomes of geographic transposition. Taxon A and B are geographically isolated taxa and taxon B is introduced into the territory of taxon A (Brasier 1995)
How geographic transposition generates novel genotypes
The transport of Phytophthora isolates can lead to the emergence of novel genotypes and species through both sexual reproduction and hybridisation events, posing another serious risk to ecosystems and agrosystems. Typically, the introduction of isolates from a single mating type of a heterothallic species establishes clonal founder populations (Peters et al. 2014). However further introductions of infected plant material increase the probability of sexual reproduction through the introduction of compatible mating types (Drenth et al. 1994; Fry et al. 1993; Peters et al. 2014).
Sexual recombination generates genetic diversity, which, when followed by episodic selection pressures, inevitably results in progeny with enhanced fitness, virulence and host range (Drenth and Goodwin 1999; Wuethrich 1998). These “Red Queens” out-compete existing genotypes within the population, and pose real threats to ecosystems and agriculture around the world (Clay and Kover 1996).
Sexual reproduction in most heterothallic Phytophthora populations is apparently rare, even when both mating types are present, and clonal populations dominate (Linde et al. 1997). There is little evidence of sexual reproduction occurring amongst populations of P. cinnamomi in Australia or South Africa (Dobrowolski et al. 2003; Linde et al. 1997), P. palmivora and P. capcisi in the tropics (Drenth and Guest 2013; Truong et al. 2010) and P. nicotianae in the US (Parkunan et al. 2010).
However following the introduction of compatible mating types of P. infestans, recombinant strains quickly replaced previously established strains across much of the Northern Hemisphere (Cooke and Andersson 2013; Drenth et al. 1994; Gavino et al. 2000; Goodwin et al. 1998). In Poland the identification of a novel A2 isolate of P. infestans in 1989 was followed by sexual recombination, and by 1991, novel genotypes dominated the pathogen population (Sujkowski 1994). In North America, the introduction of the A2 mating type in the 1990s resulted in the proliferation of clonal lineages of the novel recombinants US-8, US-11, US-22, US-23 and US-24, believed to have resulted from crosses between US-6 and US-7 (Gavino et al. 2000; Halterman and Gevens 2013; Peters et al. 2014).
Hybridisation
Hybridisation may present an even more immediate and dangerous route to the emergence of new, virulent strains of Phytophthora (Brasier 2000). Phytophthora hybrids have been identified in clades 1, 6, 7 and 8 (Bertier et al. 2013; Bonants et al. 2000; Brasier et al. 2004; Goss et al. 2011; Nirenberg et al. 2009) and as identification techniques continue to improve, these numbers are predicted to increase (Érsek and Man in’t Veld 2013).
While hybridisation is known to occur naturally in populations of Fungi and Oomycota, the chances of it occurring increase following the removal of geographic barriers because previously allopatric species often lack the genetic barriers to mating found in co-occurring species (Brasier 1995; Brasier 2000). Indeed, all the Phytophthora hybrids identified to date have been the offspring of an exotic and a native species (Érsek and Man in’t Veld 2013). In addition to heterothallic species it has also been experimentally shown that homothallic species can outbreed to produce new virulence combinations (Whisson et al. 1994) and that hybridisation can occur between different homothallic species (May et al. 2003).
Hybrids can emerge sexually (Goodwin and Fry 1994), or through mitotic or parasexual recombination (Drenth and Goodwin 1999; Érsek et al. 1995). Successful fusion depends on a number of environmental and genetic factors, including the ability of the introduced taxon to compete in the new environment, the amount of niche contact between parent taxa and finally, their vegetative or sexual compatibility (Fig. 2) (Brasier 1995). Once hybrids form, further diversity may be generated from back-crosses with one or both parents, or as the result of subsequent recombination events and chromosome losses (Fig. 2) (Érsek and Man in’t Veld 2013).
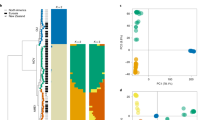
Hybrid progeny can range from being marginally different to the parent species, or they can be entirely new species with a genome that is a combination of both parents (Fig. 2) (Brasier 1991; Donahoo and Lamour 2008). The offspring that possess a combination of chromosomal material from both parent species, sometimes in allopolyploid form, sometimes hybridised in diploid form, are of the greatest interest because of their potential to adapt to new environments and hosts compared to clonally produced offspring (Text Box 1) (Fig. 3). Often a range of genomic strains may result from one set of parents, as is the case with Phytophthora alni, where we see a “swarm” of allopolyploid genotypes (Ioos et al. 2006). Unlike the nuclear DNA, the mitochondrion is uni-parentally inherited in Phytophthora (Érsek and Man in’t Veld 2013; Whittaker et al. 1991). Therefore, while mtDNA cannot be used to identify hybrids, it can be used to identify at least one of the parent species (Man in’t Veld et al. 1998).
Pathogenicity
The genus Phytophthora has proven a challenging group of pathogens to control because of its ability to overcome host resistance (Haas et al. 2009; Bertier et al. 2013). It appears that hybridisation and the often-associated state of polyploidy may enhance pathogenicity (Text Box 1) (de Cock and Lévesque 2004; Jung and Burgess 2009; Rea et al. 2010; Sansome 1977; Sansome and Brasier 1974; Sansome et al. 1991; Win-Tin and DickDick 1975). Because human interventions increase the chances of hybridisation we can anticipate the emergence of novel plant pathogens.
There are many examples of enhanced virulence in hybrid plant pathogens. Often hybrid species are adapted to the combined geographic and host ranges of their parent species. For example, when crosses were made between poplar species Populus deltoides and Populus trichocarpa in the 1980s for commercial purposes, their associated rust pathogens Melampsora medusae and M. occidentalis also appeared to have hybridised (Newcombe et al. 2000). These hybrid pathogens attack both parental species as well as the hybrid. Furthermore, introgression may extend the host ranges of each parental species (Brasier 2001).
Indeed, another risk associated with hybrids is that they can act as genetic bridges between the two previously geographically isolated parental populations, allowing the transfer of virulence genes (Newcombe et al. 2000; Brasier 2001). The two Dutch elm disease pathogens Ophiostoma ulmi and O. novo-ulmi appear to have formed transient hybrids at the epidemic fronts that transferred biologically useful loci from O. ulmi to the more virulent O. novo-ulmi (Brasier et al. 1998).
Finally, hybrids may possess novel virulence. Hybrids of Phytophthora cambivora, a pathogen of hardwoods, with P. fragariae, a pathogen of strawberry and raspberry, acquired virulence on alder, a host that neither of the presumptive parents are able to attack (Érsek et al. 1995; Brasier et al. 1999). Similarly, hybrids of P. hedraiandra and P. cactorum colonise species beyond the host range of either parent (Man in’t Veld et al. 2007).
Recently, the genetic mechanism behind the adaptability and virulence of Phytophthora has been partially unveiled. Like many plant pathogens, Phytophthora species secrete effectors that modify host plant physiology to facilitate colonisation (Haas et al. 2009; Whisson et al. 2007). Both Crinkler (CRN) cytoplasmic effectors and RxLR effector proteins are considered to be major determinants of host specificity and virulence in Phytophthora (Bertier et al. 2013). The genes controlling the relevant effectors are contained in highly variable, gene-sparse and repetition-rich regions of the genome (Haas et al. 2009). This allows homologous recombination between effector gene regions to occur without a high risk of lethality (Bertier et al. 2013). This process is likely to have led to the diversity and abundance of RxLR and CRN genes observed in Phytophthora species (Haas et al. 2009). These dynamic and expanded effector repertoires may enable the pathogenic adaptability of the genus (Vercauteren et al. 2011; Bertier et al. 2013).
Evolutionary implications
Hybridisation and allopolyploidy enable adaptation and speciation and are considered important forces in the evolution of Angiosperm plants, as well as in the evolution, speciation and pathogenicity of Phytophthora (Bertier et al. 2013). Recent analysis showing the presence of many small duplicated blocks of two or three consecutive genes indicates that an ancient whole genome duplication (WGD) created a polyploid ancestor that evolved into P. infestans, P. sojae and P. ramorum (Martens and Van de Peer 2010). Many of the duplicated genes were related to pathogenicity and infection, which suggests that the WGD contributed to the pathogenicity of the genus (Martens and Van de Peer 2010).
Further, evidence suggests newly developed hybrids continue to evolve rapidly (Goss et al. 2011). Evolving strains have now been identified in at least three Phytophthora species (P. alni subsp. alni, P. nicotianae × P. cactorum and P. hedraiandra × P. cactorum) (Man in’t Veld et al. 2007; Érsek and Man in’t Veld 2013). This continuing evolution occurs through intercrossing, introgression, mitotic chromosomal rearrangements and chromosome loss (Érsek and Man in’t Veld 2013). With their short life cycles, the rapid evolution of Phytophthora pathogens gives them a competitive advantage in the evolutionary arms race over their relatively slower evolving hosts, and biosecurity protocols (Clay and Kover 1996; Wuethrich 1998; Érsek and Man in’t Veld 2013).
Future directions
This review highlights the pressing need to improve our capacity to identify and describe Phytophthora hybrids, and exposes a serious flaw in current biosecurity protocols. Due to their rarity, atypical phenotypes and limited number of robust morphological characters, Phytophthora species are often difficult to distinguish (Brasier et al. 1999; Érsek and Man in’t Veld 2013). For example, while P. multivora was only recently described in Western Australia using ITS sequencing, it was found to be abundant in historical collections where it had been mistaken for the phenotypically similar P. citricola (Scott et al. 2009). Phytophthora multivora was found to be the second-most widespread Phytophthora species in Western Australia after P. cinnamomi, and at many sites it is P. multivora rather than P. cinnamomi that causes tree mortality. This finding has significant implications for controlling the spread of Phytophthora dieback.
Although molecular techniques provide better resolution for distinguishing hybrids and species, they are not always reliable. ITS-based techniques, for example, may homogenise hybrid rDNA arrays (Brasier et al. 1999; Bertier et al. 2013). Therefore to confidently distinguish species and hybrids it is necessary to analyse several independent loci in conjunction with morphological and physiological assessments (Man in’t Veld et al. 2007).
Additionally, following identification, there remains the issue of how to taxonomically name and define hybrids or hybrid complexes for legal and biosecurity purposes (Brasier et al. 1999). As more hybrids are described it becomes increasingly important to improve detection methods and develop consistent rules for nomenclature.
There is also much to be learnt regarding the rapid evolution and hybridisation of introduced Phytophthora species and other pathogens. For instance, a better understanding of the factors leading to hybridization, regulating virulence and host range in hybrid progeny and an understanding of how novel traits arise in hybrid progeny is required. For example, many examples of hybridisation appear to have occurred in water, either following heavy flooding or in hydroponic systems (Érsek and Man in’t Veld 2013). Such research will lead to better risk assessment and management of novel pathogen species.
Novel Phytophthora species and hybrids pose a serious biosecurity threat as trade in plant material inevitably increases. Biosecurity and control protocols focus on the risks posed by known, rather than unknown pathogens (Brasier 2008; Scott et al. 2009). Yet, the examples of P. ramorum and P. cinnamomi, amongst others, show that species that are benign in their centre of origin can become destructive in new environments as the result of episodic selection, hybridisation or perhaps through becoming part of a novel disease complex (Brasier 2008). Even when pathogens are known as minor pathogens in certain parts of the world the occurrence of hybridization among endemic and exotic species may produce hybrid species with novel pathogenicity and virulence characteristics as has been shown for the Phytophthora on alder (Brasier and Kirk 2001; Érsek et al. 1995). These organisms are difficult to detect in their native environments because native hosts show few or no symptoms of disease, and so are overlooked during the risk analysis processes. The current biosecurity system and their current basis for risk assessment must therefore be revised to consider the potential for benign microbes to cause damage when introduced to different environments.
Conclusion
The emergence of new, highly virulent Phytophthora species in unexpected places, whether they are undescribed or previously known, has fundamental implications for current biosecurity and control practices. There is now a strong body of evidence to suggest that exotic species have the capacity to evolve rapidly, sometimes via hybridisation, and that this process can lead to enhanced pathogenicity and virulence (Brasier 1995; Newcombe et al. 2000; Brasier 2000). Although hybridisation has probably always been an important evolutionary force for Phytophthora, the probability that it will happen increases as more infected and uninfected plants are traded and planted outside their natural geographic range, and new diseases may arise as a result. This review highlights the importance of continued monitoring of natural and anthropomorphised systems in order to identify novel Phytophthora species, their ecological role and if applicable, their pathogenicity. Assessment of risk and assessment of impacts due to introductions of exotic Phytophthora species and the consequences of hybridisations needs to be taken in consideration by Biosecurity agencies if they desire to make sound decisions based on established principles of invasion biology.
References
Ahmed S, de Labrouhe DT, Delmotte F (2012) Emerging virulence arising from hybridisation facilitated by multiple introductions of the sunflower downy mildew pathogen Plasmopara halstedii. Fungal Genet Biol 49(10):847–855
Arneson PA (2011) Coffee rust. Plant Health Instr. doi:10.1094/PHI-I-2000-0718-02
Bertier L, Leus L, D’hondt L, de Cock AWAM, Hofte M (2013) Host adaptation and speciation through hybridisation and polyploidy in Phytophthora. PLoS 8(12):e85385
Blair JE, Coffey MD, Park S-Y, Geiser DM, Kang S (2008) A multi-locus phylogeny for Phytophthora utilizing markers derived from complete genome sequences. Fungal Genet Biol 45:266–277
Bonants PJM, Hagenaar-de Weerdt M, Man in’t Veld WA, Baayen RP (2000) Molecular characterization of natural hybrids of Phytophthora nicotianae and P. cactorum. Phytopathology 90:867–874
Brasier CM (1991) Current questions in Phytophthora systematics: the role of the population approach. In: Phytophthora. Ed. by Lucas, JA, Shattock RC, Shaw DS and Cooke LR. Cambridge University Press, Cambridge. pp. 104–128
Brasier CM (1995) Episodic selection as a force in fungal microevolution, with special reference to clonal speciation and hybrid introgression. Can J Bot 73(1):1213–1221
Brasier CM (2000) The rise of the hybrid fungi. Nature 405:134–135
Brasier CM (2001) Rapid evolution of introduced plant pathogens via interspecific hybridization. Bioscience 51:123–133
Brasier CM (2008) The biosecurity threat to the UK and global environment from international trade in plants. Plant Pathol 57:792–808
Brasier CM (2009) Phytophthora biodiversity: How many Phytophthora species are there? In (Goheen, E.M. and Frankel, S.J. Eds).Phytophthoras in Forests and Natural Ecosystems. Proceedings of the Fourth International Union of Forest Research Organisations (IUFRO) Working Party 7.02.09. General Technical Report PSW-GTR-221. Albany, CA: USDA Forest Service, Pacific Southwest Research Station, pp. 101–115
Brasier CM, Kirk SA (2001) Comparative aggressiveness of standard and variant hybrid alder phytophthoras, Phytophthora cambivora and other Phytophthora species on bark of Alnus, Quercus and other woody hosts. Plant Pathol 50:218–229
Brasier CM, Kirk SA, Pipe N, Buck KW (1998) Rare hybrids in natural populations of the Dutch elm disease pathogens Ophiostoma ulmi and O. novo-ulmi. Mycol Res 102:45–57
Brasier CM, Cooke D, Duncan JM (1999) Origin of a new Phytophthora pathogen through interspecific hybridization. Proc Natl Acad Sci 96:5878–5883
Brasier CM, Sanchez-Hernandez E, Kirk SA (2003) Phytophthora inundata sp. nov., a part heterothallic pathogen on trees and shrubs in wet or flooded soils. Mycol Res 107:477–484
Brasier CM, Kirk SA, Delcan J, Cooke DEL, Jung T, Man in’t Veld WA (2004) Phytophthora alni sp. nov. and its variants: designation of emerging heteroploid hybrid pathogens spreading on Alnus trees. Mycol Res 108:1172–1184
Burgess TI, Webster JL, Ciampini JA, White D, Hardy GESJ, Stukely MJC (2009) Re-evaluation of Phytophthora species isolated during 30 years of vegetation health surveys in Western Australia using molecular techniques. Plant Dis 93(3):215–223
Clay K, Kover PX (1996) The red queen hypothesis and plant/pathogen interactions. Annu Rev Phytopathol 34:29–50
Comai L (2005) The advantages and disadvantages of being polyploid. Nat Rev Genet 6:836–846
Cooke DEL and Andersson B (2013) Phytophthora infestans and potato blight in Europe. Phytophthora: a global perspective Ed. Lamour, K. CABI Plant Protection Series, No.2. pp. 59–64
Cooke DEL, Duncan JM (1997) Phylogenetic analysis of Phytophthora species based on ITS1 and ITS2 sequences of the ribosomal RNA gene repeat. Mycol Res 101:667–677
Cooke DEL, Drenth A, Duncan JM, Wagels G, Brasier CM (2000) A molecular phylogeny of Phytophthora and related oomycetes. Fungal Genet Biol 30:17–32
Davison EM, Drenth A, Kumar S, Mack S, Mackie AE, McKirdy S (2006) Pathogens associated with nursery plants imported into Western Australia. Aust Plant Pathol 35:473–475
de Cock AWAM, Lévesque CA (2004) New species of Pythium and Phytophthora. Stud Mycol 50:481–487
Dobrowolski M, Tommerup I, Shearer B, O’Brien P (2003) Three clonal lineages of Phytophthora cinnamomi in Australia revealed by microsatellites. Phytopathology 93:695–704
Donahoo RS, Lamour KH (2008) Interspecific hybridization and apomixis between Phytophthora capsici and Phytophthora tropicalis. Mycologia 100:911–920
Drenth A, Goodwin SB (1999) Population structure: Oomycetes. In: Worral JJ (ed) Structure and Dynamics of Fungal Populations. Kluwer Academic Publishers, The Netherlands, pp. 195–224
Drenth A, Guest D (2013) Phytophthora palmivora in tropical tree crops. In: Lamour, K (ed) Phytophthora: a global perspective. CABI Plant Protection Series, No.2 pp. 187–196
Drenth A, Tas ICQ, Govers F (1994) DNA fingerprinting uncovers a new sexually reproducing population of Phytophthora infestans in The Netherlands. Eur J Plant Pathol 100:97–107
Érsek T, Man in’t Veld WA (2013) Phytophthora species hybrids: a novel threat to crops and natural ecosystems. In: Lamour, K (ed) Phytophthora: a global perspective. CABI Plant Protection Series, No.2. pp. 37–47
Érsek T, Ribeiro OK (2010) An annotated list of new Phytophthora species described post-1996. Acta Phytopathol Entomol Hung 45:251–266
Érsek T, English JT, Schoelz JE (1995) Creation of species hybrids of Phytophthora with modified host ranges by zoospore fusion. Phytopathology 85:1343–1347
Facon B, Genton BJ, Shykoff J, Jarne P, Estoup A, David P (2006) A general eco-evolutionary framework for understanding bioinvasions. Trends Ecol Evol 21:130–135
Förster H, Cummings MP, Coffey MD (2000) Phylogenetic relationships of Phytophthora species based on ribosomal ITS I DNA sequence analysis with emphasis on Waterhouse groups V and VI. Mycol Res 104:1055–1061
Fry WE, Goodwin SB, Dyer AT, Matuszak JM, Drenth A, Tooley PW, Sujkowski LS, Koh YJ, Cohen BA, Spielman LJ, Deahl KL, Inglis DA, Sandlan KP (1993) Historical and recent migrations of Phytophthora infestans: chronology, pathways and implications. Plant Dis 77:653–661
Gaeta RT, Chris Pires J (2010) Homoeologous recombination in allopolyploids: the polyploid ratchet. New Phytol 186:18–28
Gavino PD, Smart CD, Sandrock RW, Miller JS, Hamm PB, Lee TY, Davis RM, Fry WE (2000) Implications of sexual reproduction for Phytophthora infestans in the United States: generation of an aggressive lineage. Plant Dis 84:731–735
Goodwin SB, Fry WE (1994) Genetic analysis of interspecific hybrids between Phytophthora infestans and Phytophthora mirabilis. Exp Mycol 18:20–32
Goodwin SB, Smart CD, Sandrock RW, Deahl KL, Punja ZK, Fry WE (1998) Genetic change within populations of Phytophthora infestans in the United States and Canada during 1994 to 1996: role of migration and recombination. Phytopathology 88:939–949
Goss EM, Cardenas ME, Myers K, Forbes GA, Fry WE, Restrepo S, Grünwald NJ (2011) The plant pathogen Phytophthora andina emerged via hybridization of an unknown Phytophthora species and the Irish potato famine pathogen, P infestans. PLoS One 6:e24543
Grunwald NJ, Werres S, Goss EM, Taylor CR, Fieland VJ (2012) Phytophthora obscura sp. nov., a new species of the novel Phytophthora subclade 8d. Plant Pathol 61:610–622
Haas BJ, Kamoun S, Zody MC, Jiang RHY, Handsaker RE, et al. (2009) Genome sequence and analysis of the Irish potato famine pathogen Phytophthora infestans. Nature 461:393–398
Halterman D, Gevens AJ (2013) Phytophthora infestans in the US. In: Lamour, K (ed) Phytophthora: a global perspective. CABI Plant Protection Series, No.2. pp. 68–78
Heybroek HM (1993) The Dutch Elm Breeding Program. In: Sticklen & Sherald (Eds.) Dutch Elm Disease Research, Chapter 3. Springer Verlag, New York
Huberli D, Hardy GESJ, White D, Williams N, Burgess TI (2013) Fishing for Phytophthora from western Australia waterways: a distribution and diversity survey. Australas Plant Pathol 42:251–260
Ioos R, Andrieux A, Marçais B, Frey P (2006) Genetic characterization of the natural hybrid species Phytophthora alni as inferred from nuclear and mitochondrial DNA analyses. Fungal Genet Biol 43:511–529
Jung T, Burgess TI (2009) Re-evaluation of Phytophthora citricola isolates from multiple woody hosts in Europe and North America reveals a new species, Phytophthora plurivora sp. nov. Persoonia 22:95–110
Jung T, Stukely MJC, Hardy GESJ, White D, Paap T, Dunstan WA, Burgess TI (2011) Multiple new Phytophthora species from ITS Clade 6 associated with natural ecosystems in Australia: evolutionary and ecological implications. Persoonia 26:13–39
Kaschani F, Van der Hoorn RAL (2011) A model of the C14-EPIC complex indicates hotspots for a protease-inhibitor arms race in the oomycete-potato interaction. Plant Signal Behav 6(1):109–112
Linde C, Drenth A, Kemp GHJ, Wingfield MJ, Von Broembsen S (1997) Population structure of Phytophthora cinnamomi in South Africa. Phytopathology 87:822–827
Linderman RG, Davis EA (2006) Evaluation of chemical and biological agents for control of Phytophthora species on intact plants or detached leaves of rhododendron and lilac. In: Frankel, S.J., Shea, P.J. and Haverty, M.I. (eds) Proceedings of the Sudden Oak Death Second Science Symposium: The State of Our Knowledge. General Technical Report PSW-GTR-196. United States Department of Agriculture (USDA) Forest Service, Pacific Southwest Research Station, Albany, California, pp. 265– 268
Man in’t Veld WA, Venbaas-Rijks WJ, Ilieva E, de Cock AWAM, Bonants PJM, Pieters R (1998) Natural hybrids of Phytophthora nicotianae and Phytophthora cactorum demonstrated by isozyme analysis and random amplified polymorphic DNA. Phytopathology 88:922–929
Man in’t Veld WA, de Cock WAM, Summerbell RC (2007) Natural hybrids of resident and introduced Phytophthora species proliferating on multiple new hosts. Eur J Plant Pathol 117:25–33
Man in’t Veld WA, Rosendahl KCHM, Hong C (2012) Phytophthora serendipita sp. nov. and P. pelgrandis, two destructive pathogens generated by natural hybridization. Mycologia 104:1390–1396
Martens C, Van de Peer Y (2010) The hidden duplication past of the plant pathogen Phytophthora and its consequences for infection. BMC Genomics 11:353
May KJ, Drenth A, Irwin JAG (2003) Interspecific hybrids between the homothallic Phytophthora sojae and Phytophthora vignae. Australas J Plant Pathol 32:353–359
Mayr E (1942) Systematics and the Origin of Species, from the Viewpoint of a Zoologist. Harvard University Press, Cambridge
Newcombe G, Stirling B, McDonald S, Bradshaw JR (2000) Melampsora x columbiana, a natural hybrid of M. medusae and M. occidentalis. Mycol Res 104:261–274
Nirenberg HI, Gerlach WF, Gräfenhan T (2009) Phytophthora x pelgrandis, a new natural hybrid pathogenic to Pelargonium grandiflorum hort. Mycologia 101:220–231
Parkunan V, Johnson CS, Bowman BC, Hong CX (2010) Population structure, mating type, and mefenoxam sensitivity of Phytophthora nicotianae in Virginia tobacco fields. Plant Dis 94:1361–1365
Paun O, Fay MF, Soltis DE, Chase MW (2007) Genetic and epigenetic alterations after hybridization and genome doubling. Taxon 56:649–656
Peters RD, Al-Mughrabi KI, Kalischuk ML, Dobinson KF, Conn KL, Alkher H, Islam MR, Daayf F, Lynn J, Bizimungu B, De Koeyer D, Lévesque CA, Kawchuk LM (2014) Characterization of Phytophthora infestans population diversity in Canada reveals increased migration and genotype recombination. Can J Plant Pathol 36:73–82
Podger FD (1968) Aetiology of jarrah dieback and disease of dry sclerophyll Eucalyptus marginata Sm. forests in Western Australia. MSc Thesis, University of Western Australia
Rea A, Jung T, Burgess TI, Stukely MJC, Hardy GESJ (2010) Phytophthora elongata sp. nov. a novel pathogen from the Eucalyptus marginata forest of Western Australia. Australas Plant Pathol 39:477–491
Ristaino JB (2002) Tracking historic migrations of the Irish potato famine pathogen, Phytophthora infestans. Microbes Infect 4:1369–1377
Runge F, Telle S, Ploch S, Savory E, Day B, Sharma R, Thines M (2011) The inclusion of downy mildews in a multi-locus-dataset and its reanalysis reveals a high degree of paraphyly in Phytophthora. IMA Fungus Glob Mycol J 2:163–171
Sansome E (1977) Polyploidy and induced gametangial formation in British isolates of Phytophthora infestans. J Genet Microbiol 99:311–316
Sansome E, Brasier CM (1974) Polyploidy associated with varietal differentiation in the megasperma complex of Phytophthora. Trans Br Mycol Soc 63:461–467
Sansome E, Brasier CM, Hamm PB (1991) Phytophthora meadii may be a species hybrid. Mycol Res 95:273–277
Scott PM, Burgess TI, Barber PA, Shearer BL, Stukely MJ, Hardy GE, Jung T (2009) Phytophthora multivora sp. nov., a new species recovered from declining Eucalyptus, Banksia, Agonis and other plant species in Western Australia. Persoonia 22:1–13
Scott P, Burgess T, Hardy G (2013) Globalisation and Phytophthora. In Lamour, K (ed) Phytophthora: a global perspective. CABI Plant Protection Series, No.2. pp. 37–47
Sujkowski LS (1994) Increased genotypic diversity via migration and possible occurrence of sexual reproduction of Phytophthora infestans in Poland. Phytopathology 84:201–207
Truong NV, Liew EC, Burgess LW (2010) Characterisation of Phytophthora capsici isolates from black pepper in Vietnam. Fungal Biol 114:160–170
Tyler BM, Tripathy S, Zhang X, Dehal P, Jiang RH, et al. (2006) Phytophthora genome sequences uncover evolutionary origins and mechanisms of pathogenesis. Science 313(5791):1261–1266
Vercauteren A, Boutet X, D’hondt L, Van Bockstaele E, Maes M, Leus L, Chandelier A, Heungens K (2011) Aberrant genome size and instability of Phytophthora ramorum oospore progenies. Fungal Genet Biol 48(5):537–543
Villa NO, Kageyama K, Asano T, Suga H (2006) Phylogenetic relationships of Pythium and Phytophthora species based on ITS rDNA, cytochrome oxidase II and beta-tubulin gene sequences. Mycologia 98:410–422
Wellings CR, McIntosh RA (1990) Puccinia striiformis f.sp. tritici in Australasia: pathogenic changes during the first ten years. Plant Pathol 39:316–325
Whisson SC, Drenth A, Maclean DJ, Irwin JAG (1994) Evidence for outcrossing in Phytophthora sojae and linkage of a DNA marker to two avirulence genes. Curr Genet 27:77–82
Whisson SC, Boevink PC, Moleleki L, Avrova AO, Morales JG, Gilroy EM, Armstrong MR, Grouffaud S, van West P, Chapman S, Hein I, Toth IK, Pritchard L, Birch PR (2007) A translocation signal for delivery of oomycete effector proteins into host plant cells. Nature 450:115–118
Whittaker SL, Shattock RC, Shaw DS (1991) Inheritance of DNA contents in sexual progenies of Phytophthora infestans. Mycol Res 95:1094–1100
Win-Tin, Dick MW (1975) Cytology of Oomycetes. Arch Microbiol 105:283–293
Wuethrich B (1998) Why sex? Putting theory to the test. Science 281:1980–1982
Yang X, Copes WE, Hong CX (2014) Two novel species representing a new clade and cluster of Phytophthora. Fungal Biol 118:72–82
Acknowledgments
We thank Professor André Drenth for his insightful and constructive discussions and comments.
Author information
Authors and Affiliations
Corresponding author
Additional information
Novel pathogenic Phytophthora species are emerging as the result of the rapid evolution of introduced species, and pose an increasing biosecurity risk.
Rights and permissions
About this article
Cite this article
Callaghan, S., Guest, D. Globalisation, the founder effect, hybrid Phytophthora species and rapid evolution: new headaches for biosecurity. Australasian Plant Pathol. 44, 255–262 (2015). https://doi.org/10.1007/s13313-015-0348-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13313-015-0348-5